Research Article
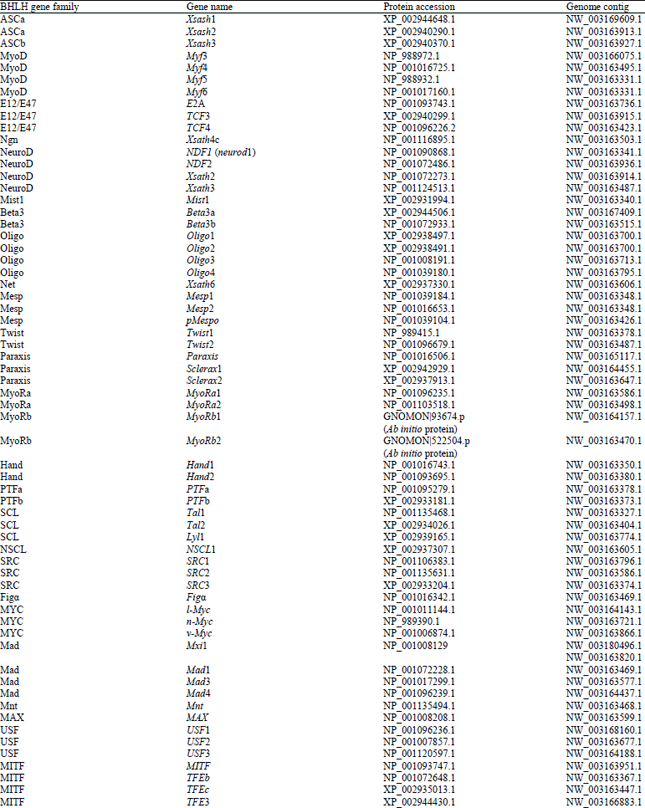
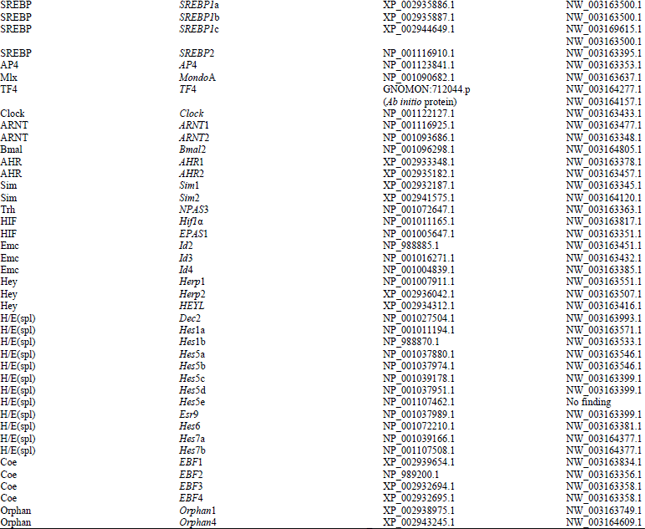
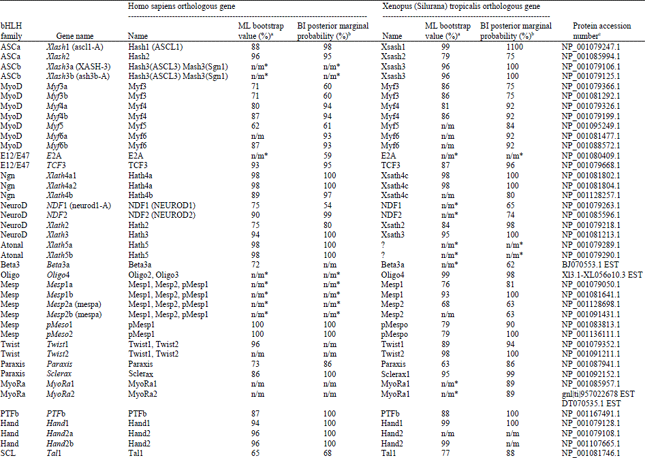
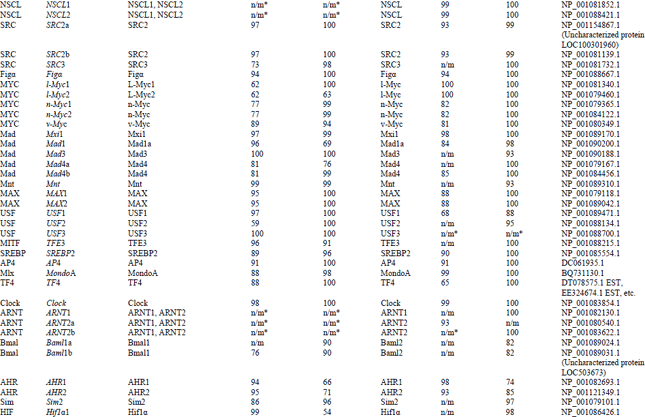
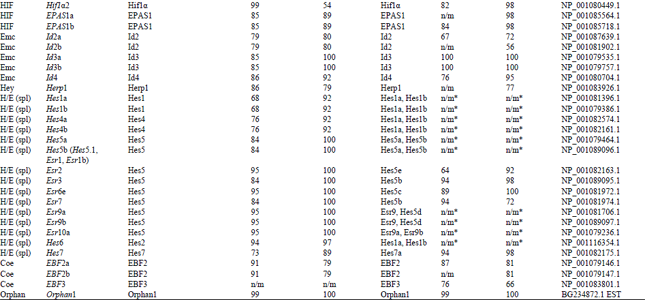
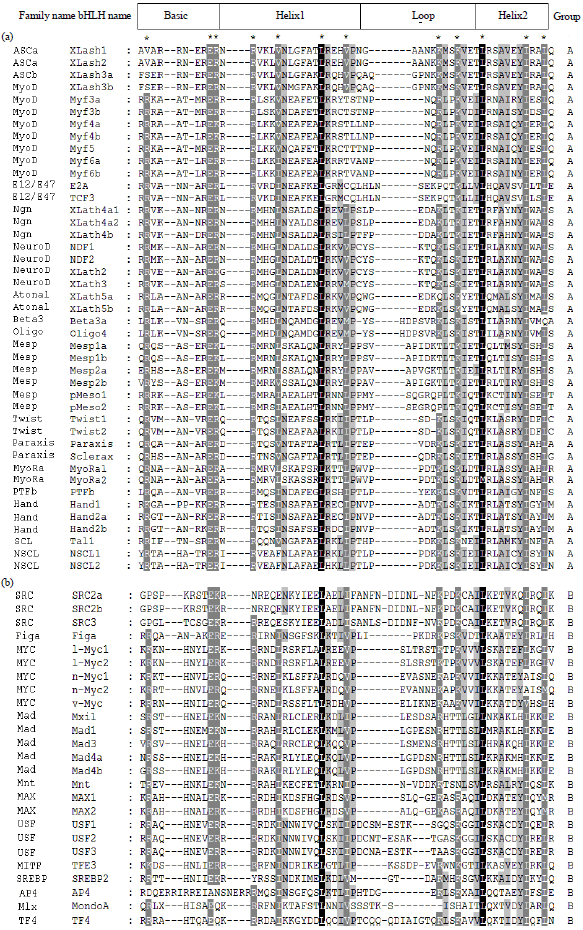
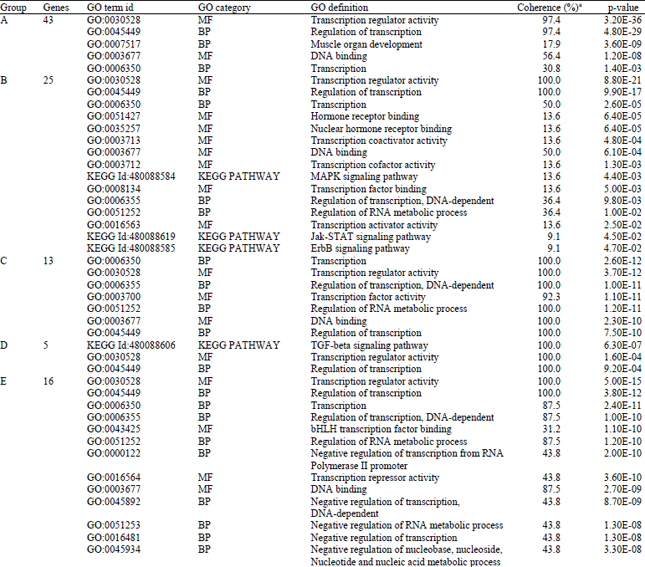
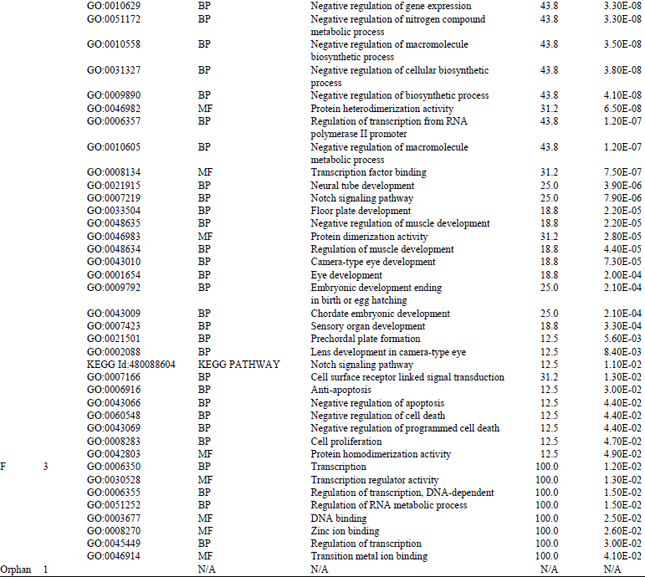
Identification and Bioinformatics Analyses of the Basic Helix-loop-helix Transcription Factors in Xenopus laevis
Department of Biology Sciences, Faculty of Biological and Food Engineering, Fuyang Normal University, Fuyang City, China
Fengmei Li
Department of Biology Sciences, Faculty of Biological and Food Engineering, Fuyang Normal University, Fuyang City, China