Research Article
Antibiotics Susceptibility Pattern and Virulence-associated Genes in Clinical and Environment Strains of Pseudomonas aeruginosa in Iraq
Department of Biology, Faculty of Science, University of Kufa, Kufa, Iraq
LiveDNA: 964.12602
Pseudomonas aeruginosa is one of the most important pathogenic bacteria, it is Gram negative bacteria, obligate, aerobic, oxidase and coagulase positive, capsulated, motile, non-lactose fermenter, cause different nosocomial infections and distribution in hospitals and different environments such as sewage water1,2. All clinical isolates of P. aeruginosa are multi-drug resistance and resistant of different classes of antibiotics such as beta-lactams, fluoroquinolones and amino glycosides3. Recently, clinical isolates of P. aeruginosa has become a serious clinical problem worldwide due to an increase resistant to 3rd and 4th generation cephalosporins4. Pseudomonas aeruginosa is considered one of the most virulent bacteria able to live in different environments include, hospitals, sewage water, soil and it is able to infect human and wide range of animals5,6. Hypermucoviscosity, biofilm formation, lipopolysaccharides formation, capsule synthesis, adhesions and iron uptake are the most important virulence factors in most pathogens7-9. Many virulence factors such as capsular polysaccharides, adhesion, siderophores and fimbriae are play an important role in pathogenicity of P. aeruginosa10. Detection of the prevalence of virulence genes in pathogenic bacteria and other pathogens by polymerase chain reaction technique is a good method to identify virulence factors in all pathogens such as Klebsiella pneumoniae (K. pneumoniae) and Escherichia coli (E. coli)11,12. These virulence factors allowing pathogenic bacteria to cause different illness such as pneumonia, burns infections and bacteremia13. Any microorganism especially pathogenic bacteria can be a virulent when a single factor presented, sometimes the presence of various factors at the same time is required to decide the bacterial ability of causing infections14. Some previous and recent studies outside Iraq suggested that there was positive correlation between the high presence of virulence genes and high resistance of antibiotics in pathogenic bacteria isolated from clinical and environment sources15-18. In Iraq, there was no studies focus on this relationship, therefore, the main aim of the present study was to compare between 60 clinical and 60 sewage water isolates of P. aeruginosa according to antibiotics susceptibility pattern and the prevalence of virulence genes in these isolates.
Samples and study design: This is a case control study performed in Department of Biology, Faculty of Science, University of Kufa, Iraq. A total of 120 isolates of P. aeruginosa were collected from two sources during the period from January-October, 2017; 60 isolates were collected from sputum of patients infected with pneumonia was considered as case study and 60 isolates were collected from sewage water was considered as a control study.
Sample collection and P. aeruginosa identification: Non-duplicate 60 sputum samples were collected in sterile containers (Himedia, India) from patients infected with pneumonia (diagnosis by specialized physician) admitted in Al-Najaf hospital in Al-Najaf City, Iraq (mouth should be rinsed out with distilled water, breath and cough deeply, coughed up from the lungs, not saliva or mucous, early in the morning upon rising) and homogenized with distilled water. Non-duplicate 60 samples of sewage water was collected in sterile containers (Himedia, India) from purified water station in Al-Najaf City, Iraq, during the period from January- October, 2017. Immediately, all collected samples were transferred to the Laboratory of Microbiology, Department of Biology, Faculty of Science, University of Kufa. All sputum and sewage water samples were streaked (by sterile swab, Bioanalyse, Turkey) onto blood agar (Oxoid, UK) surface, MacConkey agar (Oxoid, UK) surface and chocolate agar (Oxoid, UK) surface and incubated 24 h at 37°C. Any bacterial growth was identified by standard biochemical tests according to MacFaddin19 such as Gram positive, oxidase positive, coagulase positive, motility positive and growth on mannitol salt agar.
Antibiotics susceptibility testing: Disc diffusion method was used for antibiotics susceptibility test according to method by Kirby-Bauer20. Ten different antibiotics were used provided from Bioanalyse Company, Turkey as follows: Amoxicillin (25 μg), amoxiclav (30 μg), cefotaxime (30 μg), ceftazidime (30 μg), ceftriaxone(30 μg), amikacin (30 μg), tobramycin (10 μg), gentamicin (15 μg), tetracycline (30 UI) and imipenem (10 μg). Clinical and Laboratory Standards Institute21 was used as a guideline of antibiotic susceptibility and resistance according to bacterial zone diameter. According to the results of antibiotics sensitivity test, any bacterial isolate was resist to at least three different types of antibiotics classes was considered as a MDR, if was susceptible to only one type of antibiotic class was considered as a XDR and when resist to all types of antibiotics classes was considered as a PDR (CLSI21).
Phenotypic detection test of extended spectrum beta-lactamase: This test was done according to method by Aljanaby and Alhasnawi11 as follow:
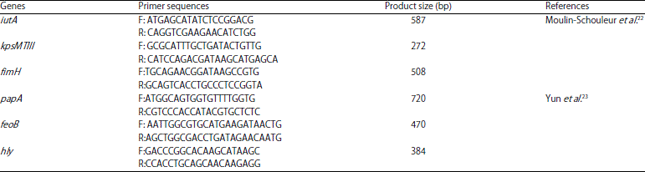
| Table 1: | All genes and primer sequences used in this study |
 | |
Amoxiclav disc (30 μg) was placed in the center of Mueller Hinton agar plate (Oxoid, UK). Ceftriaxone disc (30 μg), cefotaxime disc (30 μg) and ceftazidime disc (30 μg) were placed with distance of 15 mm from center to center of amoxiclav disc and incubated at 37°C overnight. If inhibition zone was increased towards the amoxiclav disc then considered as positive result.
DNA extraction, primer sequences and thermo cycling conditions: This method was done according to procedure by Aljanaby and Alhasnawi11. Briefly, all P. aeruginosa isolates were heated in boiling water at 100°C for 30 min and centrifugation at 8000 rpm for 10 min, the supernatant was considered as a DNA template. The primer sequences are mentioned in Table 1. The multiplex PCR program consisted of an initial denaturation step at 95°C for 4 min, followed by 35 cycles of DNA denaturation at 52°C for 60 sec, primer annealing at 55°C for 60 sec extension 72°C for 60 sec. After the last cycle, a final extension step at 72°C for 5 min. Five microliter aliquots of PCR product were analyzed by gel electrophoresis with 2% agarose22,23.
Statistical analysis: GraphPad prism computer software version 8 was used in this study for comparison between samples according to Chi-squared test. The p<0.05 was considered statistically significant.
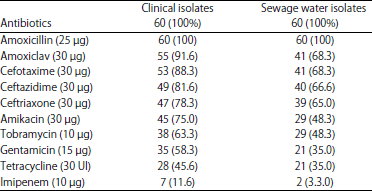
Antibiotics susceptibility: All clinical isolates 100% (60/60) were resistance to amoxicillin, that were resistant to other antibiotics with different percentages as follow: Amoxiclav 91.6% (55/60), cefotaxime 88.3% (53/60), ceftazidime 81.6% (49/60), ceftriaxone 78.3% (47/60), amikacin 75% (45/60), tobramycin 63.3% (38/60), gentamicin 58.3% (35/60), tetracycline 45.6% (28/60) and imipenem 11.6% (7/60).
| Table 2: | Numbers and percentages of P. aeruginosa strains isolated from clinical source and sewage water that were resistant to antibiotics |
 | |
| Table 3: | Numbers and percentages of MDR, XDR and PDR of P. aeruginosa strains isolated from clinical source and sewage water |
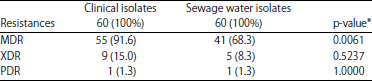 | |
Compare between clinical isolates and sewage water isolates, MDR: Multi-drug resistance, XDR: Extensive-drug resistance, PDR: Pan-drug resistance | |
Also, all sewage water isolates 100% (60/60) were resistance to amoxicillin, while, was resistant to amoxiclav and cefotaxime with percentage 68.3% (41/60), ceftazidime 66.6% (40/60), ceftriaxone 65% (39/60), amikacin and tobramycin 48.3% (29/60), gentamicin and tetracycline 35% (21/60) and imipenem3.3% (2/60) (Table 2). Out of 60 clinical isolates, there were 55 isolates (91.6%) were MDR, 9 isolates (15%) were XDR and 1 isolate (1.3%) was PDR. Out of 60 sewage water isolates, there were 41 isolates (68.3%) were MDR, 5 isolates (8.3%) were XDR and 1 isolate (1.3%) was PDR. The results proved that there was significant differences (p-value = 0.0061) in the prevalence of MDR of clinical isolates as compared with sewage water isolates (Table 3). While, there was no significant differences (p-value = 0.3282) between clinical isolates (18/60, 30%) and sewage water isolates (11/60, 18.4%) in the prevalence of ESBL-producing isolates (Table 4).
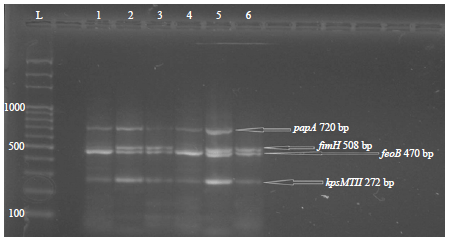 | |
| Fig. 1: | Gel-electrophoresis of PCR amplification of P. aeruginosa DNA using papA, fimH, feoB and kpsMTII genes specific primers show positive results at (720, 508, 470 and 272) bp, respectively |
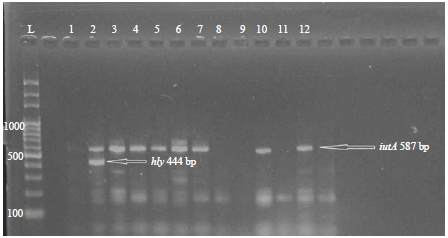 | |
| Fig. 2: | Gel-electrophoresis of PCR amplification of P. aeruginosa DNA using iutA and hly genes specific primers show positive results at (587 and 444) bp, respectively |
| Table 4: | Numbers and percentages of ESBL-producing P. aeruginosa strains isolated from clinical source and sewage water |
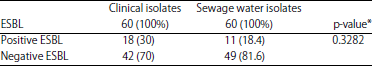 | |
Compare between sputum and sewage water isolates, ESBL: Extended spectrum beta-lactamase | |
| Table 5: | Numbers and percentages of prevalence of virulence genes in P. aeruginosa strains isolated from clinical source and sewage water |
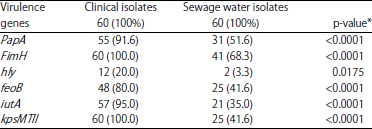 | |
Compare between clinical isolates and sewage water isolates | |
Genotypic detection of virulence genes: The results proved that there were significant increase (p<0.05) in the prevalence of virulence genes in clinical isolates as compared with sewage water isolates (Table 5). The PapA, FimH, feoB, iutA, hly and kpsMTII genes were prevalence in clinical isolates with percentages 91.6, 100, 80, 95, 20 and 100%, respectively, while the same genes were prevalence in sewage water isolates with percentages, 51.6, 68.3, 41.6, 35, 3.3 and 41.6%, respectively (Fig. 1 and 2).
This study showed that the clinical and sewage water isolates of P. aeruginosa were MDR with percentages 91.6 and 68.3%, respectively with significant increase (p = 0.0061) in the prevalence of clinical isolates as compared with sewage water isolates. Multi-drug resistance pathogens such as P. aeruginosa, K. pneumoniae and S. aureus become one of the most important medical problems worldwide11. Pseudomonas aeruginosa is one of the most important multi-drug resistance bacteria found in hospitals environments, water, soil and cause different kinds of infections include pneumonia and liver abscess24. Mostly encountered MDR bacterial strains such as 3rd and 4th generation cephalosporins resistance P. aeruginosa is common and prevalent in a patients infected with pneumonia and water environments25,26. Water and air borne pathogens such as P. aeruginosa and E. coli could cause different kinds of illness such as pneumonia and heavy diarrhea27,28. Multi-drug resistance P. aeruginosa is inherently resistant to different drug classes and is able to acquired resistance to all effective antibiotics29 and elaborates inactivating enzymes that make β-lactams ineffective such as extended spectrum β-lactamases30. The antibiotics susceptibility pattern of sewage water isolates of P. aeruginosa is mostly overlooked and rarely studied. One of the most important of antibiotics resistance strategies in P. aeruginosa is efflux pumps, this resistance mechanism has been increasingly recognized as a major component of resistance in P. aeruginosa31. Eighteen clinical isolates (30%) and 11 sewage water isolates (18.4%) were ESBLs-producing strains according to phenotypic test in this study. Extended spectrum β-lactamases (ESBLs)-producing P. aeruginosa have been associated with different infections such as pneumonia, liver abscess and burns infections and became a serious health problem in worldwide lead to treatment failure in hospitals32. Extended spectrum β-lactamases are plasmid mediated enzymes that hydrolyze broad spectrum β-lactams antimicrobials. These ESBLs are transmitted by plasmids between bacteria to bacteria in different environments and hospitals lead to emergence a new ESBLs-producing bacteria11. If infections were accrued caused by ESBLs-producing bacteria, can be predicated by the clinical characteristics of patients, can lead to a batter selection of antimicrobials and help to improve the outcome of infections33. The efflux system in P. aeruginosa is mostly encoded by chromosomal genes and capable of ejecting of antibacterial agents and fall into five families, multi-drug and toxic compound extrusion, ATP binding cassette, major facilitator superfamily, small multi drug resistance and the resistance nodulation division34. Because the efflux system is encoded by chromosomal genes and these genes are able to transfer from bacteria to other in hospitals environment and between person to person, therefore, in this study, clinical isolates of P. aeruginosa were high MDR isolates as compare with sewage water isolates.
Infections caused by P. aeruginosa are well known to be persistent and resilient in hospitals and environments due to the tendency of the species to form biofilms35. Biofilms and outer membrane lipoproteins are play an important role in protecting Gram negative bacteria from different antimicrobials effect36. In this study, all 60 clinical isolates (100%) were harbored fimH gene, 55 isolates (91.6%) were harbored PapA gene and 12 isolates (20%) were harbored hly gene. While sewage water isolated were harbored the same genes in low prevalent, 41 isolates (68.3%), 31 isolates (51.6%) and 2 isolates (3.3%), respectively. Many studies suggested that outer membrane lipoproteins and biofilms act as biodegradable effect on β-lactamase antibiotics. Beta-lactamases enzymes are secreted and maintain their activity inside of biofilm matrix and decompose β-lactam antibiotics before these antibiotics reach the bacterial cells7-37. In the present study, all 60 clinical isolates (100%) were harbored kpsMTII gene that responsible for capsule synthesis while only 25 isolates (41.6%) were positive for this gene. Almost P. aeruginosa produce large amounts of extra-capsular polysaccharides and mucopolysaccharide mass to produce strain with more virulent38. The kpsMTII is important gene have an essential role in capsule synthesis and pathogenicity of P. aeruginosa39,40. Pseudomonas aeruginosa is able of producing and secreting siderophores for iron uptake, siderophores are weakly soluble and found in most environments such as sewage water. In this study, feoB gene was found in 80 and 41.6% of clinical isolates and sewage water isolates, respectively. While, iutA gene was predominated in 95 and 35% of clinical isolates and sewage water isolates, respectively. This results are in agreement with previous studies proved that the most mucoidgram-negative bacteria isolated from clinical isolates were harbored virulence gene responsible for siderophores synthesis7-41. Siderophore are compounds produced by bacterial cells to iron uptake from iron-binding proteins of the host and they have strong-affinity extracellular ferric chelators, which, they have an important role in pathogenesis and virulence of bacteria, virulence of these compounds are remain unclear42. Most Gram negative bacteria such as Pseudomonas aeruginosa are able to produce many types of toxins, one of the most important toxins is hemolysins. These toxins are proteins and lipids that cause lysis of red blood cells by destroying their cell membrane43. In the present study, 12 clinical isolates (20%) and 2 sewage water isolates (3.3%) were harbored hly gene. Hemolysins are able to enhance the production of outer membrane vesicles and release of the cytolysin44. In Gram negative bacteria, outer membrane turnover and outer membrane vesicles production provide a general mechanism of protection against clinical and environment source of envelope stress like antimicrobials and agents45,46.
Clinical strains of P. aeruginosa isolated from sputum of patients infected with pneumonia were harbored virulence genes and were high antibiotics resistance more than those isolated from sewage water. Therefore, clinical strains of P. aeruginosa are more virulent than those isolated from environment isolates. It is recommended to analyze the antibiotic susceptibility pattern and virulence-associated genes of other pathogens such as K. pneumonia and S. aureus and conduct the comparison to get more pronounced data on virulence genes.
This study discovered that the environment isolates of P. aeruginosa has new virulence-associated genes don’t found before. So, this study could be beneficial in determining the detail mechanism of virulence gene and formulation of antibiotics to treat the diseases related to it. This study will help researchers to discover more details about virulence genes in clinical and environments isolates of P. aeruginosa and also to modify the antibiotics according to the virulence genes discovered to treat the disease. Thus, best theory on it may be arrived at.