Research Article
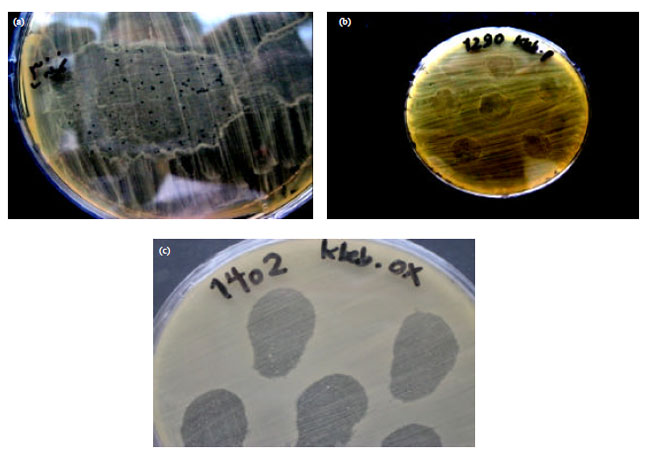
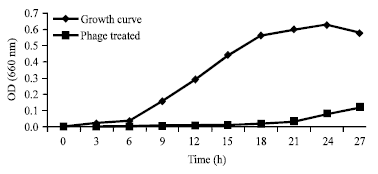
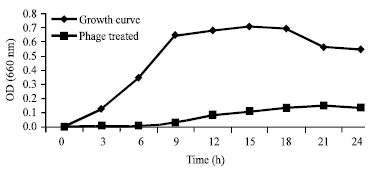
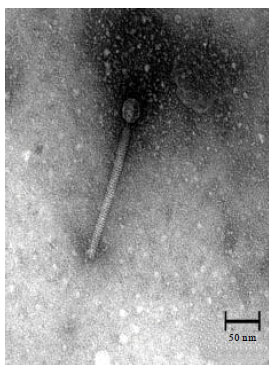
Isolation and Identification of Klebsiella pneumonia and Klebsiella oxytoca Bacteriophages and their Application in Wastewater Treatment and Coliform’s Phage Therapy
Department of Microbiology, Faculty of Biological Sciences, Falavarjan Branch, Islamic Azad University, Isfahan, Iran
Abbas Soleimani Delfan
Department of Microbiology, Faculty of Biological Sciences, Falavarjan Branch, Islamic Azad University, Isfahan, Iran
Sharareh Salmanizadeh
Department of Microbiology, Faculty of Biological Sciences, Falavarjan Branch, Islamic Azad University, Isfahan, Iran