Research Article
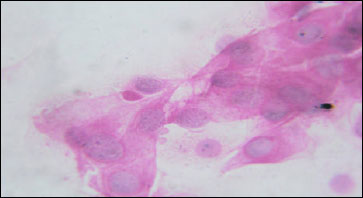
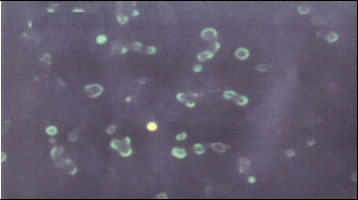
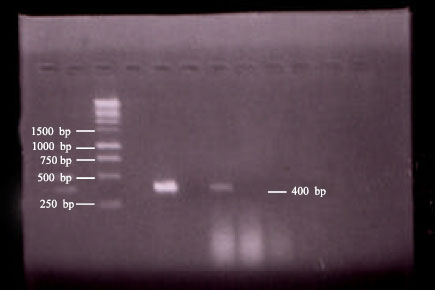
Isolation and Characterization of PI-3 Virus from Sheep and Goats
Animal Health Research Institute, Dokki, Giza, Egypt
H.A. Hussein
Faculty of Veterinary Medicine, Cairo University, Giza, 12211, Egypt
I.M. Reda
Faculty of Veterinary Medicine, Cairo University, Giza, 12211, Egypt