Research Article
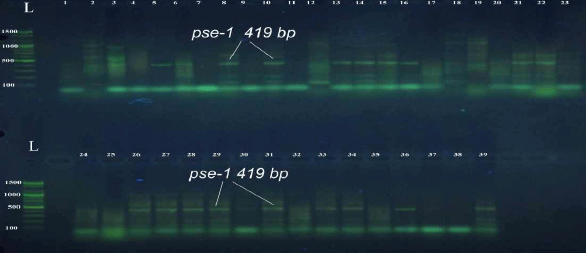
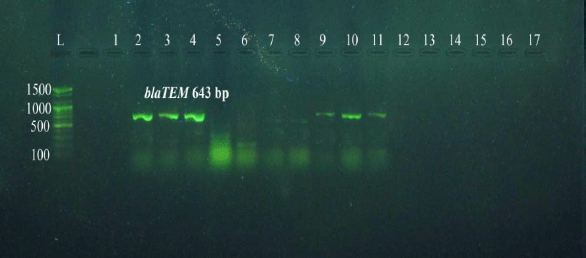
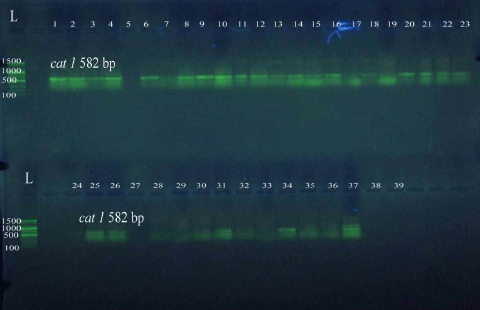
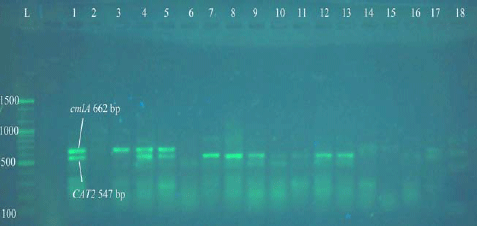
Prevalence of Some Antimicrobials Resistance Associated-genes in Salmonella typhi Isolated from Patients Infected with Typhoid Fever
Department of Microbiology, Faculty of Science, University of Kufa, Al-Najaf City, Iraq
LiveDNA: 964.12602
Ali Reyadh Medhat
Department of Microbiology, Faculty of Science, University of Kufa, Al-Najaf City, Iraq