ABSTRACT
In recent years, the research on the Y chromosome of Drosophila melanogaster has made great progress. The Y-linked, testis-expressed Su (Ste) repeats are thought to be suppressors of Stellate transcription. New in this study is the recent advance on studies of the structure and functions of Su (Ste) repeat units on Y chromosome; We expressed the hypothesis about the cosuppression mechanism of Su (ste) and Stellate: Su (ste) antisense RNA combined with Stellate RNA to form double strands RNA (dsRNAs), which restrains Stellate expression; Su (ste) sense-chain RNA reacts on Su (ste) anti-sense RNA, to form dsRNA that restrains Su (ste)’ transcription, which called negative autogenously regulation, now we call it RNAi; The mechanism of the Su(Ste) repeats’ suppression on meiotic-driving is also discussed: In the spermatocyte of male Drosophila which delete the Su (ste), Stellate mRNA and Stellate protein were highly expressed, which caused the crystal formation and the change of meiosis, then resulted in the male sterility. The mechanism of how the expression of Stellate protein causing the abnormal of meiotic is possibly because the restrain of gene CK2 activity. The molecular evolution of the Su(ste) repeats has been reviewed in this paper too, the unique testis-expressed euchromatic gene was suggested to be an ancestor of both types of amplified paralogous repeats, SLL originates from CK2 through reverse-transcription, Su(Ste) and Stellate at the same time originate from SLL, then the ancestor sequence translocated to Y- chromosome.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/je.2005.46.51
URL: https://scialert.net/abstract/?doi=je.2005.46.51
INTRODUCTION
Fruit fly is a favorite research tool for genetic study since the beginning of this century. Because It has only 4 chromosomes and breed very quickly, the female produces hundreds of eggs at a time and a new generation come along every two weeks. Studying about it not only influence the fruit production, but also bring many discoveries in Genetics, A.H. Sturtevant (1913) suggested that genes were arranged in linear order on chromosome. TH Morgan used fruit Fly to prove it successfully, and found the linkage inheritance, crossing-over and recombination.
But the Queensland fruit fly, Bactrocera tryoni, has a diploid of 12 chromosomes, five pairs of autosomes and a XX/XY sex chromosome pair[1]. Queenland fruit fly caused millions of dollars damage to Australia horticulture every year, it infests almost every commercial vegetable and fruit crop[2]. To minimize the use of chemical insecticide, biological control is necessary and considerable effort has been done. The fruit fly research center in the university of Sydney has developed a preliminary Sterile Insect Techniques (SIT). The challenge is to develop genetic sexing separation system with genetic and molecular techniques, which allow rising of the male-only broods. Some visible genetic markers in Q-fly has been isolated, gene mapping in the Q-fly, Bacterocera tryoni has been explored for several years. The B. Tryoni Y-autosome translocations using a recessive marker have been constructed and such translocations can be utilized to construct genetic sexing strains[1,3].
The most acceptable theory about the sex determination nowadays is that the sex chromosomes determine the sex of fruit fly. Female has homologous sex chromosome, two X chromosomes; male has heterogeneous sex chromosomes, one X chromosome and one Y chromosome. But according to recent study, there are some genes involved in the sex determination, it is the genes not the entire sex chromosome that determines fruit fly’s sex. More and more evidences can be found now to prove this gene determination theory. In the male Drosophila melanogaster, Y chromosome takes part of the 12% of entire gene group. Scientists try to research on the heredity of Y chromosome for a long time and have gotten many important accomplishments, especially have gotten significant progress in recent years on molecular level. Livak first discovered that the 2L1 sequence of the kl-1 and kl-2 districts on Y-chromosome’s long arm is related to the Ste position of X chromosome[4]. Repetitive unit Su (Ste) [suppressor of stellate] is located in Crystal seats of 2L1 sequence. The X-linked Stellate repeats, encoding a putative regulatory subunit of protein kinase CK2, are expressed in XO male testis. In the early stage of the spermatogenesis, if Su (Ste) fragment is lost, the sperm of Drosophilae will be abnormal and unusually cause its male sterility. It was in 1992 that Balakireva has cloned complete Su(Ste) repeat sequence[5]. In the following 10 years, research on the structure, functions and evolution of Su (Ste) repeat has got some important accomplishments.
The structure and functions of Su (Ste) repeats: The Su (Ste) repeat is a X –linked Stellate multiple-copy gene, which only expressed in the testis of male XO type Drosophila, it codes a kind of polypeptide highly homologous with the β subunit of protein kinase CK2 and Stellate protein is tested in vitro, which showed it could interact with protein kinase CK2β and it can control the action of CK2β[6]. The multicopies of Su (Ste) on Y chromosome are considered as Stellate expression inhibitor, its position is the Crystal seat of Y chromosome. In early stage of spermatogenesis, the deletion of Su(Ste) caused the Stellate gene over-expression, which resulted in the meiotic division abnormal; the abnormal meiotic division caused the accumulation of crystalline and the crystalline accumulation then caused the male sterility[7]. The Su (Ste) repeat sequence is composed of 3 areas (Fig. 1): part one has 90% nucleotides similar with Stellate and include the promoter sequence and coding sequence; Part 2 is a Y chromosome specific sequence with no relation to Stellate; part 3 is a 1360bp inserted transposition element[5,8]. By the comparison of the stellate-like sequence between the multicopies of Su(Ste), it is discovered that the stellate-like coding district has damaged ORF, which means the selection pressure is very low, it does not code protein, there is no PolyA sequence in Su(Ste) which exists in Stellate, but the PolyA signal replaced by the Y-specific sequence[9].
In 1991, Danilevskaya[10] based on the homologues analysis of the Y-linked Su (Ste) and Stellate sequences, has made an assumption that the interference of antisense chain of Su (Ste) RNA silents the Stellate gene’s expression, so as to maintain the reproduction ability of male fruit fly. However, Kalmykova et al.[11,12] has not detected the antisense chain of Su (Ste) RNA by RT-PCR technique, but obviously detected the existence of sense chain of RNA. Accordingly to this, the transcribed sense chain of RNA of Su (Ste) repeats can code polypeptide, it has conservative zinc-finger structural region and through DNA-Protein interaction it can restrain the transcription of Stellate. Newest research shows that sense chain of RNA, antisense chain of RNA and dsRNA of Su (Ste) can be detected at the same time in Drosophila melanogaster. Since antisense chain of RNA without Oligo dTs cannot be detected by RT-PCR with Oligo dT primer. This has explained the result of Kalmykova who did not detect the antisense chain of RNA. Accordingly to this result and the development of RNAi technique, we can certify the assumption of Stellate/Su (ste) cosuppression (Fig. 2).
Su (ste) transcriptional antisense chain of RNA combined with Stellate RNA to form double strands RNA (dsRNAs), which restrains Stellate expression; Sense chain of Su (ste)’s transcriptional product (RNA) reacts on antisense chain of Su (ste)’s transcriptional product (RNA), to form dsRNA that restrains Su (ste)’ transcription, which called negative autogenously regulation. Now we call it RNAi. In fruit fly, dsRNA can be cut by nuclease DICER to become the small RNA fragments of 21-23 bp[13]. In normal testis of Su (ste)+/ Stellate+ Drosophila melanogaster, the little RNA fragment of 25-27 bp can be detected. This has proved that dsRNA can be degraded by special enzyme. The mechanism of silencing of testis expressed X-linked Stellate repeats by homologous Y-linked Suppressor of Stellate [Su(Ste)] repeats localized in the crystal locus[14]. The dsRNA and siRNA (small interference RNA) as a product of symmetrical transcription of Su (Ste) repeat were revealed the mechanism of Stellate silencing by RNA interference (RNAi). The RNAi mechanism of Stellate silencing as a result of the transcription of Stellate and Su (Ste) repeat sequences is shown in Fig. 2. The RNAi phenomenon was first discovered in nematode (Caenorhabditis elegans)[15] then cosuppression was discovered in plant’s gene silencing[16] which also gets enough discussion in transgenic fruit fly[17]. At present, the phenomenon of RNAi has been found in many kinds of animals and plants, but the interaction of Stellate and Su (ste) repeat may represent the first discovered natural cosuppression control mechanism of gene expression.
The process of Stellate silenced by Su (ste) repeat, is normally happened after transcription. But also cannot exclude the silencing of Stellate by Su (ste) in the transcription course.
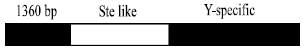 | |
| Fig. 1: | The structure of Su (Ste) unit[18] |
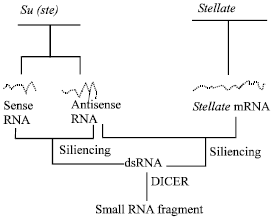 | |
| Fig. 2: | The cosuppression [RNAi] mechanism of Su(ste) and Stellate |
Because Su (ste)’s promoter probably act on Stellate’s DNA, hinder the transcription of Stellate, this phenomenon has been discovered in plant recently[18]. So whether Stellate/Su (ste) has this possibility is worth for further research.
Su (ste)/Stellate and meiotic drive of chromosome: Meiotic drive is taking place in male’s testis to produce sperms without Y chromosome; it causes to breed excessive female descendants. The mechanism of this phenomenon is not understood now; it usually supposed that it originated from the change of driving position and linkage decoration, or the direct result caused by the chromosome’s feature[19]. The huge heterochromatin district on Y chromosome has been considered as to prevent the pairing and the recombination of X chromosome and Y chromosome. The behavior of Y chromosome is the foundation of sex ratio. Abnormal behavior of Y chromosome is the signal of meiotic-drive, which causes the failure of producing differential gametes[20]. McKee et al.[21] has studied the relation of rDNA’s IGS district and the meiotic-drive and also concluded that the aberration of Y chromosome is possibly the cause of meiotic-drive.
In the spermatocyte of male Drosophila melanogaster that deleted the Su (ste), Stellate mRNA and Stellate protein were highly expressed. The highly expression of Stellate caused the crystal formation of Stellate protein; crystal formation caused the change of meiosis which result in male sterility of Drosophila melanogaster. The absence of crystal is controlled by Su (ste), its shape is decided by the copy number of stellate[4]. Now the mechanism of how the expression of Stellate protein caused the abnormal of meiotic is not known, possibly because the restrain of gene CK2β activity. Hurst assumed that Stellate is the meiotic drive inhibitor[22,23]. Recently in Drosophila melanogaster the additional 3 genes sting, scratch and homeless have been discovered[24], they can restrain the expression of Stellate, their mutations also can raise the occurrence of meiotic-driving, but the mechanism is not very clear. Seemingly, meiotic drive is a kind of very complex chromosome hereditary behavior, because there is many proteins’ participation. It can infer that the interaction of Su (ste) and Stellate in Drosophila melanogaster has played a key role in the meiotic-drive.
Homeless mutations were previously found to cause female sterility with numerous defects in oogenesis, including improper formation of both the anterior-posterior and dorsal-ventral axes and failure to transport and localize key RNAs required for axis formation. One homeless mutation was also found to affect male meiosis, causing X-Y chromosome pair’s nondisjunction. Stapleton W reported that these mutations cause a variety of defects in male meiosis, including: nondisjunction of the X-Y chromosome pair, nondisjunction of the chromosome 2 pair, Y chromosome marker loss, chromosome breakage, chromatin bridges at anaphase and tripolar meiosis. In addition, homeless mutations interact with an X chromosome can cause complete male sterility. These phenotypes are similar to those caused by deletion of the Suppressor of Stellate [Su(Ste)] locus. Like Su(Ste) deficiencies, homeless mutants also exhibit crystals in primary spermatocytes and derepression of the X-linked Stellate locus. Their functions along with Su(Ste) and other recently described genes to repress the Stellate locus in spermatocytes and that it may also play a role in repressing certain other repeated sequences[24].
In the spermatocyte cell of male fruit fly which delete Su (Ste) repeat sequence, Stellate protein highly expressed, cause the change of meiotic division and the male sterility of fruit fly. In this course, the mechanism of Su (Ste) and Stellate worth further research, especially whether Su (Ste) is expressed and whether it’s RNA directly act on Stellate DNA. How the over expression of Stellate causes the change of meiotic division and their relation to chromosome meiotic division are still in research. If these problems have been solved, we probably know the mechanism of meiosis and meiotic drive and the function of heterochromatin on Y chromosome, etc. This can help us to understand the reproduction principle and to solve the reproduction problem, then breed only male offspring and control the population of fruit fly, reduce their damage to agriculture.
The Evolution of Su (ste) repeats: Kalmykova has cloned a gene-SLL(suppressor-of-Stellate-like) on autosome which is highly homologous with CK2β, Su(Ste) and Stellate[11]. SLL protein has 45% similarity with CK2β, the similarity with Stellate is 53%. Molecular pedigree analysis shows (Fig. 3): SLL originated from CK2β through reverse transcription; Su(Ste) and Stellate originated from SLL at the same time, the 3’ of non-transcription ancestor’s DNA sequence translocated to Y-chromosome, but no ancestor’s sequence translocated to X chromosome, so the copies on Y chromosome increased. Kogan found a Su (Ste) Orphan sequence on the Y-chromosome[8], it has Su (Ste) and Stellate sequence feature at the same time, it can be inferred that the Su (Ste) on Y chromosome is an evolution record of ancestor’s sequence. A Su (Ste)-like orphan was localized on a Y chromosome, outside of the Su (Ste) cluster. Several diagnostic molecular markers peculiar for the both types of diverged Stellate and Su (Ste) units were detected in the orphan sequence. The orphan was suggested to be a close relative of the immediate ancestor of both types of paralogous repeats which initiated evolution on the Y chromosome. Selection pressure on the level of translation was shown as a driving force in the evolution of Su (Ste) repeats, which are considered as more ancient derivatives of the ancestor euchromatic gene than Stellate repeats. In a vicinity of 12E Stellate cluster, the undamaged recently originated euchromatic Stellate orphan was found at 12D, providing the poly (A) signal for the bendless gene. P-element mediated transformations revealed that the fragments of cloned Stellate and Su (Ste) clusters are able to induce variation of a reporter mini-white gene. The observed variation phenomenon has peculiar features: a significant increase of trans-activation of a reporter mini-white gene in homozygous state; absence of effects of several conventional modifiers of position effect variation (PEV) and independence of a severity of variation on a distance between insertion and centromere region[25]. Su (Ste) may get certain translation selective pressure, which make the copy number keep homogenous, but there are 6.4% replacing difference between repetitive fragments, the homogenization degree of Stellate heterochromatin on X chromosome is 5 to 10 times higher than Su (Ste) repetitive fragment on Y chromosome, but they have only 2.5% nucleotides replacing difference[8].
In the past 10 years, scientists attempt to detect the molecular effect of Su (Ste) on sexual selection, which showed that genes relating to male reproduction were evolving very quickly among species[26,27].
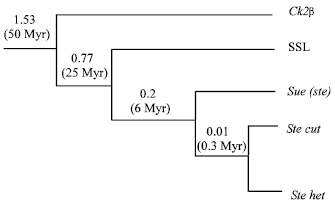 | |
| Fig. 3: | Gene genealogy for the CK2β, SLL, Su(Ste) and Stellate of D. melanogaster |
According to the comparative study of Su (Ste) multiple sequences, we discovered that Stellate-like coding area has damaged ORF and have certain difference between sequences, which showed that Su (Ste) has very weak selective pressure and the evolution is very rapid, because Su (Ste) is related with male reproduction. According to the research of other Y-chromosome genes, the coding regions of Y-chromosome are under great selective pressure, but they are very conservative, such as Dhc-Yh[28], which lack nucleotides polymorphism. But group hybridization research shows, Y chromosome has very high polymorphism, this is coincident with Haldane's rule[29]. We can infer from above that maybe the functional non-coding regions on the Y chromosome have higher polymorphism, as IGS, its coding region has very strong conservation. This also prove Su (Ste) repeat sequence may be a non-coding sequence.
Molecular evolution and divergence of paralogous tandem heterochromatic repeats Stellate and Su (Ste) located on the X and Y-chromosomes respectively. These repeats appear to emerge as a result of amplification of a unique autosomal euchromatic gene encoding the regulatory β-subunit of the CK2 protein kinase. The autosomal gene and the clusters of heterochromatic repeats are transcribed in testes. A high level of the Stellate expression leads to partial male sterility and abnormal meiosis[30]. The Stellate expression and its adverse effects are suppressed by homologous Su (Ste) repeats. In genome evolution, the Open Reading Frames (ORF) of Stellate and Su (Ste) putative ancestor were maintained by translation selection. Then the coding Su (Ste) function has been damaged and switched to a new one resulting in antisense Su (Ste) transcription and silencing of the Stellate genes. Symmetrical (sense and antisense) Su (Ste) transcription causes the formation of a double-strand RNA, which by RNA interference (selective elimination of the homologous gene expression) suppresses the Stellate genes[31]. Now only on the Y chromosome of the D. melanogaster, D. simulans and D. mauritiana have determined the existence of Su (ste), not detected the stellate-like sequence in the D. erecta, D. tessieri and D. yakuba of the same genus of Drosophila melanogaster. Hurst inferred that the Stellate in Drosophila melanogaster comes from the invading by external group and then Su (Ste) repeats invaded in.
It can be concluded that the study of the structure, function, distribution, formation and molecular evolution of Su (Ste) and Stellate repeats in Drosophila melanogaster is very meaningful and can be used in the controlling of the population of fruit fly to do good to our agriculture.
ACKNOWLEDGMENTS
The authors would like to thank Yu H, Wangyan, Xoumei, Frommer M, Robson MK, Meats AW, Shearman DC, Sved JA of Fruit Fly Research Center, School of Biological Sciences, University of Sydney, for their helpful discussions and guidance for us; Thank the University of Sydney for giving me OPRS and IPA scholarships to support my study; We are grateful to our families, colleagues and many friends for their support of my work.
REFERENCES
- Bozzetti, M.P., S. Massari and P. Finelli, 1995. The Ste locus, a component of the parasitic cry-Ste system of Drosophila melanogaster, encodes a protein that forms crystals in primary spermatocytes and mimics properties of the beta subunit of casein kinase 2. Proc. Natl. Acad. Sci. USA., 92: 6067-6071.
- Tang, G.L., B.J. Reinhart, D.P. Bartel and P.D. Zamore, 2003. A biochemical framework for RNA silencing in plants. Genes Dev., 17: 49-63.
Direct Link - Bass, B.L., 2000. Double-stranded RNA as a template for gene silencing. Cell, 101: 235-238.
CrossRefDirect Link








