Research Article
Use of Starter Cultures of Lactic Acid Bacteria and Yeasts in the Preparation of Kisra, a Sudanese Fermented Food
Food Research Centre, P.O. Box 213, Sudan
Muna M. Mustafa
Food Research Centre, P.O. Box 213, Sudan
Lactic Acid Bacteria (LAB) and yeasts have been reported to be predominant microorganisms in most of the African indigenous fermented foods (Nout, 1991; Halm et al., 1993; Hounhouigan et al., 1993; Sanni, 1993; Steinkraus, 1996; Olasupo et al., 1997; Nago et al., 1998 and Kunene et al., 2000). Stable co-metabolism between LAB and yeasts is common in many foods, enabling the utilization of substances that are otherwise non fermentable (for example starch) and thus increasing the microbial adaptability to complex food ecosystems (Gobbetti et al., 1994; Stolz et al., 1995 and Gobbetti and Corsetti, 1997).
It has been suggested that the proliferation of yeasts in foods is favoured by the acidic environment created by LAB while the growth of bacteria is simulated by the presence of yeasts, which may provide growth factors, such as, vitamins and soluble nitrogen compounds (Nout, 1991). The association of LAB and yeasts during fermentation may also contribute metabolites, which could impart taste and flavour to foods (Akinrele, 1970; Halm et al., 1993; Brauman et al., 1996 and Hansen and Hansen, 1996).
According to Sanni (1993) and Kimaryo et al. (2000), the use of the starter cultures would be an appropriate approach for the control and optimization of the fermentation process in order to alleviate the problems of variations in organoleptic quality and microbiological stability observed in African indigenous fermented foods. No LAB starter cultures are commercially available yet for the small-scale processing of traditional African foods (Holzapfel, 1997).
The development of starter cultures is one of the prerequisites for the establishment of small-scale industrial production of fermented foods in Africa (Sanni, 1993). The primary consideration before introducing starter cultures for traditional small-scale fermentations should be whether these would significantly contribute to an improvement of processing conditions and product quality with respect to: rapid or accelerated acidification, an improved and more predictable fermentation process, desirable sensory attributes, improved safety and reduction of hygienic risks (Holzapfel, 1997). Therefore, a thorough understanding of the fermentation process is required. The knowledge gained with the controlled starter culture may also benefit those operating at a very small scale and practicing back slopping.
Kisra is an indigenous staple food of the majority of Sudanese people. It is a pancake-like bread made from sorghum or millet flour. Kisra fermentation is a traditional process, whereby sorghum or millet flour is mixed with water in a ratio of about 1:2 (w/v), usually a starter is added by a back-slopping using mother dough from a previous fermentation as a starter at a level of about 10%. Fermentation is completed in about 12-19 hours by which time the pH drops from about six to less than four. Due to the tedious process of kisra preparation, most of the population abandoned kisra consumption and shifted to bread.
Enumeration and characterization of LAB and yeasts: Duplicate samples of traditional starter (10 ml) were homogenized with 90 ml sterile peptone water. The homogenate was decimal diluted and the relevant dilutions surface plated. MRS agar (Oxoid) plated were incubated anaerobically (BBL Gas Pak, Becton Dickinson, Cockeysrille, MD, USA) for 48 h at 37oC for the enumeration and isolation of Lactic Acid Bacteria (LAB). Total of 42 representative colonies were randomly picked from higher dilution plates of various fermentation stages and confirmed to be Gram-positive and catalase-negative. For subsequent purification and sub-culturing MRS agar and broth were used. The pure bacterial cultures were inculcated into appropriate broth (MRS broth), incubated for 24 h at 37oC, centrifuged at 3000 rpm for 15 min and the supernatant decanted. The cell pellets were re-suspended in sterile MRS broth containing 10% (v/v) glycerol. The suspension was aseptically transferred into sterile cryo-tubes containing acid-washed glass beads and stored at -80oC until required for identification. Yeasts were enumerated and isolated after incubation for 3-4 days at 28oC on malt extract agar containing 0.01% (w/v) chloramphenicol (selective supplement, Oxoid).
Purification and sub-culturing was done using malt extract broth. The purified yeast cultures were stored on malt extract slants at 4oC until required for identification and further use.
The bacteria were characterized by microscopic examination and by conventional biochemical and physiological tests. The cultures were examined for colony and cell morphology; motility, cell arrangement, Gram reaction; catalase reaction; growth in broth at 10, 15, 40 and 45oC; growth in presence of 2, 4 and 6.5% (w/v) NaCl; production of amino from arginine; production of dextran from sucrose and production of carbon dioxide from glucose using Gibson's litmus milk. These tests were done according to procedures described by Harrigan and McCance (1976). Preliminary grouping for selection of 42 isolates for API tests was based on the above-mentioned morphological, physiological and biochemical characteristics. The fermentation pattern among carbohydrates was determined by using the API 20A gallery with the API 20 A medium (Bio Merieux, Marcy, France). Anaerobiosis in the inoculated tubes was obtained by overlaying with sterile paraffin oil. The inculcated galleries were incubated at 37oC and the observations were made after 24 and 48 h. The identification of isolates was facilitated by the use of a computer programme, API LAB PLUS, version 3.2.2 (Bio Merieux) and reference to Bergey's Manual of Systematic Bacteriology (Sneath et al., 1986) and Wood and Holzapfel (1995).
Pure isolates of yeasts were identified according to Barrnett et al. (1983) and Barrnett et al. (1990). The culture was examined microscopically after incubation at 28oC for 72 h. The shapes of the yeast cells and the form of budding were observed and registered (Barrnett et al., 1983). Each isolate was then inoculated in-Potato Dextrose Agar (PDA) and incubated at 28oC for one to four weeks and then they were examined microscopically for ascospore formation (Barrnett et al., 1983). YMA media was inoculated with fresh culture of yeast and incubated aerobically at 37oC and 42oC (Harrigan and McCance, 1976). Resistance to cyclohexamide and the fermentation patterns among Glucose, Maltose, Galactose, Sucrose, Lactose and Fructose were carried out according to (Harrigan, 1998).
Preparation of starter cultures: The cultures of LAB (Lactobacillus fermentum, Lactobacillus amylovorus, Lactobacillus brevis) and yeast (Saccharomyces cerevisiae) isolated from household fermented dough were used. The cultures had been stored at -80°C in sterile cryo-tubes containing MRS broth with 10% (v/v) glycerol and acid-washed glass beads until required. LAB were cultivated by streaking on MRS agar (Oxoid) and incubated anaerobically (BBL, Gas Pak, Becton Dickinson) at 37oC for 24 h. A colony was picked from each pure culture plate, grown successively in MRS broth before centrifugation at 655 g/15 min. The pellet was washed in sterile distilled water centrifuged again and redistributed in distilled water. This procedure achieved a culture preparation containing 109 colony- forming units cfu/ml, checked as viable count on MRS agar. Pure cultures of Saccharomyces cerevisiae were cultivated by streaking on malt extract agar (Oxoid), incubated at 37oC for 48 h and the picked colony was inoculated into 10 ml of malt extract broth (Oxoid) and incubated at 28oC for 24 h. These cultures were centrifuged and washed as described above.
This procedure achieved a culture preparation containing 107 cfu/ml, as viable count on malt extract agar. Yeast cultures had been stored on malt extract agar slants at 4oC until required. The number of LAB and yeasts was monitored during fermentation by serial dilution of the samples, using the media described above.
Preparation of samples: This was carried out under sterile conditions and ingredients according to the traditional procedure. The samples were prepared by mixing 50 gm sterile sorghum flour with 100 ml sterile water in a rate of 1:2 (w/v).
Fermentation: In controlled fermentations, 100 ml of sterile sorghum dough was inoculated with 10 ml of previously obtained lactobacillus bacteria. The resultant fermented dough was used as a stock starter for another dough prepared as follows:
| • | To 500 g of sterile flour, 1 litre of sterile water was added. The dough was throughly mixed with glass rod at room temperature in a controlled sterile atmosphere. |
| • | 10% (50 g) from the stock starter were added, mixed and incubated at 37oC for 4 h. |
At zero, 2, 4, 6, 8 and 10 h of fermentation intervals, sub-samples were aseptically drawn for pH determination and microbiological analysis. Another 100 ml of sample was prepared using mixture of Lactobacillus and yeast and subjected to pH and microbiological analysis at 2 h intervals.
Statistical analysis: All determinations were performed in triplicate. All statistical analysis were conducted using nonparametric one-way ANOVA procedures to separate the responses of different microorganisms. All differences were reported at a significance level of α = 0.05.
Phenotyping of starter cultures isolated from fermented sorghum dough: Among the 42 lactic acid bacteria isolated 52% (22 isolates) produced gas from glucose and were referred to as hetero-fermentative, whereas 48% (20 isolates) were homo-fermentative. All of the 22 hetero-fermentative lactic isolates were arginine positive. Six of these were able to grow at 15oC, fermented glucose, maltose and xylose and they were tentatively identified as L. brevis. The remaining 16 of the arginine positive hetero-fermentative lactic isolates were not able to grow at 15oC and fermented mannose tentatively identified as L. fermentum.
From 42 isolates obtained from the fermented sorghum (both traditionally and laboratory-prepared), 20 did not produce gas from glucose and showed homo-fermentative characteristics as mentioned above. These homo-fermentative rod shaped isolates were tentatively identified as L. amylovorus. (Table 1).
The lactic acid bacteria identified in fermented dough have been reported in other fermented foods. L. ermentum and L. brevis have been suggested to be the predominating microorganisms during fermentation of fufu and ogi, two Nigerian foods (Adegoke and Babalola, 1988), kisra a Sudanese sorghum fermented flat bread (Mohammed et al., 1991; Abd Elmoniem et al., 1994), kenkey, a Ghananian fermented maize dough (Halm et al., 1993).
L. fermentum and L. amylovorus have been suggested to be the predominating microorganisms during fermentation of sorghum dough in Sudanese kisra (Halm et al., 1993). In togwa, a Tanzanian fermented food prepared from sorghum, maize, millet and maize- sorghum, the microorganisms were tentatively identified as Lactobacillus plantarum, Lactobacillus brevis, Lactobacillus fermentum, Pediococcus pentosaceus and yeast Issatchenkia orientalis, Saccharomyces cerevisiae and Candida tropicalis (Mugula et al., 2003) and in ogi, a fermented porridge prepared from sorghum in West Africa, the main microorganisms involved were Lactobacillus spp. particularly Lactobacillus plantarum and two yeasts Candida krusei and Debaryomyces hensenii (Odunfa and Adeyele, 1985).
These dominant microorganisms in kisra fermentation seem to be found in other indigenous fermented foods in the genus but not the species. Lei and Jakobsen (2004) reported that the dominating lactic acid bacteria from five Koko production sites in Northern Ghana prepared from millet were found to be Weisella confuse and Lactobacillus fermentum, followed by the less dominant Lactobacillus salivarius and Pediococcus spp.
The yeast populations observed were Saccharomyces cerevisiae. The association of yeasts and lactic acid bacteria is known from a wide variety of traditional food and beverage fermentations (Soni et al., 1985; Adegoke and Babalola, 1988; Sakai and Caldo, 1985).
Yeasts have also been reported to make a useful contribution to the improvement of flavour and acceptability of fermented cereal gruels Banigo et al. (1974), Odunfa and Adeyele (1985) and Akinrele (1970) reported the contribution of S. cerevisiae and Candida mycoderma to the flavour acceptability of ogi. S. cerevisiae proliferated at the beginning while the latter was predominant at the end of fermentation.
| Table 1: | Differential characteristics of four isolates from fermented sorghum based on API 20 A CHL analysis |
 | |
| +, positive reaction; -, no reaction | |
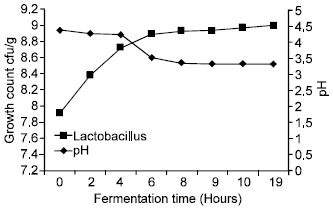 | |
| Fig. 1: | Laboratory fermentation using pure Lactobacillus in dabar flour |
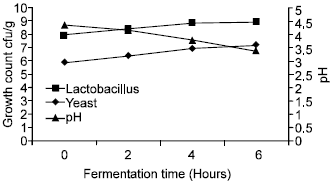 | |
| Fig. 2: | Laboratory fermentation using pure Lactobacillus and yeasts in dabar flour |
pH and microbial numbers: The initial pH of the dough was 4.33, when Lactic acid bacteria growth as a monoculture, pH decreased from 4.33-3.47 after 6 h. Similar findings were obtained by Farahat (1998) who found that using different strains of Lactobacillus to ferment dabar resulted in dropping pH to 3.7 (Fig. 1). When using starter dough from the isolated Lactobacillus strains and pure yeast, the microbial population in a significant increase from 8.31 x 107 cfu/g initial count of Lactobacillus to 8.20 x 108 cfu/g and the pH values in a significant decrease from 4.32-3.75 after 4 h in dabar flour (Fig. 2). These encouraging results show potential efficiency of using pure starter culture dough instead of the traditional starter where fermentation takes about 19 h to reach these values.
The number of LAB and yeasts differed among household samples and increased during laboratory fermentation. Gobbetti et al. (1994) and Steinkraus (1996) proposed that lactic acid bacteria create an acidic environment conducive to yeast proliferation while the yeasts provide vitamins and other growth factors such as amino acids for the lactic acid bacteria. The simultaneous increase in numbers of both LAB and yeasts may therefore be attributed to their symbiotic association. The results are in agreement with those reported by other authors (Mbugua, 1984; Odunfa and Adeyele, 1985; Mohammed et al., 1991; Nche et al., 1994). Melaku and Faulks (1988) also indicated that numbers of LAB increased during the first stages of the natural fermentation with a slight reduction in number during the later stages of fermentation.
The decrease in pH and increase in lactic acid followed the same trend as reported for other traditionally fermented foods (Mohammed et al., 1991; Choi et al., 1994; Dziedzoaze et al., 1996).