Research Article
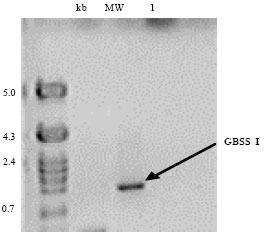
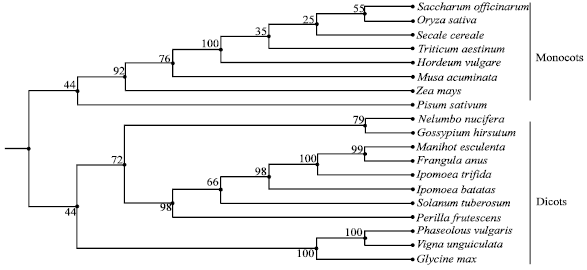
Isolation of Genomic Clones Encoding Granule-bound Starch Synthase (GBSS I) in Cassava (Manihot esculenta Crantz)
Department of Crop Production and Protection, Obafemi Awolowo University, Ile-Ife, Nigeria
O.O. Oyelakin
Central Biotechnology Laboratory, International Institute of Tropical Agriculture (IITA), Ibadan, Nigeria; c/o Lambourn Ltd, Carolyn House, 26 Dingwall Road, Croydon CR9 3EE, UK
O.A. Akinyemiju
Department of Crop Production and Protection, Obafemi Awolowo University, Ile-Ife, Nigeria
I.L. Ingelbrecht
Central Biotechnology Laboratory, International Institute of Tropical Agriculture (IITA), Ibadan, Nigeria; c/o Lambourn Ltd, Carolyn House, 26 Dingwall Road, Croydon CR9 3EE, UK