Research Article
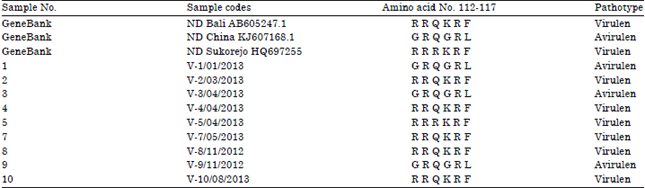
Molecular Pathotyping of Newcastle Disease Virus from Naturally Infected Chickens by RT-PCR and RFLP Methods
Department of Biochemistry and Molecular Biology, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Jl. Fauna 2, Karangmalang, Yogyakarta, 55281, Indonesia
LiveDNA: 62.12752
Vera Wati
Division of Biotechnology, Animal Disease Investigation Center Wates, Jl. Yogya-Wates Km. 27, Wates, Daerah Istimewa, Yogyakarta, Indonesia
Suhendra Pakpahan
Department of Biochemistry and Molecular Biology, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Jl. Fauna 2, Karangmalang, Yogyakarta, 55281, Indonesia
Nastiti Wijayanti
Laboratory of Animal Physiology, Faculty of Biology, Universitas Gadjah Mada, Jl. Teknika Selatan, Yogyakarta, 55281, Indonesia