Short Communication
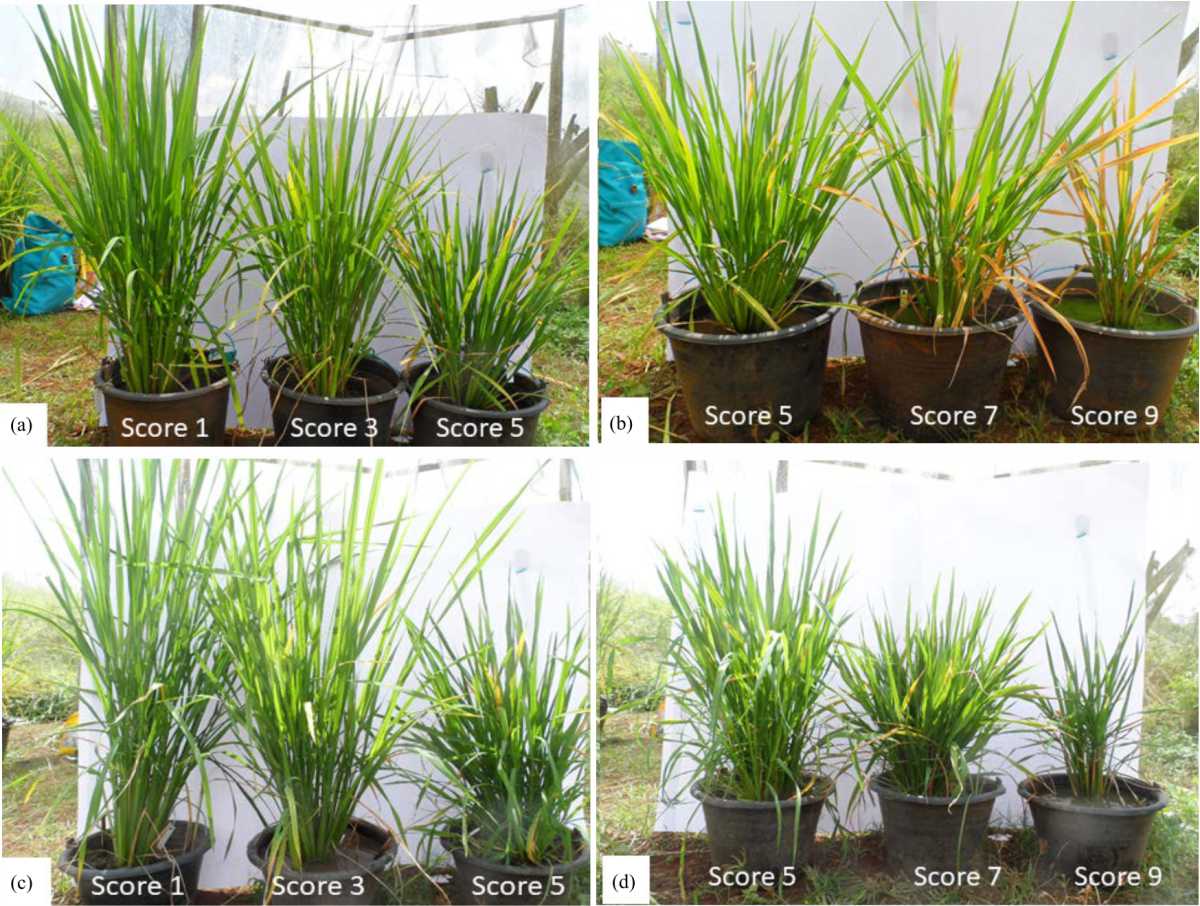
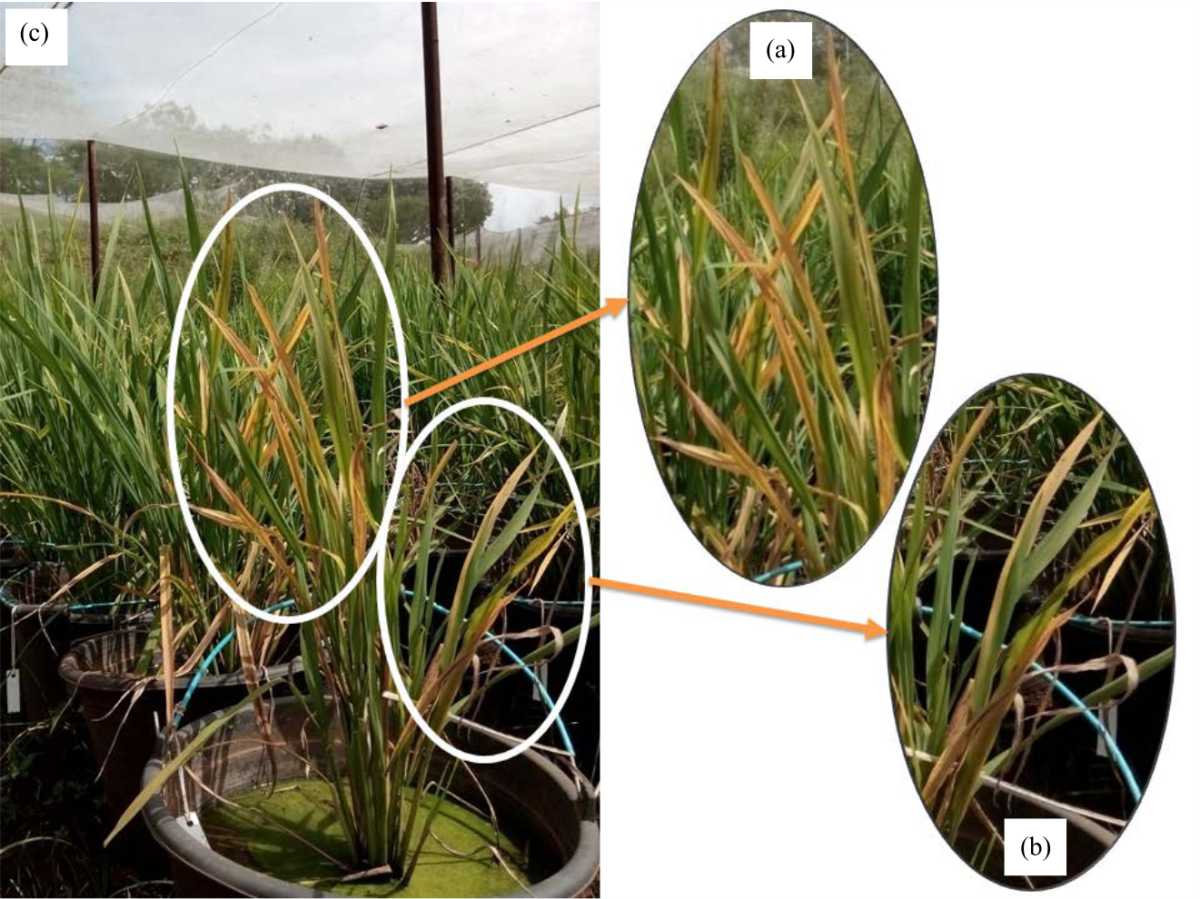
Segregation Ratios of Tungro Virus Resistance on F2 Progenies Derived from Utri Merah and ARC12596
Master Program of Crop Science, Faculty of Agriculture, Universitas Padjadjaran, 45363 Jatinangor, Indonesia
Fitri Widiantini
Faculty of Agriculture, Universitas Padjadjaran, 45363 Jatinangor, Indonesia
Santika Sari
Faculty of Agriculture, Universitas Padjadjaran, 45363 Jatinangor, Indonesia
Nono Carsono
Faculty of Agriculture, Universitas Padjadjaran, 45363 Jatinangor, Indonesia
LiveDNA: 62.31343