Research Article
Diagnosis of Neonatal Bacterial Sepsis by Polymerase Chain Reaction
Department of Pediatric, Cairo University, Cairo, Egypt
Wahiba A. Zarouk
Department of Molecular Human Genetics, National Research Center, Egypt
Sepsis is a serious disease with high mortality in newborns, particularly in preterm, low birth weight infants. A fast and correct diagnosis, followed by rapid treatment, plays an important role in the reduction of infant mortality resulting from sepsis (Kaufman and Fairchild, 2004).
The incidence of neonatal sepsis ranges from 1 to 8 cases for every 1000 infants, but much higher values are found in preterm, in low birth weight newborn infants and generally, in newborn infants admitted in neonatal intensive care units (Hervas et al., 1993). Several studies have previously tried to find a clear-cut definition of sepsis in neonates, based on a combination of clinical signs and laboratory parameters (Benitz et al., 1998). However, diagnosing neonatal sepsis is difficult since being exposed to known risk factors for sepsis is not a necessity (Thomas and Baker, 1995) clinical signs are often vague and laboratory parameters are unspecific. Although, CRP serves as a good example of haw biochemical markers help the neonatologist in distinguishing infected from non infected infants, it is not useful as an early phase infection marker but has good diagnostic utilities in the later phase of sepsis as both sensitivity and negative predictive value in early evaluation of sepsis are 14 and 75%, respectively. Also, the wide range of normal variation of total and differential white cell counts plus the subjective nature of white cell morphology make the routine hematological investigation in diagnosis sepsis limited (Lam and Ng, 2008). Conventional blood culture is considered the gold standard in the etiological diagnosis of neonatal bacterial sepsis (Gerdes, 1991). Although, in some studies have shown that the culture sensitivity was low (Jordan et al., 2006). The reasons for this low sensitivity were small blood volume obtained from neonates and wide use of prophylactic intrapartum antibiotics for high risk deliveries (Alfa et al., 1995). Because, the preliminary report of blood culture is not available before 48-72 h, it is customary to start treatment at birth in all neonates with risk factors for early sepsis and/or with alterations of some laboratory tests (i.e., white cell count, total neutrophil count, I/T ratio, CRP). However, the sensitivity and specificity of each laboratory test are far from 100% (Gerdes and Polin, 1987; Reier Nilsen et al., 2009). For these reasons, there is an obvious need for a more rapid, yet precise, tool which with the use of a limited blood volume can detect bacteremia if present and exclude it when absent.
Polymerase Chain Reaction (PCR) techniques have been used routinely for a number of years for detection of a wide range of microorganisms from different samples (Fenollar and Raoult, 2004). The nucleic acid amplification test is now considered the gold standard test for diagnosing neonatal meningitis due to herpes simplex virus (Jordan et al., 2006).
DNA sequence present in all bacteria such as portions of the DNA encoding the 16-S ribosomal-RNA has been used to define an organism as a bacterium. (Angert et al., 1993). Those sequences are amplified with PCR, allowing a rapid diagnosis (McCabe et al., 1995).
PCR detection of blood borne bacteria using highly conserved targets within the 16S ribosomal RNA gene (16S rDNA) has been described previously by Jordan and Dues (2002) published the first two large-scale studies for detecting sepsis in the neonate. These studies compared the results of culture to conventional PCR in which whole blood was first pre-enriched in Tryptic Soy Broth (TSB) for between 5 and 10 h at 35°C before sample preparation and testing occurred (Jordan and Dueso, 2005). In another study PCR with sensitivity of 100% and specificity of 95.6% is a useful tool in diagnosis of neonatal sepsis because of the short time required to reach confirmation. The PCR has shown 100% correlation with microbiological methods in all infected cases and was positive in four additional cases with positive sepsis screen but with sterile blood cultures (Yadav et al., 2005).
In a more recent study Wu et al. (2008) developed a method of simultaneous detection and gram classification of clinically relevant bacterial pathogens causing sepsis directly from blood samples with gram stain-specific-probe-based real-time PCR (GSPBRT-PCR) and it showed a sensitivity of 100% and specificity of 97.1% when blood culture was used as a control. Another study of Dutta et al. (2009) in diagnosis of neonatal sepsis using universal PCR before and after starting parenteral antibiotics, it was found that universal PCR can accurately diagnose neonatal sepsis before but not after antibiotic therapy.
The aim of this study was to find a rapid yet conclusive method for diagnosis of neonatal sepsis; in neonates with suspected sepsis instead of starting parenteral antibiotics while waiting at least 72 h till the results of blood culture will be available. PCR could provide valuable rapid information to rule out sepsis and this can help us to avoid the increasing risk antibiotics resistance and giving unnecessary antibiotics so PCR was compared to conventional blood culture for detecting bacterial DNA encoding 16S RNA in blood samples from neonates with suspected sepsis, also the relationship between clinical signs and laboratory parameters in cases with suspected and proven sepsis were explored.
The study was done from July 2008 till December 2008 in the Neonatal Intensive Care Unit (NICU) Cairo University. During this period the NICU received 180 neonates. The study included 75 newborns who were admitted to NICU with suspected sepsis. Inclusion criteria were signs of infection leading to initiation of treatment with intravenous antibiotics and doing blood culture as decided by the neonatologist, as the symptoms and signs of suspected bacterial sepsis were multiple and non specific patients with any of the following criteria were enrolled, feeding difficulties with or without abdominal distension, irritability or lethargy, temperature instability, development of apnea, respiratory rate more than 60 min-1, poor capillary refill ( more than 3 sec), heart rate more than 160 min-1. All patients were subjected to proper assessment of gestational age, birth weight, apgar scoring at 5 min, full clinical examination. Newborns with sepsis who started antibiotics prior to sampling, newborns with major congenital anomalies or history suspecting inborn errors of metabolisms or hypoxic ischemic encephalopathy and surgical problems were excluded from this study. Also, those with severe illness with expected death before 24 h of inclusion in the study were excluded.
A venous blood sample was withdrawn from each baby prior to start of antibiotics, 2 mL venous blood was collected for total leucocytes count, peripheral blood smear with differential count and C reactive protein an acute phase reactant (qualitative CRP), sepsis screen was considered positive if the ratio of immature neutrophils (Band cells and myelocytes ) to total neutrophils ( I/T ) > 0.2 and CRP was positive (Yadav et al., 2005; Philip, 1994). One milliliter for conventional BACTEC blood culture in the same time another 1 mL venous blood was taken on EDTA in vaccutainer for application of PCR technique.
Genomic DNA extraction: Genomic DNA was extracted from peripheral blood leukocytes by a salting out procedure (Lahiri et al., 1992). Briefly, 10 mL of blood were mixed with triton lysis buffer (0.32 M sucrose, 1% Triton X-100, 5 mM MgCl2.H2O, 10 mM Tris-HCl, pH 7.5). Leukocytes were spun down and washed with H2O. The pellet was incubated with proteinase K at 56°C and subsequently salted out at 4°C using a substrate NaCl solution. Precipitated proteins were moved by centrifugation. The DNA in supernatant fluid was precipitated with ethanol. The DNA pellet was dissolved in 400 μL H2O.
Synthesis of primers: Oligonucleotide primers were designed to be complementary to the highly conserved regions of the 3’ end of the 16s regions of the ribosomal RNA operons based on a computer alignment of ribosomal RNA sequences available through gene bank sequences of primers.
Amplification: Presence of bacterial DNA in blood samples was evaluated by amplifying the DNA region encoding 16 rRNA (861 base pair) using the following primers: F-5-AGAGTTTGAT-CCTGGCTCAG-3’ (15) and R-5’GGACTACCAGGGTACTT AAT-3’. A small volume of the sample (2 μL) was mixed in the PCR reaction (briefly, 50 mM KCI, 10 mM Tris-HCL (pH-8), 1.5 mM MgCl2, 200 μL each d NTP, 60 pM each primer and 2.5U Taq polymerase) (Laforgia et al., 1997). The amplification used an initial modifying denaturation step (3 min at 94°C and 3 min at 55°C for 3 times) followed by 30 cycles (1 min at 95°C, 1 min at 54°C and 1 min at 72° C). A final extension cycle of 72°C for 7 min was carried out. The amplified DNA was separated by agarose gel electrophoresis stained with ethidium bromide and visualized under UV transilluminator. Analysis was carried out keeping blood culture as the gold standard (Jun-Fen et al., 2002).
PCR amplification in 2% agarose gel electrophoresis is considered positive if bacterial DNA region encoding for 16S RNA can be amplified at 860 base pair (bp) band.
Statistical analysis: The statistical program SPSS Version 12.01 was used for data analysis. Numerical data was expressed as a Mean±SD. Comparison of means for changes in variables was performed using the Paired Student t-test, while the non-Paired Student t-test was used for numeric comparison between two different groups. Nonparametric test for independent samples (the Mann-Whitney test) was used to compare linear variables between groups. For categorical variables the Fisher exact test was used. A p-level of <0.05 was considered statistically significant.
Of 75 newborns who were enrolled in the study 41 newborns were blood culture positive 2 of them were excluded as their blood cultures were positive for fungal infection and a 34 newborns were blood culture negative.
Table 1 shows the distribution of gestational age, birth weight and 5 min Apgar score among both groups, it showed no statistically significant difference between both groups as regards gestational age and birth weight however 5 min apgar score was significantly lower in blood culture positive group (p = 0.000).
Table 2 shows comparison of clinical and laboratory parameters in patients with blood culture positive and patients with blood culture negative. There was no statistical significant difference between both groups as regards gender. It showed that the most common presentation was feeding difficulty (p = 0.000) with abdominal distension (p = 0.003).
| Table 1: | Comparison of gestational age, birth weight and 5 min Apgar score among blood culture positive and negative patients |
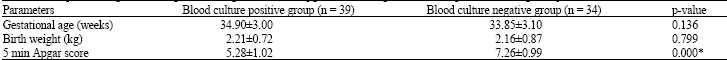 | |
| *p-value of <0.05 was considered statistically significant | |
| Table 2: | Comparison of clinical and laboratory parameters in patients with blood culture positive and patients with blood culture negative |
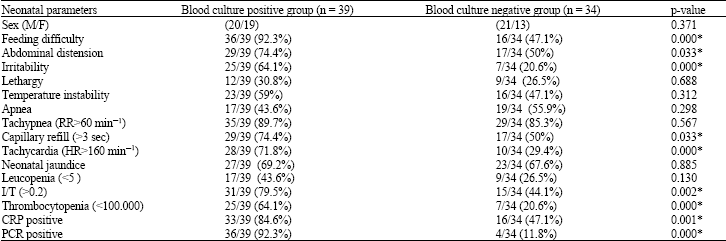 | |
| *p-value <0.01 | |
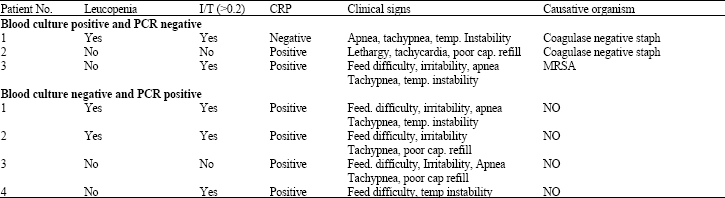
| Table 3: | Profile of patients with discrepant blood culture and PCR results |
 | |
There was statistically significant difference between both groups as regard, tachycardia (p = 0.000) and poor capillary refill (p = 0.033). Also, presence of irritability was statistically significant in blood culture positive group with (p = 0.000).
However, neither lethargy, temperature instability, apnea, tachypnea nor neonatal jaundice showed any significant difference between both groups. Regarding laboratory results we found that there was statistically significant difference between both groups as regard (I/T>0.2) was present in 79.5% of blood culture positive group and 44.1% of blood culture negative group with (p = 0.002), thrombocytopenia less than 100.000 was found in 64.1% of blood culture positive group and 20.6% of blood culture negative group with (p = 0.000), CRP was positive in 84.6% of blood culture positive and 47.1% of the blood culture negative group with (p = 0.001) and PCR was positive in 92.3% of blood culture positive group and 11.8% of blood culture negative group with (p = 0.000). while leucopenia showed no statistically significant difference between both groups.
In the present study both Klebsseila and Staph coagulase negative were the commonest causative organisms for neonatal sepsis followed by Acinobacter and Pseudomonas as shown in Fig. 1.
Of blood culture positive patients’ 36 patients have positive PCR while three patients have negative PCR and of blood culture negative group four patients have positive PCR and the reminder 30 have negative PCR. As shown in Table 3, the three patients with blood culture positive and PCR negative had Coagulase negative staph in two patients and MRSA in one. Whereas the other four patients with blood culture negative, PCR positive had sepsis screen positive and they present with signs and symptoms of neonatal sepsis.
Considering the blood culture as the gold standard in diagnosis of neonatal sepsis, sepsis screen had specificity more than sensitivity as shown in Table 4. The sensitivity, specificity, positive predictive value and negative predictive value of sepsis screen in detecting bacteremia in relation to blood cultures were 26/39 (66.6%), 27/34 (79.4%), 26/33 (78.7%) and 27/40 (67.5%), respectively.
| Table 4: | Comparison between sepsis screen and conventional blood culture |
 | |
| Table 5: | Comparison of broad range 16S rDNA and conventional blood culture |
 | |
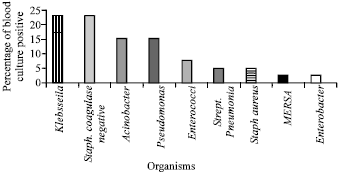 | |
| Fig. 1: | Common organisms causing neonatal sepsis in the studied groups |
Considering the blood culture as the gold standard in diagnosis neonatal sepsis, PCR had sensitivity more than specificity as shown in (Table 5). The sensitivity, specificity, positive predictive value and negative predictive value of PCR in detecting bacteremia relative to blood cultures were 36/39 (92.3%), 30/34 (88.2%), 36/40 (90%) and 30/33 (91%), respectively.
Figure 2 and 3 represent sample of PCR amplification. In Fig. 2 and 3 M lane represent 100 bp DNA ladder (the size marker) and some of studied population were represented in lanes each lane represent one patient.
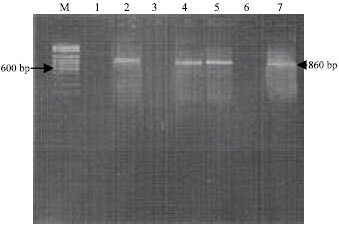 | |
| Fig. 2: | Agarose gel electrophoresis of samples run after Polymerase Chain Reaction (PCR): Lane M 100 bp pair DNA ladder; Lane 2, 4, 5, 7: Correspond to the yield of DNA’s patients amplifiable with the 16S RNA primer pair |
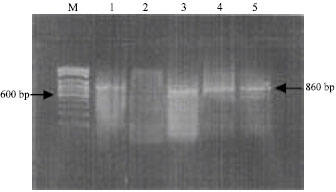 | |
| Fig. 3: | Agarose gel electrophoresis of samples run after Polymerase Chain Reaction (PCR): Lane M 100 bp pair DNA ladder; Lane 1, 3, 4, 5: Correspond to the yield of DNA’s patients amplifiable with the 16S RNA primer pair |
In Fig. 2 lanes 2, 4, 5, 7 PCR amplification yielded that 860 bp band so they were positive and those in lanes 1, 3, 6 did not yield that 860 bp band so they were negative.
In Fig. 3 patients in lanes 1, 3, 4, 5 PCR amplification yielded 860 bp band so they were PCR positive while the patient in lane 2 PCR amplification did not yielded 860 bp band so was PCR negative.
Early identification of infected neonate is fraught with difficulties as the clinical feature during the early phase of infection may be non specific and inconspicuous, routine microbiological and hematological investigations that are currently in common use have substantial limitations; the wide normal variation of differential white cell count, subjective nature of the white cell morphology, also CRP although still used during initial evaluation for sepsis, recent studies have concluded that CRP is best used through serial measurements when deciding whether to continue antibiotic therapy after initiation. Difficulty in obtaining culture specimens from various tissues, low sensitivity of the blood culture in neonates and often long period of time required for clinically useful results by conventional culture techniques (Lam and Ng, 2008). This diagnostic uncertainty leads to a greater use of broad spectrum antibiotics, a practice known to promote antibiotic drug resistance. Changing this practice will require newer diagnostic tests that are both rapid and accurate, which can provide results in a timely manner that can impact on the care of the patient (Jordan and Dueso, 2005). In addition, no available test can enable accurate diagnosis of bacterial sepsis in neonates exposed to antibiotic agents before collection of blood for culture. Universal primer Polymerase Chain Reaction (PCR) has been suggested as a potentially useful test in the diagnosis of neonatal sepsis in these settings (Weitkamp and Aschner, 2008). In the present study, exploration of some clinical presentation of neonates with suspected sepsis revealed that feeding difficulties with or without abdominal distension, tachycardia, poor capillary refill and irritability were more common in blood culture positive group however respiratory problems including tachypnea and apnea, temperature instability and jaundice were not specific for blood culture positive group and it is difficult to be distinguished from those of prematurity,that was supported by other studies (Reier Nilsen et al., 2009).
In this study sepsis screening was a poorer test than PCR in the diagnosis of blood culture positive sepsis The sensitivity, specificity, positive predictive value and negative predictive value of sepsis screen in detecting bacteremia relative to blood cultures were 26/39 (66.6%), 27/34 (79.4%), 26/33 (78.7%) and 27/40 (67.5%), respectively. The same was found in the study of Dutta et al. (2009) who found that sensitivity, specificity and positive predictive value of PCR were significantly higher than those of sepsis screening (Dutta et al., 2009). In the present study compared to blood culture the diagnosis of bacterial sepsis in newborns by PCR revealed sensitivity of (92.3%), specificity of (88.2%), positive predictive value of (90%) and negative predictive value of (91%),which were lower than those found in the study of (Laforgia et al., 1997), who found that PCR in diagnosis of neonatal sepsis has sensitivity of (100%), specificity of (93%), positive predictive value of (67%) and negative predictive value of (100%) their study included 33 newborns at risk for early sepsis and only 4 neonates had positive blood culture and all of them had positive PCR, PCR was positive in another 2 cases with negative blood cultures none of them had clinical signs of sepsis however in the current study 75 newborns with suspected sepsis and admitted in NICU were included so difference in sample size and inclusion criteria may be the cause of higher sensitivity in the study of Laforgia et al. (1997). However, study of Makhoul et al. (2005) showed sensitivity (69.2%) much lower than we found, specificity (100%), positive predictive value (100%) and negative predictive value (98%). This study was done on neonates with late onset sepsis and samples of DNA were PCR amplified by using a primer pair specific for staphylococcal 16S RNA and this cause of higher specificity.
In the study of Wu et al. (2008) PCR showed sensitivity of 100% and specificity of 97.1% when blood culture used as a control however the PCR used was gram stain specific probe based real time PCR which is different from the one used in the current study. Also that of Reier-Nielsen et al. (2009), found that the sensitivity was 66.7%, specificity 87.5%, positive predictive value 95.4% and negative predictive value was 75% with lower sensitivity than the current study and this could be explained by the lower number of positive blood culture as only six newborns were blood culture positive and 9 newborns were PCR positive out of 48 newborns. This discrepancy in sensitivity between 100 and 66.7% may be relayed to the pre enrichment step which was used in some previous studies Jordan et al showed a higher level of agreement between the two methodologies when preincubation was performed before PCR testing. They used 200-500 microlitters. EDTA-full blood preincubated at 37°C for 5 hours before PCRtesting and found 96% sensitivity, 99.4% specificity and 88.9% positive and 99.8% negative predictive values for PCR compared with the culturing of 0.5-1.0 mL full blood with BACTEC 9240.
However, a drawback with this procedure is that only live bacteria, able to grow in blood culture bottles will be detected. There is sufficient reason to show that speed is one of the potential advantages of PCR, an advantage that is somewhat lost if preculturing is used (Jordan et al., 2006; Jordan and Dueso, 2005). The PCR by itself cannot distinguish between asymptomatic bacteremia, contamination and true infection. The availability of clinical details and sepsis screening in this study served to better categorize the cases in which the PCR and blood culture reports were discrepant (Yadav et al., 2005; Jordan and Durso, 2000).
In the present study we had seven newborns with discrepancy between PCR and blood culture results. There were three newborns with blood culture positive PCR negative, their blood culture showed staph Coagulase negative in two of them and MRSA in the third one. This observation points to a need to change sample preparation protocols. Effective lysis of bacterial cells to liberate their DNA is crucial in achieving an assay with adequate sensitivity. Gram-positive organisms are more difficult to lyse than gram-negative ones because of their cell wall composition. Perhaps including an additional enzymatic treatment targeting gram-positive bacteria, using laybiase or achromopeptidase 31 or using glass beads to disrupt the cell wall more effectively would improve cell lysis (Niwa et al., 2005). Contamination of blood samples during blood drawing has been suggested to have led to an overestimation of the incidence of Staph coagulase negative-associated late onset sepsis (Karlowicz et al., 2000). However, in this study clinical course and sepsis screen of these three newborns were in favour of diagnosing them as neonatal sepsis and not contamination. This study showed that, there were four newborns with blood culture negative and PCR positive this can represents false positive results secondary to amplification of contaminated DNA because laboratory contamination is a well known problem in PCR methods (Ohlin et al., 2008). It is important to consider that PCR amplification is capable of identifying bacterial DNA, even if a very small amount is present in the blood, a quantity that could be not enough to determine a visible growth in usual culture conditions. Besides, according to recent data it has been demonstrated that when a small amount of blood is drawn and then subsided in different culture media, it is possible that up to 27% of sepsis are not diagnosed, particularly during the neonatal age (Laforgia et al., 1997). Analysis of data of these four neonates showed that their clinical picture and sepsis screen were consistent with sepsis although contamination can not be excluded. Specificity is difficult to interpret as false positive PCR may actually be false negative four blood culture. However, present study is limited as we included neonates with both early onset sepsis (less than 72 h of life) and late onset sepsis (more than 72 h of life) and 16S rDNA PCR performances. May vary in these two subpopulation. Although maternal data are important we were obliged to exclude them from statistical analysis as some of them were missed also we did not measure serial and quantitative CRP. Although, PCR technique is quite coasty and needs trained personnel, however its value in diagnosis of neonatal sepsis is important even in areas with poor resources as this coast is not comparable to the stay length in NICU, start of empirical antibiotics with raising risk of antibiotic resistance and the side effects associated with the antibiotic use, however large studies are recommended before PCR can be used for routine clinical application.
Keeping the blood culture as the god standard for diagnosis of neonatal sepsis, PCR method used in this study had correctly identified bacterial DNA encoding 16S rDNA in the majority of cases of positive blood culture neonates within 4 h with considerable sensitivity and when compared to blood culture PCR showed more sensitivity and specificity than sepsis screening in identifying bacterial sepsis. PCR has potential as a method for earlier detection of bacteria but this technology needs to be further developed and improved by using Gram Stain-Specific-Probe-Based Real-Time PCR. Blood culture is still irreplaceable at present, since pure isolates are essential for antimicrobial drug susceptibility testing.