Research Article
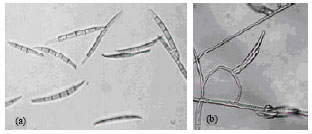
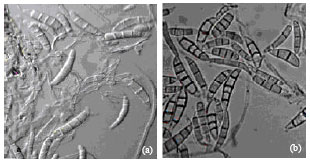
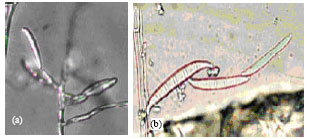
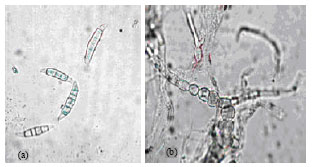
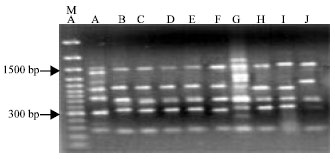
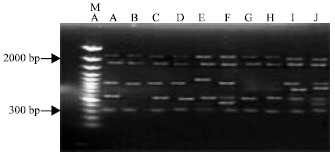
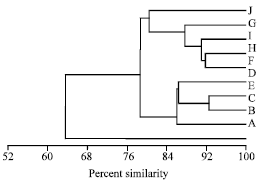
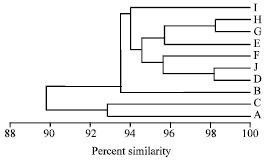
RAPD-PCR Analysis of Genetic Variation among Isolates of Fusarium graminearum and Fusarium culmorum from Wheat in Adana Turkey
Department of Plant Protection, Faculty of Agriculture, University of Suleyman Demirel, Isparta, Turkey
Namik Kemal Koc
Department of Plant Protection, Faculty of Agriculture, University of Cukurova, Adana, Turkey