Research Article
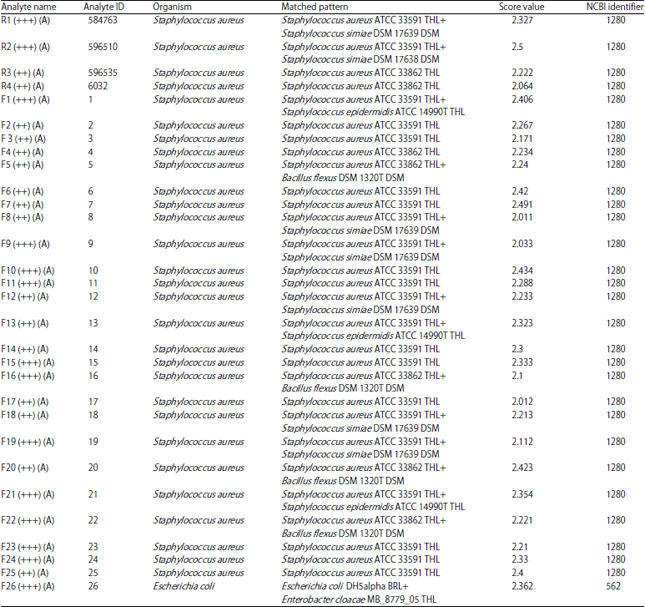
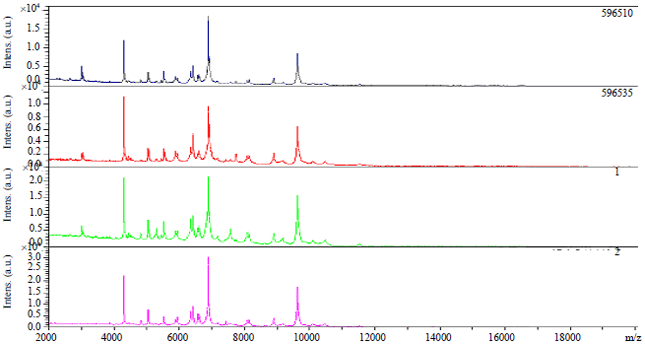
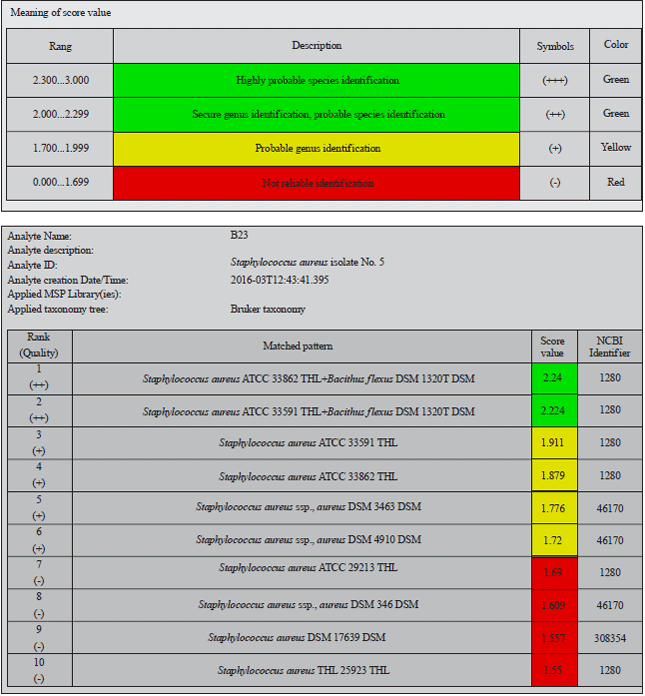
Identification of Staphylococcus aureus Causing Bovine Mastitis using MALDI-TOF Fingerprinting
Central Laboratory for Evaluation of Veterinary Biologics (CLEVB),
M. L. Sayed
Central Laboratory for Evaluation of Veterinary Biologics (CLEVB),
Aatia A. El-Gedawy
Animal Health Research Institute (AHRI),
Ghada M. El Sadek
Central Laboratory for Evaluation of Veterinary Biologics (CLEVB),
A. A. Samy
Department of Microbiology and Immunology, National Research Center (NRC), Dokki, Egypt
Abdelhakam M.M. Ali
Central Laboratory for Evaluation of Veterinary Biologics (CLEVB),