Research Article
Classification of Halophilic Heterotrophic Bacteria Thriving in the Jordanian Dead Sea Littoral Zone
Department of Biological Sciences, Faculty of Sciences, Al al-Bayt University, Mafraq, Jordan
Halophiles are interesting group of organisms thriving at high salinity. These organisms can be broadly classified based on their optimum salinity for growth into mild halophiles (1-6%, w/v NaCl), moderate halophiles (7-15%) and extreme halophiles (15-30%) (Madigan and Martinko, 2006). Extensive research has shown that halophiles are not restricted to any of the life domains, they can be eukaryotes (Gunde-Cimerman et al., 2000; Zalar et al., 2005), or prokaryotes belonging to the domains of Bacteria and Archaea. The algal genus Dunaliella and the brine shrimp Artemia (Boetius and Joye, 2009) are examples of halophilic eukaryotes, whereas Halobacterium and Salinibacter are examples of halophilic prokaryotes. Success of halophiles in surviving in highly salty environments is due to unique physiological adaptations, like ion pumping strategy and organic solutes accumulation (Oren, 2006; Madigan and Martinko, 2006). At the genomic and proteomic levels, halophiles are characterized by high GC content and proteins characterized by low hydrophobicity, over-representation of acidic residues, lower propensities of helix formation and higher propensities of coil structure (Paul et al., 2008).
Halophiles are now gaining more access to industrial microbiology and biotechnology because halophiles grow at high salt concentration and this minimizes the risk of contamination during cultivation (Oren, 2006). Few examples of biotechnological applications are the use of Micrococcus varians to produce nuclease H (Kamekura el al., 1982) and the use of the halophilic Tetragenococcus strains in the production of soy sauce and the production of some enzymes including hydrolases (amylases, nucleases, phosphatases and proteases) (Oren, 2006). Halophiles are also important in biodegradation and bioremediation since many halophiles are able to degrade hydrocarbons and other toxic compounds (Ventosa et al., 1998). Halophiles can also produce polymers used as enhancers of oil recovery because of their surfactant activity and bioemulsifying properties (Oren, 2006).
Halophiles flourish in environments where salinity reaches high levels, such as oceans, solar salterns and natural salt lakes (Oren, 2007). The Dead sea of Jordan is one of largest truly hypersaline inland salt lakes in the world (Boetius and Joye, 2009; Oren, 2007; Madigan and Martinko, 2006). In addition to high salinity; salt, concentration of over 340 g L-1 (Oren, 2007), the Dead sea is unique by its high barometric pressure (800 mmHg) due to very low altitude below sea level, partial oxygen pressure (PIO2) of 8% more than at sea level, unique UV radiation, low humidity (below 40%) and paucity of rain (Avriel et al., 2011).
This study was conducted to classify the halophilic heterotrophic bacterial species thriving in the littoral zone of the Dead sea of Jordan. Microbial classification is based on colonial and cell morphology, as well as similarity in 16S rRNA gene.
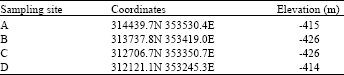
Sampling: Dead sea water samples were collected from four littoral zones (Fig. 1) in March, June and October, 2011. Geographic coordinates and elevation of the sampling sites are shown in Table 1. Geographic coordinates and elevation were determined for each location by (eTrex Legend C, Taiwan). Dead Sea water samples were collected in 1 L clean sterile glass bottle leaving enough head space in the bottle and transported immediately to the lab.
Physicochemical analysis of the samples: Temperature, pH, Total Dissolved Solids (TDS) and Biological Oxygen Demand (BOD) of water samples were determined. Water temperature, pH and salinity were measured in situ. Water temperature and pH were measured by a portable pH meter (Microcomputer pH meter T19000, Trans Instruments). Salinity was measured by a handheld salinity refractometer. BOD was measured in (Water, Environment and Arid Regions research Center at Al al-Bayt University, Jordan). Water samples were transferred to a new glass bottle, then 1 mL of phosphate buffer, magnesium sulfate, calcium chloride, iron chloride solutions per liter water sample were added. The sample was then brought to temperature 20±3°C and saturated with organic-free filtered air. The pH of the sample was checked. If the sample was not in the range of 6.5-7.5, sulfuric acid or sodium hydroxide was added to bring the sample to the required pH range (6.5-7.5) and the added concentration did not dilute the sample more than 0.5%. The sample was brought to temperature 20°C before making dilutions. Suitable volume of the sample was then transferred to BOD bottles. The bottle was filled with enough dilution water to displace all air leaving no bubbles. Dissolved Oxygen (DO) was determined using a DO analyzer and any displaced contents were replaced with dilution water and stopper tightly. The bottle was incubated for 5 days at 20°C. After that, DO was determined and BOD was calculated as following the difference between initial and the final DO over volume.
Heterotrophic viable bacterial count: To count bacteria in our samples, we used the viable plate count method. Ten microliters of water sample were transferred to solid high salinity medium (ingredients per liter distilled water: MgCl2.6H2O, 5,67 g; MgSO4.7H2O, 6,8 g; NaHCO3., 0.19 g, CalCl2.2H2O, 1,47 g; KCl, 0,72 g; KH2PO4, 0.5 g, peptone, 10 g; glycerol, 3 g; yeast extract, 2 g; NaCl, 30 g, agar, 18 g. Inoculated plates were incubated at 30°C in dark. After incubation, colonies were counted as CFU mL-1 original sample.
| Table 1: | Geographic coordinates of the sampling sites |
 | |
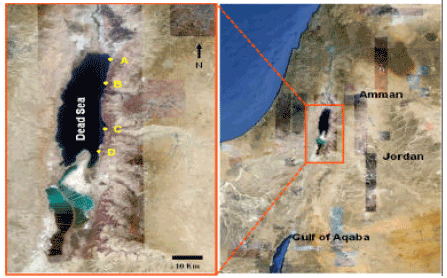 | |
| Fig. 1: | Map of the Dead Sea showing the four sampling sites (in yellow): A, B, C and D. The maps is retrieved from Google Earth |
Enrichment, isolation and gram staining: To enrich water samples, Dead sea water (10 mL) was transferred to 90 mL of liquid high salinity medium and incubated at 30°C in dark with shaking (100 rpm). After about 12 h, a loopful of enrichment culture was streaked on modified solid mineral salt medium and the separated colonies were then subcultured. Glycerol stocks of the isolates were also prepared and stored at -20°C for further analysis. Cells were Gram-stained and examined under the microscope.
16S rRNA gene sequencing and analysis: 16S rRNA gene sequence amplification and sequencing was carried out by Macrogen Inc., Seoul, Korea, according to the following method: young colonies of the bacterial strains were suspended in a 1.5 mL centrifuge tube containing 0.5 mL sterile saline solution and then centrifuged at 10,000 rpm for 10 min the supernatant was removed and the pellet was suspended in 0.5 mL of InstaGene Matrix (Bio-Rad, USA) and incubated at 56°C for 30 min and then heated to 100°C for 10 min. After heating, supernatant was used for PCR. PCR was carried out by mixing 1 μL of template DNA with 20 μL of PCR reaction solution. 27F/1492R primers (27F: 5’-AGA GTT TGA TCM TGG CTC AG-3’, 1492R: 5’-TAC GGY TAC CTT GTT ACG ACT T-3’) were used for amplification and then 35 amplification cycles were performed at 94°C for 45 sec, 55°C for 60 sec and 72°C for 60 sec. Unincorporated PCR primers and dNTPs were removed from PCR products by Montage PCR Clean up kit (Millipore). The purified PCR products of approximately 1,400 bp were sequenced by 518F/800R primers (518F: 5’-CCA GCA GCC GCG GTA ATA CG-3’, 800R: 5’-TAC CAG GGT ATC TAA TCC-3’). Sequencing was performed by using Big Dye terminator cycle sequencing kit v.3.1 (Applied BioSystems, USA). Sequencing products were resolved on an Applied Biosystems model 3730XL automated DNA sequencing system (Applied BioSystems, USA) at the Macrogen, Inc., Seoul, Korea.
The sequences were analyzed and compared to the public nucleotide database using the NCBI BLAST website (http://blast.ncbi.nlm.nih.gov/blast/Blast.cgi). Sequences of the closest relatives were then retrieved from the database and used to construct a phylogenetic tree using MEGA5 (Tamura et al., 2011). GC content the sequences were calculated by Oligo Calculator (http://mbcf.dfci.harvard.edu/docs/oligocalc.htmL).
Physicochemical properties of the samples: Dead Sea water samples were very saline ranging from (36-38%). The pH values of the samples were low and range from 5.6 to 6.3. This indicates the slightly acidic property of sea water. The BOD value were very low (1-2 mg O2 L-1) indicating very low organic material content in the sample. The physicochemical properties are shown in Table 2.
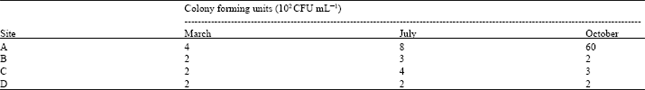
Heterotrophic viable bacterial count: Results of viable plate count revealed very low number of colony forming units in each sample (200-6000 CFU mL-1). The CFU count is shown in more details in Table 3.
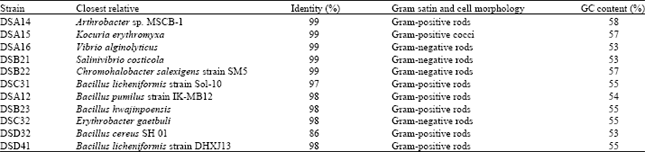
Isolation and classification of halophilic heterotrophic bacteria: In this study, we have isolated 44 halophilic heterotrophic bacterial strains. Eleven bacteria strains out of 44 were considered different based on colonial morphology, Gram staining and cell morphology. Seven out of eleven different strains were Gram-positive and 4 out of 7 were Gram negative. The bacterial strains were identified based on 16S rRNA gene analysis and they were found to belong to the domain of bacteria. The closest relative of each isolated bacterial strain is shown in Table 4 and Fig. 2. All sequences showed a relatively High GC content (up to 58%).
| Table 2: | Physicochemical properties of Dead Sea water samples collected from 4 different sites, during March, July and October, 2011 |
 | |
| Table 3: | Heterotrophic halophilic bacterial count of the samples |
 | |
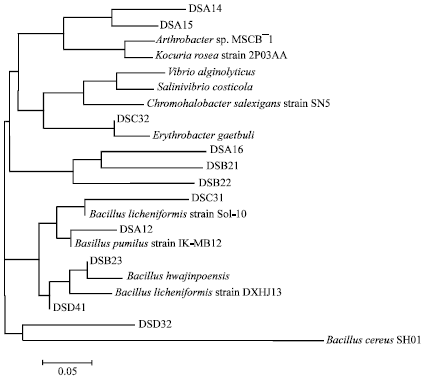 | |
| Fig. 2: | Phylogenetic tree of the isolated strains and their closest relatives based on 16S rRNA gene. The sequences were retrieved from NCBI website and the tree was constructed by MEGA5 software (Tamura et al., 2011). The evolutionary distances were computed using the maximum composite likelihood method (Tamura et al., 2004) |
| Table 4: | Closest relatives of the isolated strains with their percentage identity. Strains with more than 97% are considered strains of the same species |
 | |
DISCUSSION
The physicochemical properties of Dead sea water samples were determined and analyzed. The salinity percentage was found to be very high (up to 38%). This is a typical characteristic of Dead sea water making it one of the famous “athalassohaline” brines (Oren, 2007). The pH of dead sea water was found to be slightly acidic (5.6-6.3) as compared to pH (7.5-8) in thalassohaline brines (Oren, 2007). BOD of the samples were very low (1-2 mg O2 L-1) reflecting low organic material in water that could be utilized by heterotrophic bacteria. These physicochemical conditions undoubtedly affect negatively the microbial diversity. Therefore the number of viable cells in the tested samples was very low (not more than 6x102 CFU mL-1) as compared to open sea water. Counts of heterotrophic bacteria in marine waters are usually in the order of 105 to 106 bacteria mL-1 (Zweifel and Hagstrom, 1995; Madigan and Martinko, 2006). These numbers are derived from unspecific fluorescent staining techniques (Zweifel and Hagstrom, 1995) which usually gives higher numbers than viable plate count method. The first publications on bacterial count in dead sea was done by microscopy. The cell number was about 1.9x106 cells mL¯1 and during a bloom of red halobacteria the population densities reached 1.9x107 but declined after moths to 5x106 (Oren, 1983).
Research on halophilic organisms inhabiting Dead Sea started very early in 1892 when bacteria was isolated from the genus Clostridium from mud (Oren, 2002). Later, in a short article published in 1936, gave the first description of an indigenous microbial community adapted to the extremely harsh conditions of the Dead sea (Oren and Ventosa, 1999). Since that time, our knowledge about the biological aspects of the Dead sea is expanding and accumulating. We conducted this research to expand our knowledge about the halophilic heterotrophic bacteria thriving in littoral zone of the Jordanian Dead sea. We have isolated different bacterial species. Subsequently, we isolated and identified 11 different species of halophilic bacteria. Most isolates were Gram-positive (7 out of 11). Even though Gram positive bacteria posses important adaptations enable them to coup with environmental stress such as high salinity (Battistuzzi and Hedges, 2009). It is not clear in the literature whether Gram-positive bacteria or Gram-negative bacteria are dominant in hypersaline environments. In one study on prokaryotic halophiles recovered from sediments from the shallow El-Djerid salt lake in Tunisia, Hedi et al. (2009) found that the dominant bacterial population belongs to Gram-positive spore-forming bacteria.
All of the isolated strains in this study belong to the domain of Bacteria. They belong to 7 different genera in the domain. Five out of seven Gram-positive bacterial isolates belong to genus Bacillus. This genus was established in 1872 to include three species, but now there are 142 named Bacillus species listed in Bergey’s Manual of Systematic Bacteriology (Logan and de Vos, 2009). Bacillus strains are typically soil strains. Bacillus strains isolated in this study belong to the following species B. licheniformis, B. pumilus, B. hwajinpoensis and B. cereus. These strains were encountered in different salty environments (Miranda et al., 2008; Parvathi et al., 2009; Yoon et al., 2004; Al-ZaZaee et al., 2011). For instance, B. licheniformis was previously recovered from marine sediments by Miranda et al. (2008), while B. pumilus was isolated from marine organisms like oysters, crab and fish in addition to sediments by Parvathi et al. (2009). B. hwajinpoensis was recovered from seawater of the East sea and Yellow sea in Korea (Yoon et al., 2004). And finally B. cereus, a common soil bacterium, was encountered in sewage water but with halophilic properties (Al-ZaZaee et al., 2011). Thick cell wall, high peptidoglycan content (more than 90% of cell wall) (Madigan and Martinko, 2006), high GC content and resistant-spore formation are the main reasons for Bacillus survival in harsh conditions like high salinity. It should be noted that strain DSD32 (identified as B. cereus) has the very low similarity with its closest relative B. cereus. This strain may represent a new species in the genus Bacillus based on 16S rRNA gene.
The other two Gram-positive bacteria are strains of Arthrobacter sp. and Kocuria rosea. Arthrobacter species are widely distributed in nature especially in soil (Funke et al., 1996) but is not very common in hypersaline environments. Kocuria rosea is considered as moderately halophilic species and it was recovered from different saline environments like saline open shallow water and some strains of Kocuria rosea grow optimally at NaCl concentration of 30% (Wright and Tanaka, 2002).
Gram negative bacteria were less frequent in our samples (4 out of 11). Nevertheless, published literature shows that these isolates are not uncommon in salty environments. One of the isolated strains was identified as Vibrio alginolyticus. The latter species is actually common in marine samples (Molitoris et al., 1985) and was encountered in marine sea water. The strain has medical importance because it may cause complicated skin and soft tissue infection (Sganga et al., 2009). More importantly, some strains of Vibrio alginolyticus were found to produce tetrodotoxin, a strong neurotoxin (Noguchi et al., 1987). Another Gram-negative bacterium identified in this study is Chromohalobacter salexigens. This species was first isolated and described as the moderately halophilic species Halomonas elongata and then proposed as new species of Chromohalobacter (Arahal et al., 2001). Erythrobacter gaetbuli is another species identified in this study. This strain is also not uncommon for saline habitats. It was recently isolated from tidal flat of the yellow sea in Korea and it was described as halophilic species by Yoon et al. (2005). The last strain belong to Salinivibrio costicola. This species was first described as Vibrio costicola but its phenotypic and genotypic characteristics distinguished it from species of the genus Vibrio. Therefore, the strain was placed in new separate genus named Salinivibrio (Mellado et al., 1996). Salinivibrio costicola is moderately halophilic and was originally isolated from salted food and grow optimally in media containing 10% salts (Mellado et al., 1996). A member of the genus Salinibacter, S. ruber, is an interesting model for the study of the adaptation of microorganisms to life at high salt concentrations (Oren, 2007).
Dead Sea water from littoral zone is characterized by high salinity, low pH, low organic material content (low BOD) and restrain different species of heterotrophic bacteria belonging to both Gram-positive and Gram-negative genera including Arthrobacter, Kocuria, Vibrio, Salinivibrio, Chromohalobacter, Bacillus and Erythrobacter.
This study was supported by the deanship of academic research at Al al-Bayt University, Jordan, decision of the Scientific Research Council in meeting number 2/2010/2011. Thus, the author would like to appreciate the financial assistance provided by the University.