Research Article
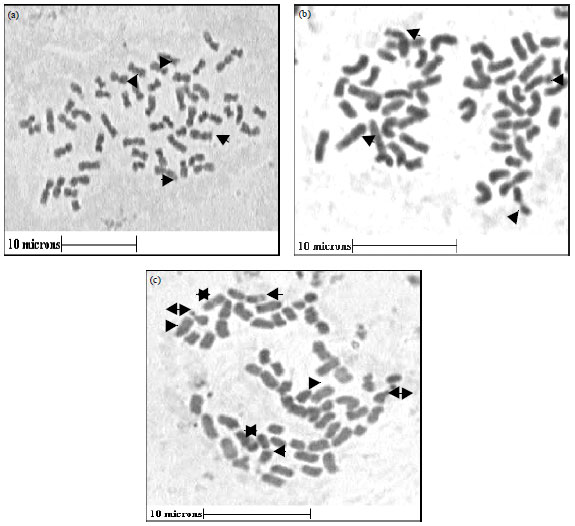
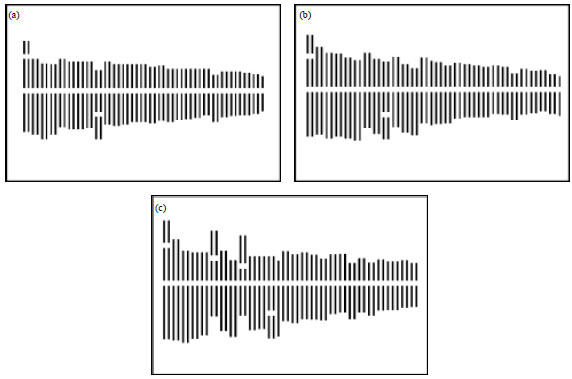
A Comparative Study of Karyomorphology among Three Populations of Garcinia indica (Clusiaceae) (Thomas-Dupetite) Choisy
K.E.T`s V.G.Vaze College, Mulund (East), Mumbai-400 081, India
Neetin Desai
Padmashree Dr. D.Y. Patil University, CBD Belapur, Navi Mumbai-400 614, India
Manjushri Deodhar
K.E.T`s V.G.Vaze College, Mulund (East), Mumbai-400 081, India