Research Article
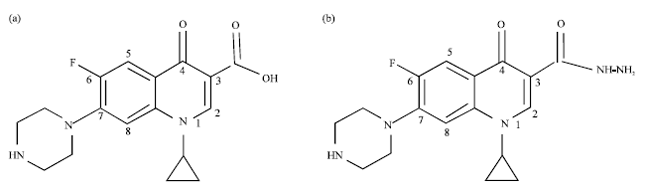
Effects of Substitution of Carboxyl with Hydrazide Group on Position 3 of Ciprofloxacin on its Antimicrobial and Antitumor Activity
Department of Molecular Biology and Biochemistry, Xinxiang Medical University, Xinxiang,Henan, 453003, People�s Republic of China
Yu Zhang
Department of Molecular Biology and Biochemistry, Xinxiang Medical University, Xinxiang,Henan, 453003, People�s Republic of China
Sufeng Zhou
Clinical Skill Training Center, Xinxiang Medical University, Xinxiang,Henan, 453003, People�s Republic of China
Youxun Liu
Department of Molecular Biology and Biochemistry, Xinxiang Medical University, Xinxiang,Henan, 453003, People�s Republic of China
Jiangang Wang
Department of Physiology, Xinxiang Medical University, Xinxiang,Henan, 453003, People�s Republic of China
Yali Wang
Department of Physiology, Xinxiang Medical University, Xinxiang,Henan, 453003, People�s Republic of China
Chengbiao Lu
Department of Physiology, Xinxiang Medical University, Xinxiang,Henan, 453003, People�s Republic of China
Changzheng Li
Department of Molecular Biology and Biochemistry, Xinxiang Medical University, Xinxiang,Henan, 453003, People�s Republic of China