Research Article
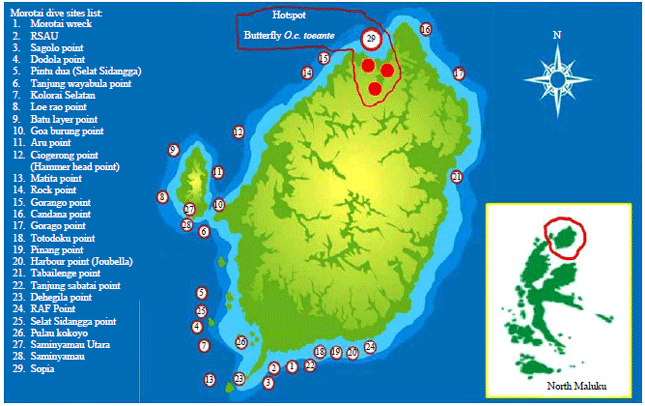
Genetic Variability of Ornithoptera croesus toeantei Endemic Butterfly in Morotai Island, Based on Morphology and Molecular
Department of Biology Education, Faculty of Teacher Training and Education, Universitas Khairun, Street Pertamina Kampus II Unkhair Gambesi, 97728 South Ternate, North Maluku, Indonesia
LiveDNA: 62.29963
Sundari
Department of Biology Education, Faculty of Teacher Training and Education, Universitas Khairun, Street Pertamina Kampus II Unkhair Gambesi, 97728 South Ternate, North Maluku, Indonesia
Nurhasanah
Department of Biology Education, Faculty of Teacher Training and Education, Universitas Khairun, Street Pertamina Kampus II Unkhair Gambesi, 97728 South Ternate, North Maluku, Indonesia
Chumidach Roini
Department of Biology Education, Faculty of Teacher Training and Education, Universitas Khairun, Street Pertamina Kampus II Unkhair Gambesi, 97728 South Ternate, North Maluku, Indonesia
Mohamad Amin
Faculty of Mathematics and Natural Sciences, State University of Malang, Street Semarang 5, Sumbersari, Lowokwaru District, 65145 Malang, East Java, Indonesia
Fatchur Rohman
Faculty of Mathematics and Natural Sciences, State University of Malang, Street Semarang 5, Sumbersari, Lowokwaru District, 65145 Malang, East Java, Indonesia
Alisi
Conservationists, Endemic Butterflies From Bacan, 97728 North Maluku, Indonesia