ABSTRACT
Full length cDNA of wheat MCM2 was used to synthesize the complete MCM2 protein in E. coli. The polyclonal antibodies against the synthesized protein was generated in the rabbit and used for cellular localization of the MCM2 protein during cell cycle. The immunostaining of root cells of wheat shows that MCM2 was localized inside the nucleus and exhibits cyclic synthesis and degradation during cell cycle. On the other side, the over expression of wheat MCM2 protein in Saccharomyces cerevisiae was associated with increase in cell size and protrusion of several buds at different points on the cell surface.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajps.2007.1093.1097
URL: https://scialert.net/abstract/?doi=ajps.2007.1093.1097
INTRODUCTION
MCM genes (minichromosomal maintenance) are a gene family of six members MCM2-7 which were first identified in the budding yeast and the fission yeast either as genes required for replication of minichromosomes or genes required for the progression of the cell division cycle (Takahashi et al., 1994; Okishio et al., 1996). The MCM proteins share common structures like the presence of zinc binding motifs (Fletcher et al., 2003) and conserved MCM box motifs (Hingorani and O’Donnell, 2000; Kearsey and Labib, 1998). However, both MCM2 and MCM3 have special structures like the presence of Nuclear Localization Sequences (NLS) (Young et al., 1997). The study on MCM2 protein suggests that the NLS of MCM2 is essential for its translocation and perhaps cotranslocation of other members of the MCM2-7 family (Tye, 1999).
All MCM mutants appear to affect the same set of chromosomal Autonomous Replicating Sequences (ARSs), leading to the suggestion that these proteins may function as a hexameric complex (Gibson et al., 1990; Lei et al., 1997). The complex assembly is essential for function, since no mutants that disrupt complex assembly are viable (Dalton and Hopwood, 1997; Lei et al., 2002; Sherman et al., 1998). The controlling mechanism of MCM protein function still not clear and the recent research results suggest the participation of two mechanisms for such regulation. The nuclear envelope allows entry of MCM protein into the nucleus, the other part of regulation includes posttranslation regulations which determine their binding to chromatin (Madine et al., 1995; Young et al., 1997; Aparicio et al., 1997).
This study is aimed to analysis the over expression of wheat MCM2 on the shape and division of yeast cell. The purified recombinant protein is used for antibody detection the nuclear localization of MCM2 protein in the root cells of wheat.
MATERIAL AND METHODS
This study was done at the Genetic Engineering and Biotechnology Research Institute, Menoufiya University, Egypt.
The ectopic expression of Bacterial expression of fusion proteins: The full length cDNA (Gene Bank accession number AY532594) was generated using differentially expressed EST sequence (Balzer et al., 1996; Matzk et al., 1997). The fusion protein was expressed in E. coli HMS (DE3)103 cells after induction with 0.5 M isopropyl-1-thio-β-D-galactoside (IPTG) for 10 minutes at 37EC. Proteins were extracted after sonication in 1/10 volume washing buffer (4.29 mM Na2HPO4, 1.47 mM KH2PO4, 0.27 mM KCl, 0.137 M NaCl, 0.1% Tween 20, 0.002% sodium azide, pH 7.3). Following centrifugation, fusion protein was isolated by Ni-MAC chromatography Purification Kit (Novagen). After buffer exchange (10 mM Tris, 0.05 mM EDTA) and concentration with polyethylene glycol, the protein concentration was determined by Bradfrod method (Bradford, 1979). For antibody production, 100 μg of the purified protein was used for rabbit injection three times with three week interval.
Yeast transformation: The ectopic expression effect of WMCM2 was tested using yeast model. The full length WMCM2 cDNA was directionally ligated to Yeast expression vector pYes2 (invetrogen) through Kpn1/Xba1 restriction sites. The transformed yeast with pYes2 parental vector was used as a negative control. The competent yeast cell were prepared using the method of Dohmen et al. (1991).
PAGE of WMCM2 protein expressed by E. coli: Protein samples were separated by SDS-PAGE in 12% polyacrylamide gels according to Laemmli (1970) and Schagger et al. (1988).
Immunocytochemistry and cell staining: The root tips of wheat seedling were fixed with 3.7% (w/v) formaldehyde for 1h and then squashed on coverslips coated with poly-L-lysine. The cells were then treated with an enzyme solution containing 0.1% pectolyase Y23 and 1% cellulase Y-C (both from Seishin Co., Tokyo, Japan) for 5min and then with a detergent solution containing 1% Nonidet P-40 for 15min, as described in Hasezawa and Nagata (1991). Antibodies against recombinant WMCM2 expressed in E. coli (WMCM2pAb) were raised in rabbit and affinity purified by protein A column chromatography (pharmacia). The WMCM2pAb were applied at dilution 1:20 together with anti α tubulin mouse monoclonal antibody (mAb) at dilution 1:300 in PBS 3% BSA. After a 12 h incubation of slides containing squashed root tip meristems in primary antibody solution at 4°C, the slides were washed for 15 min in PBS and incubated in rhodamineconjugated antirabbit IgG and FITC conjugated antimouse IgG 1:80 in PBS 3% BSA for 1 h at 35°C both from Dianova (Hamburg, Germany). After final washes in PBS, the preparations were mounted in anti fade containing DAPI as a counterstain (1 mg mL-1 in PBS buffer) for 1 min to permit visualization of nuclear DNA and were finally stained with 0.01%. Toluidine blue solution to reduce autofluorescence. The images were captured with a Zeiss Axioplan 2 epifluorescence microscope equipped with a Spot 2e CCD camera. Images were pseudocolored and merged using Adobe Photoshop 6.0 software (Adobe).
Microscopy: The pYES2 vector plasmid was used for the expression of the recombinant WMCM2 in yeast. The expression of the introduced gene is induced by shifting the transformed cells to the media that contains galactose as the sole carbon source. The empty vector transformed cells were used as a control. Microscopic investigation were carried out at OD600 ~1. The following conditions were used for confocal laser stereo microscope stemi 200C (Zeiss, Jena, Germany) was used for yeast micrographs.
RESULTS
E. coli expression of complete WMCM2 cDNA: The full length WMCM2 cDNA of 2907bp was used to express the recombinant WMCM2 protein in the bacterial cells. Figure 1 shows the 108 kDa of the recombinant protein as a major protein in the total cell lysate. The lane C shows the purified recombinant protein after Ni column chromatography. The purified protein amount of 100 μg was used to inoculate rabbit for polyclonal antibody production.
Polyclonal antibody staining: The actively dividing cells of wheat root tips were stained with florescent pAWMCM2 (red) to localize the WMCM2 protein during cell division. The florescent monoclonal antibody for α-tubulin (green) was used to detect the cellular anaphase and Dabi staining for DNA (blue) were used to demonstrate the nuclear DNA replication through the cell cycle. the immunostaining did not show presence of WMCM2 protein in the cytoplasm rather it was restricted in the nucleus. At the onset of interphase the WMCM2 were located on the nucleus side of the nuclear membrane. The intensity of signal was gradually decreased as mitosis progressed (Fig. 2A).
Ectopic expression of WMCM2 in yeast cell: The conserved sequence of the MCM proteins throughout the eukaryotes provide the possibility to use yeast model for analyzing the over expression of the WMCM2 protein (Forsburg, 2004). The induction of the recombinant WMCM2 protein synthesis was turned on by adding a new media containing 2% of galactose sugar. The control phenotype was the transformed cells with the empty pYes2 yeast vector. The microscopic investigations demonstrate that cell size display significant variability in cell transformed with WMCM2 and those that transformed with empty vector (Fig. 3). The second effect of WMCM2 accumulation was the protrusion of several budding cells at different peripheral points on the cell surface and have similar size.
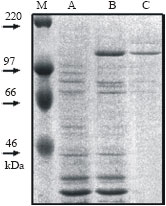 | |
| Fig. 1: | SDS-PAGE separation of E. coli expressed WMCM. (1) the E. coli total cell protein, (2) E. coli total cell protein after recombinant protein induction and (3) the affinity chromatography purified WMCM2 |
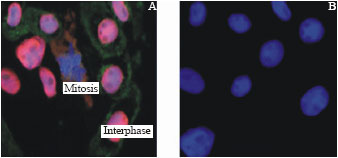 | |
| Fig. 2: | Subcellular localization of WMCM2 in the root cell of wheat. (A) the root cells, stained with anti-WMCM2 antibodies and rhodamine-conjugated secondary antibody (red), showing the nuclear localization of the WMCM2 at the interphase, immunofluorescence localization of α-tubulin using FITC-conjugate secondary antibody (green) and DAPI staining of DNA (blue). (B) root cells with DAPI staining |
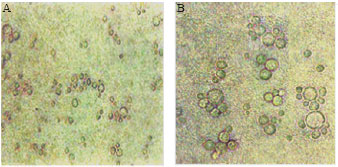 | |
| Fig. 3: | Micrographs of the S. cerevisiae (Broker C13 ABYS86 Strain) (A) the native strain. The cells were cultivated in minimal media with 2% Galactose and grown at 30°C for 16 h. (B) the transformed cells with WMCM |
DISCUSSION
The immunohistochemistry of polyclonal antibodies against WMCM2 shows that the WMCM2 is localized in the nucleus but it is not in the cytoplasm. Furthermore, it exhibits a cyclic appearance during the cell cycle (Fig. 2). The western data of MCM proteins during HeLa cell cycle demonstrate that MCM2 concentration decreases faster than other MCM species when the cell undergoes G1-S transition. However, the fluctuation pattern of MCM2 is quit similar to the pattern of origin recognition complex ORC protein (Kinoshita and Johnson, 2004). This results seems to be paradox with the early yeast work of (Thommes et al., 1992; Young and Tye, 1997) where the cyclic activity of MCM proteins were referred to the cyclic migration of the protein from the nucleus to the cytoplasm and reverse. The present result support the idea of proteolytic degradation of MCM2, actually the cyclic degradation of many cell cycle proteins is well established. The G1 cyclin, G2 cyclin and Anaphase Promoting Complex (APC) are integrated in an alternative pattern to control the cyclic activation of the cell cycle proteins (Tyers et al., 1993; Blondel and Mann, 1996; Nasmyth, 1999). In this context, this work suggest that MCM2 is controlled by the G1 cyclin. The other MCM species are stayed much longer in the nucleus (Kinoshita and Johnson, 2004), this suggest that those MCM species may regulated by other cell cyclins.
The continuous ectopic expression of WMCM2 in yeast is associated with relatively large cell with multiple bud formation as shown in Fig. 3. The process of new bud emergence is controlled by the cell wall synthetic enzymes together with fusion of actin-directed secretary vesicles with specific bud plasma membrane receptors (Mulholland et al., 1994; Harold, 1995). As a part of cell cycle progress, The bud formation emerges at the start point (at G1-S transformation), the point at which MCM2 plays its role for DNA replication. Actually the G1 cyclins control both functions as well as other cell cycle evidences that are initiated at the G1-S transition. In this context, Nasmyth (1999) demonstrates that cln3 is the cellular sensor of cell size and couples the cell size with cell division. On the other part, cln3 and other G1 cyclins are involved in the activation of CDK and the subsequent proteolysis processes that lead to cell cycle transition. The yeast cln3 mutant executes start point at about twice size of wild type cell (Dirick et al., 1995). So the large size phenotype and multibud structure may mediated by functional disruption of the G1 cell cycle cyclins that link metabolic state (cell size) and DNA replication. The ectopic expression of WMCM2 did not disturb the DNA replication as the flow cytometry assay did not show differences in DNA content between the WMCM2 transformed yeast, empty vector transformed cells and untransformed cells (data not shown). The continuous expression of the WMCM2 keep a base line concentration of MCM2 protein in the cell. This constant protein concentration seems to be save for DNA replication but disturb the other processes of bud initiation.
ACKNOWLEDGMENTS
I would to thank Dr. H. Baumlein and Dr. A. Houben from the Institute of Plant Genetic and Crop Plant Research (IPK) in Gatersleben, Germany for their technical support and helpful advices. I would also thank Prof. Dr. M.E. Nassr and Prof. Dr. Adel A. Guirgis, Molecular Biology Dept., Genetic Engineering and Biotechnology Research Institute, Minufiya University. for critical revision of the manuscript.
REFERENCES
- Aparicio, O.M., D.M. Weinstein and S.P. Bell, 1997. Components and dynamics of DNA replication complexes in S. cerevisiae: Redistribution of MCM proteins and Cdc45p during S phase. Cell, 91: 59-69.
Direct Link - Dalton, S. and B. Hopwood, 1997. Characterization of Cdc47p-minichromosome maintenance complexes in Saccharomyces cerevisiae: Identification of Cdc45p as a subunit. Mol. Cell. Biol., 17: 5867-5875.
Direct Link - Dirick, L., T. Bohm and K. Nasmyth, 1995. Roles and regulation of Cln/Cdc28 kinases at the start of the cell cycle of Saccharomyces cerevisiae. Embo J., 14: 4803-4813.
Direct Link - Dohmen, R.J., A.W. Strasser, C.B. Honer and C.P. Hollenberg, 1991. An efficient transformation procedure enabling long-term storage of competent cells of various yeast genera. Yeast, 7: 691-692.
Direct Link - Gibson, S.I., R.T. Surosky and B.K. Tye, 1990. The phenotype of the minichromosome maintenance mutant mcm3 is characteristic of mutants defective in DNA replication. Mol. Cell. Biol., 10: 5707-5720.
Direct Link - Hasezawa, S. and T. Nagata, 1991. Dynamic organization of plant microtubules at the three distinct transition points during the cell cycle progression of synchronized tobacco BY-2 cells. Bot. Acta, 104: 206-211.
Direct Link - Hingorani, M.M. and M. O`Donnell, 2000. A tale of toroids in DNA metabolism. Nat. Rev. Mol. Cell Biol., 1: 22-30.
Direct Link - Kinoshita, Y. and E.M. Johnson, 2004. Site-specific loading of an MCM protein complex in a DNA replication initiation zone upstream of the c-MYC gene in the HeLa cell cycle. J. Biol. Chem., 279: 35879-35889.
Direct Link - Laemmli, U.K., 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227: 680-685.
CrossRefDirect Link - Lei, M., Y. Kawasaki, M.R. Young, M. Kihara, A. Sugino and B.K. Tye, 1997. Mcm2 is a target of regulation by Cdc7-Dbf4 during the initiation of DNA synthesis. Genes Dev., 11: 3365-3374.
Direct Link - Lei, M., I.H. Cheng, L.A. Roberts, M.A. McAlea and B.K. Tye, 2002. Two mcm3 mutations affect different steps in the initiation of DNA replication. J. Biol. Chem., 277: 30824-30831.
Direct Link - Madine, M.A., C.Y. Khoo, A.D. Mills and R.A. Laskey, 1995. MCM3 complex required for cell cycle regulation of DNA replication in vertebrate cells. Nature, 375: 421-424.
Direct Link - Matzk, F., H.M. Meyer, C. Horstmann, H.J. Balzer, H. Baumlein and I. Schubert, 1997. A specific alpha-tubulin is associated with the initiation of parthenogenesis in 'Salmon' wheat lines. Hereditas, 126: 219-244.
Direct Link - Mulholland, J., D. Preuss, A. Moon, A. Wong, D. Drubin and D. Botstein, 1994. Ultrastructure of the yeast actin cytoskeleton and its association with the plasma membrane. J. Cell Biol., 125: 381-391.
Direct Link - Okishio, N., Y. Adachi and M. Yanagida, 1996. Fission yeast nda1 and nda4, MCM homologs required for DNA replication, are constitutive nuclear proteins. J. Cell Sci., 109: 319-326.
Direct Link - Sherman, D.A., S.G. Pasion and S.L. Forsburg, 1998. Multiple domains of fission yeast Cdc19p (MCM2) are required for its association with the core MCM complex. Mol. Biol. Cell, 9: 1833-1845.
Direct Link - Takahashi, K., H. Yamada and M. Yanagida, 1994. Fission yeast minichromosome loss mutants mis cause lethal aneuploidy and replication abnormality. Mol. Biol. Cell, 5: 1145-1158.
Direct Link - Thommes, P., R. Fett, B. Schray, R. Burkhart, M. Barnes, C. Kennedy, N.C. Brown and R. Knippers, 1992. Properties of the nuclear P1 protein, a mammalian homologue of the yeast MCM3 replication protein. Nucleic Acids Res., 20: 1069-1074.
Direct Link - Tyers, M., G. Tokiwa and B. Futcher, 1993. Comparison of the Saccharomyces cerevisiae G1 cyclins: Cln3 may be an upstream activator of Cln1, Cln2 and other cyclins. EMBO J., 12: 1955-1968.
Direct Link - Young, M.R. and B.K. Tye, 1997. Mcm2 and Mcm3 are constitutive nuclear proteins that exhibit distinct isoforms and bind chromatin during specific cell cycle stages of Saccharomyces cerevisiae. Mol. Biol. Cell, 8: 1587-1601.
Direct Link - Young, M., K. Suzuki, H. Yan, S. Gibson and B.K. Tye, 1997. Nuclear accumulation of Saccharomyces cerevisiae Mcm3 is dependent on its nuclear localization sequence. Genes Cell, 2: 631-643.
Direct Link








