Research Article
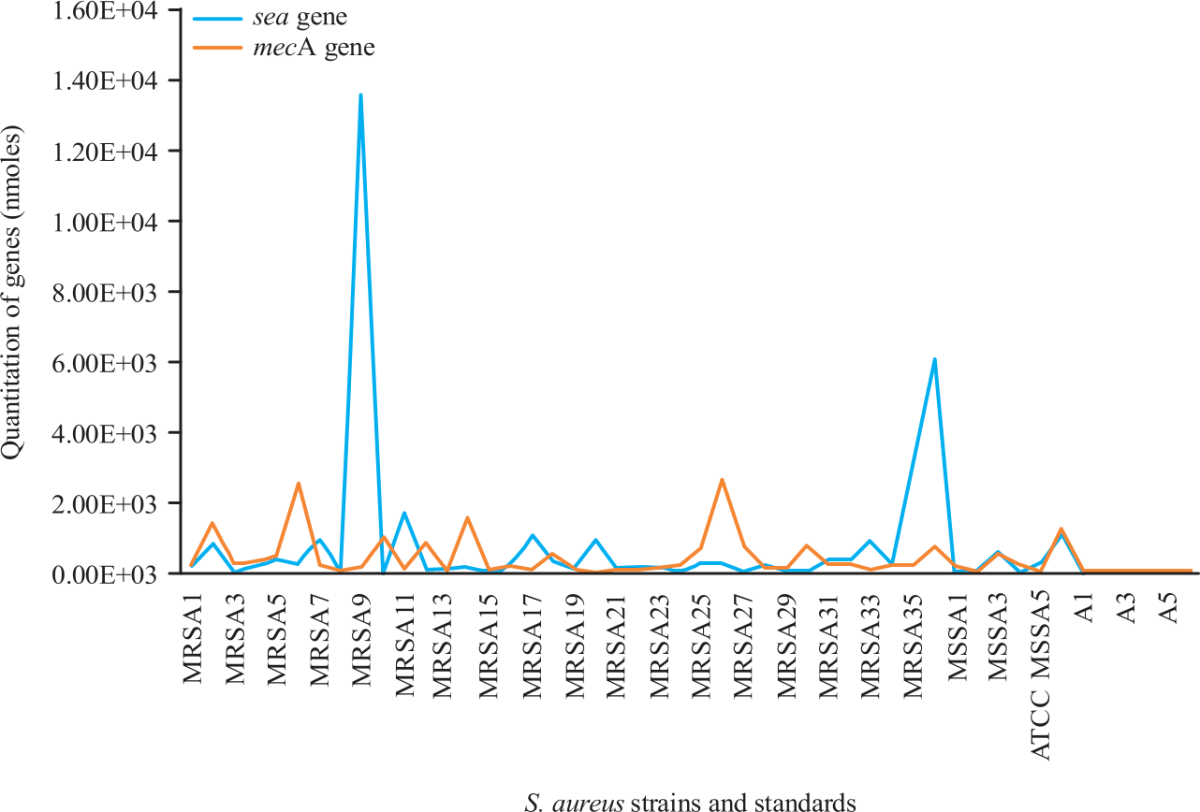
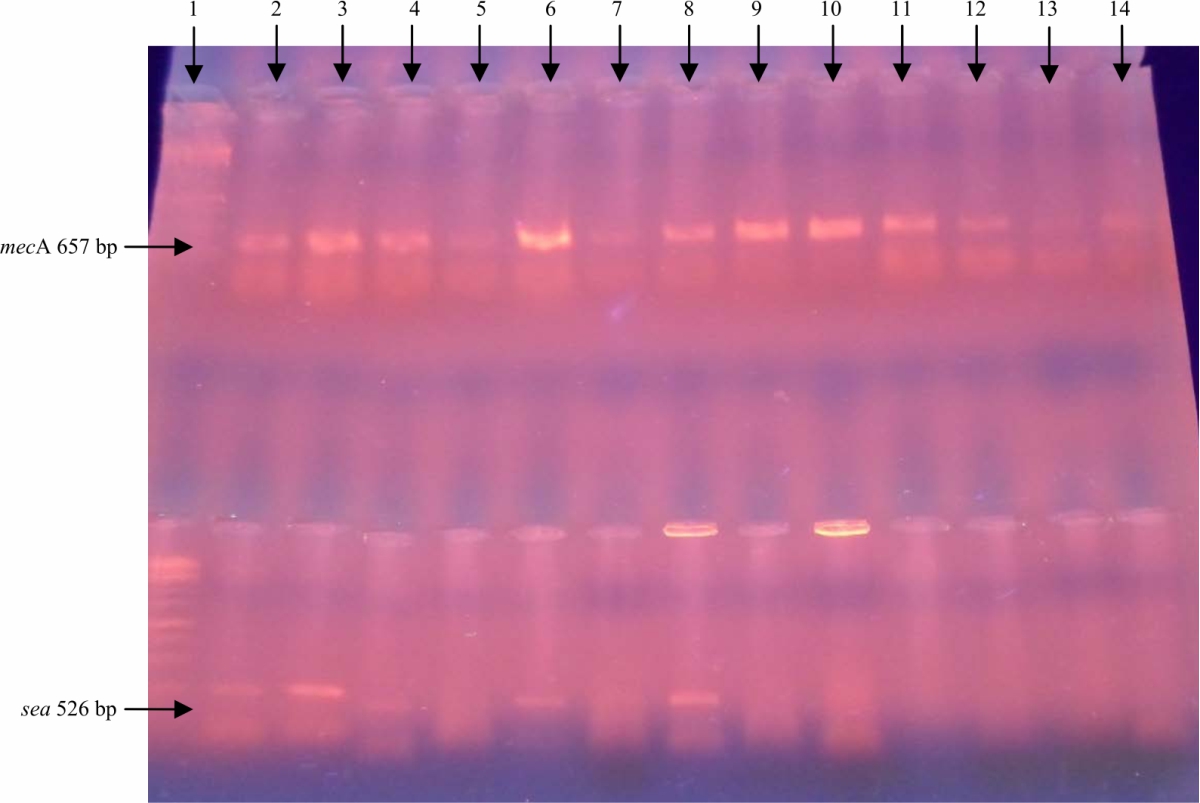
Quantitation of mecA and sea genes on Staphylococcus aureus using Quantitative PCR Assay
Department of Microbiology, Ambrose Alli University, Ekpoma, Edo State, Nigeria
LiveDNA: 234.18715
F.I. Esumeh
Department of Microbiology, Ambrose Alli University, Ekpoma, Edo State, Nigeria
G.T. Sunmonu
Department of Microbiology, Federal University Oye-Ekiti, P.M.B. 373, Oye-Ekiti, Ekiti State, Nigeria