Research Article
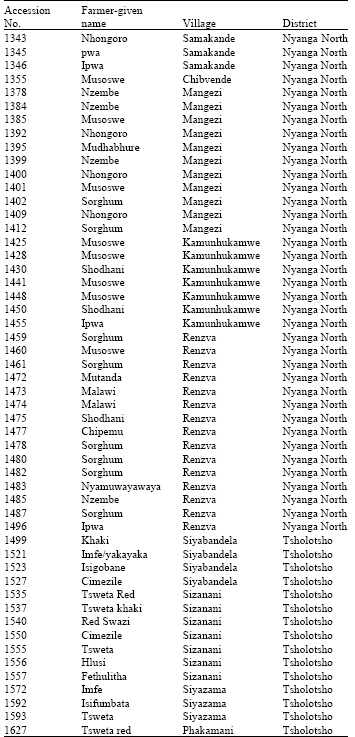
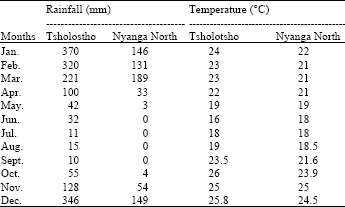
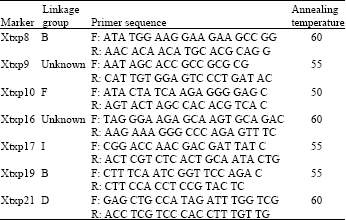
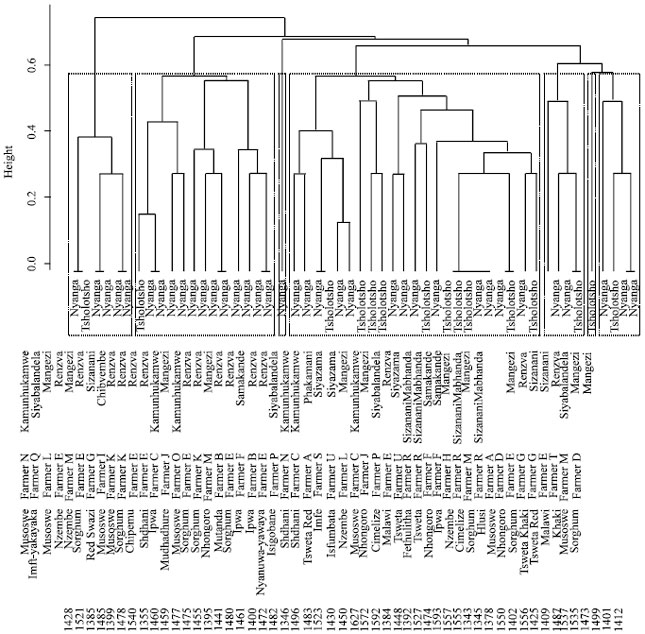
Genetic Diversity Assessment of Sorghum Landraces in Zimbabwe Using Microsatellites and Indigenous Local Names
ACCI, Plant Pathology, University of Natal, PI Bag XOl, Scottville, KwaZulu Natal, South Africa
P. Tongoona
ACCI, Plant Pathology, University of Natal, PI Bag XOl, Scottville, KwaZulu Natal, South Africa
E. A. Matibiri
Kutsaga Research Station, Airport Ring Road, Harare, Box 1909, Harare, Zimbabwe
M. Grum
Sub-Saharan Africa IPGRI, c/o ICRAF, P.O. Box 30677, Nairobi, Kenya