Research Article
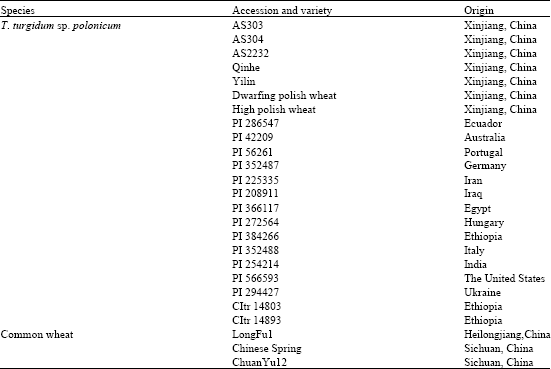
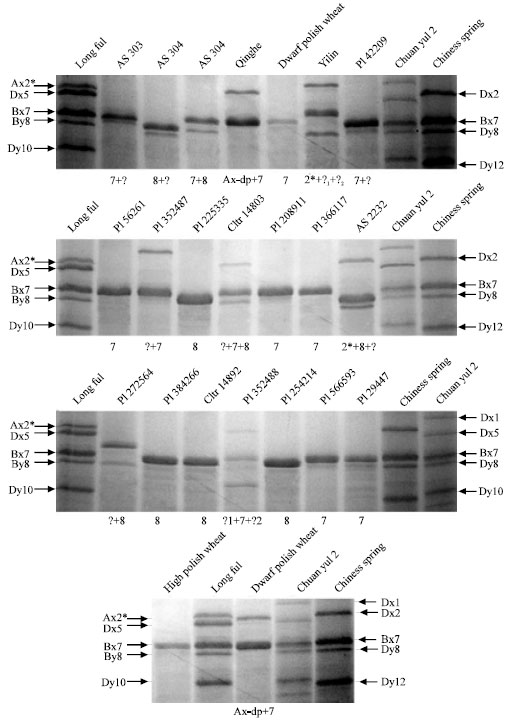
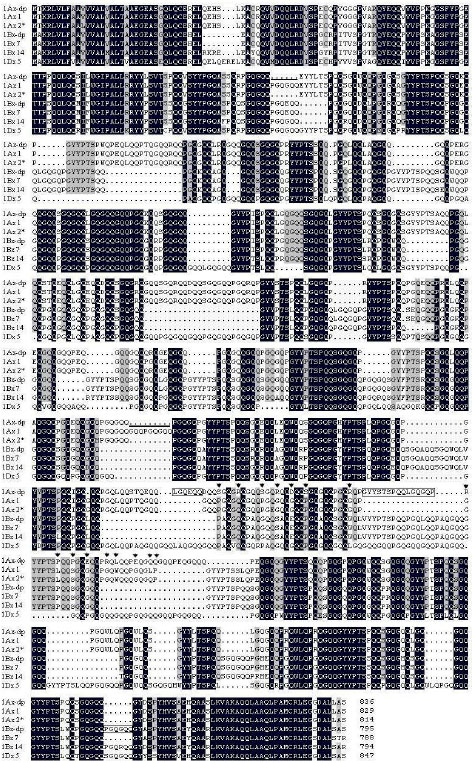
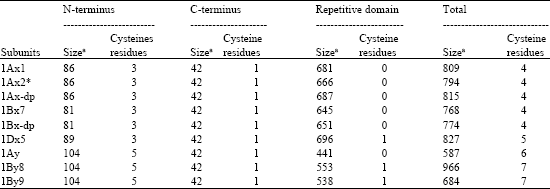
Identification and Characterization of HMW Glutenin Subunits and their Coding Sequences in Dwarfing Polish Wheat
Triticeae Research Institute, Sichuan Agricultural University, Wenjiang 611130, Sichuan, China
Zi-Jian Song
Triticeae Research Institute, Sichuan Agricultural University, Wenjiang 611130, Sichuan, China
Hou-Yang Kang
Triticeae Research Institute, Sichuan Agricultural University, Wenjiang 611130, Sichuan, China
Rui-Wu Yang
College of Biology and Science, Sichuan Agricultural University, Yaan 625014, Sichuan, China
Yong-Hong Zhou
Triticeae Research Institute, Sichuan Agricultural University, Wenjiang 611130, Sichuan, China