Research Article
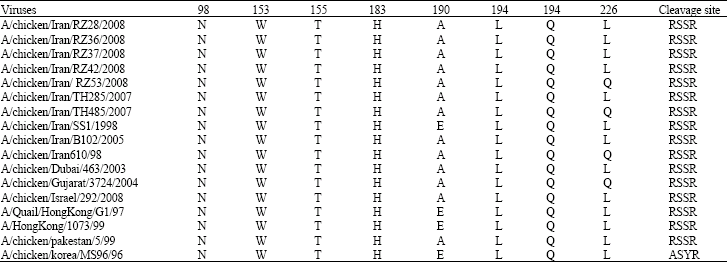
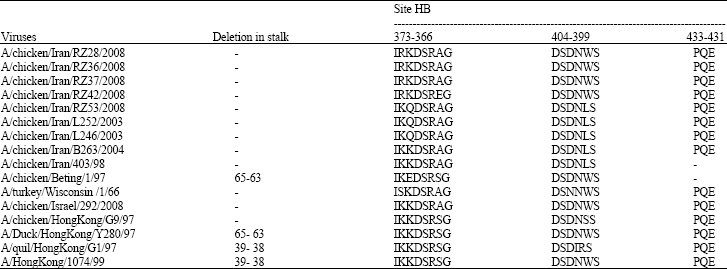
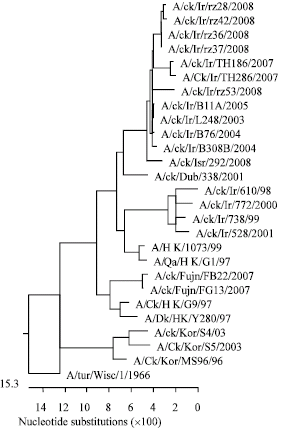
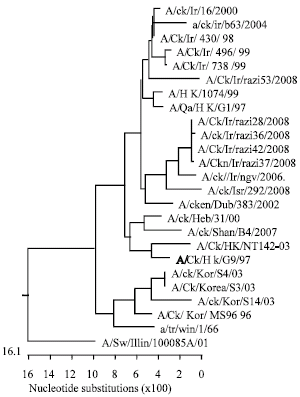
Molecular Characterization of Hemagglutinin and Neuraminidase Genes of H9N2 Avian Influenza Viruses Isolated From Commercial Broiler Chicken in Iran
Ph.D Student, Science and Research Branch, Isamic Azad University (IAU), Tehran, Iran
H. Shoushtari
Division of Avian Diseases and Research, Razi Vaccine and Serum Research Institute, Karaj, Tehran, Iran
M. Bozorgmehrifard
Division of Avian Diseases, Tehran University, Iran
S. Charkhkar
Division of Avian Diseases, Science and Research Branch, Isamic Azad University, Tehran, Iran
F. Eshratabadi
Division of Avian Diseases and Research, Razi Vaccine and Serum Research Institute, Karaj, Tehran, Iran