Research Article
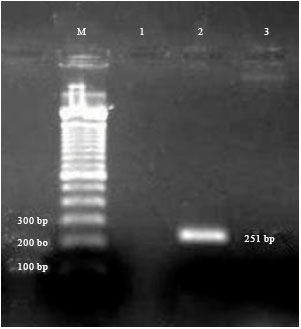
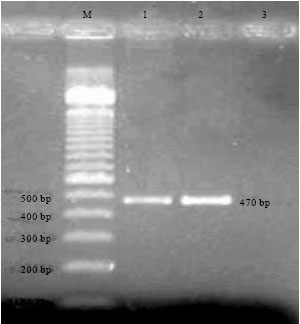
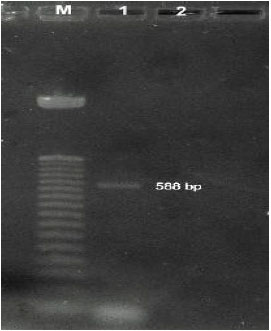
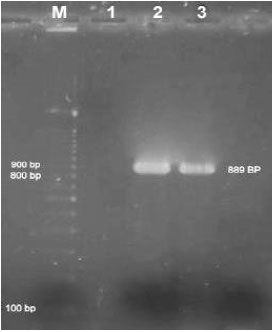
Multiplex PCR (Polymerase Chain Reaction) Assay for Detection of E. coli O157:H7, Salmonella sp., Vibrio cholerae and Vibrio parahaemolyticus in Spiked Shrimps (Penaeus monodon)
Industrial Microbiology Laboratory, Institute of Food Science and Technology, Bangladesh Council of Scientific and Industrial Research, Dhaka, Bangladesh
Mahmuda Sultana
Biotechnology Program, Department of Mathematics and Natural Sciences, BRAC University, Dhaka, Bangladesh
Monzur Morshed Ahmed
Industrial Microbiology Laboratory, Institute of Food Science and Technology, Bangladesh Council of Scientific and Industrial Research, Dhaka, Bangladesh
Abhijit Chowdhury
Industrial Microbiology Laboratory, Institute of Food Science and Technology, Bangladesh Council of Scientific and Industrial Research, Dhaka, Bangladesh
Naiyyum Choudhury
Biotechnology Program, Department of Mathematics and Natural Sciences, BRAC University, Dhaka, Bangladesh