Research Article
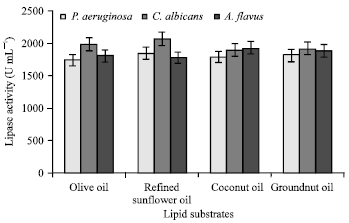
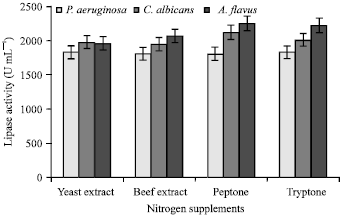
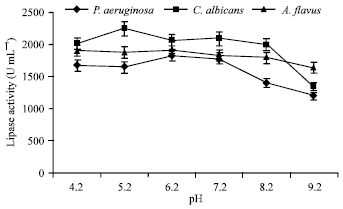
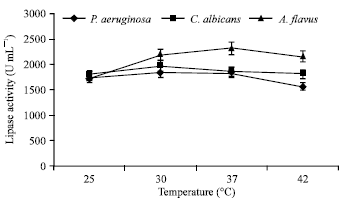
Optimization of Process Parameters Influencing the Submerged Fermentation of Extracellular Lipases from Pseudomonas aeruginosa, Candida albicans and Aspergillus flavus
Department of Microbiology, Genohelix Biolabs A Division of Centre for Advanced Studies in Biosciences, Jain University, India
Arijit Das
Department of Microbiology, Genohelix Biolabs A Division of Centre for Advanced Studies in Biosciences, Jain University, India
Sourav Bhattacharya
Department of Microbiology, Genohelix Biolabs A Division of Centre for Advanced Studies in Biosciences, Jain University, India