Research Article
Diversity and Morphology of Bacterial Community Characterized in Topsoil Samples from the Gaza Strip, Palestine
Department of Biology, Faculty of Science, Al Azhar University Gaza, P.O. Box 1277 Gaza, Palestine
The soil of the Gaza Strip was classified into three types: sand, clay and loess according to Jury et al. (1991). The sandy soil is located along the coastline along the Gaza Strip as sometimes hilly sand dunes with a thickness ranging from 2 to 50 m. Clay soil is found in the north eastern part of the Gaza Strip where loess soil is found around valleys (Wadis) of approximate thickness ranging from 25 to 30 m. A recent and more specific study categorized the soil in the Gaza Strip into several types including (1) Arenosolic regosols (sandy regosols), (2) Calcaric Arenosols (Loessial sandy soils) which can be found some 5 km inland in the central and southern part of the Strip, in a zone along Khan Younis toward Rafah, parallel to the coast, (3) Calcaric soils (Loess) are found in the area between the city of Gaza and the Wadi Gaza, (4) Arenosolic Calcaric (Sandy loess) these soils can be found in the depression between the Calcareous (Kurkar) ridges of Deir El Balah, (5) Arenosols calcaric over Calcaric (sandy loess over loess) that have been covered by a layer (0.2-0.5 m) of dune sand. These soils can be found east of Rafah, (6) Luvisols, Xerosols (Dark brown/Silty clay), which are dominated by loamy clay textures, are found on the slopes of the northern depressions between Beit Hanoun and Wadi Gaza (Shomar et al., 2010).
Soil furnishes an appropriate habitat for many microorganism included bacteria. Bacteria and other microorganisms play an important role in various natural, biological, chemical and geological activities (Molin and Molin, 1997; Wall and Virginia, 1999). One gram of soil contains more than one billion bacteria with probably one thousand various species (Torsvik and Ovreas, 2002; Torsvik et al., 1990; Wise et al., 1997).
Consequently, soil morphology and characteristics are highly influenced by various microbial vital activities (Batisson et al., 2009; Dastager et al., 2009; Cairney, 2000; Klironomos et al., 2000; Van der Heijden et al., 1998), which change the soil structure (Wright and Upadhyaya, 1998; Dodd et al., 2000) and the soil fertility (Yao et al., 2000; O’Donnell et al., 2001).
This project is unique since it is the first study to identify the allocation of bacterial communities and their characteristics in various topsoil samples existing in the Gaza Strip.
The Gaza Strip is a highly congested area of 1.38 million residing in a small piece of land of 365 km2 (L of 40xW of 7-12 km) (The Palestinian Central Bureau of Statistics, 2008). It is divided into five governorates, the northern area, Gaza, the middle area, Khan Younis and Rafah (Fig. 1) (Bashir and Adnan, 2011). The Gaza Strip is located east of the Mediterranean and has a four-season climate of a hot and humid summer and mild rainy winter.
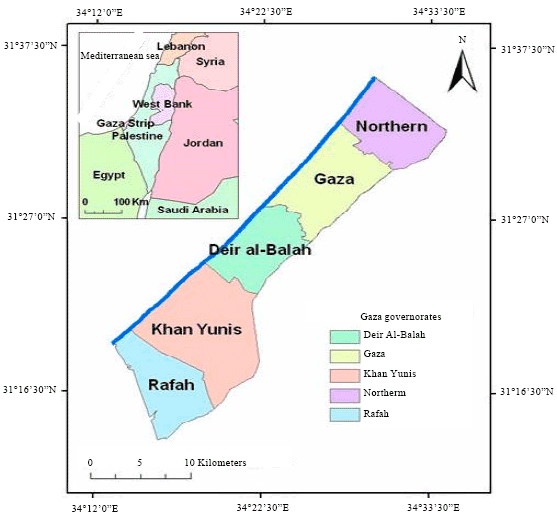 | |
| Fig. 1: | Map of Gaza Strip |
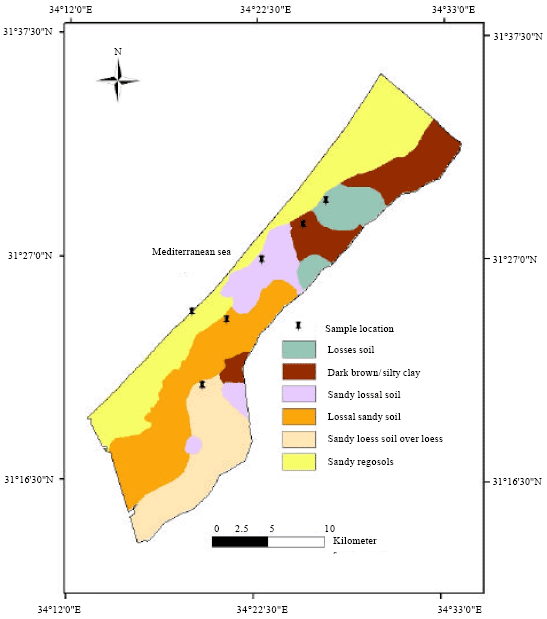 | |
| Fig. 2: | Soil map of Gaza Strip |
Soil sampling: Soil samples were collected according to the European soil sampling guidelines (Theocharopoulos et al., 2001).
Surface soil samples obtained were selected from different locations in the Gaza Strip applying the Global Positioning System (GPS) technique (Fig. 2).
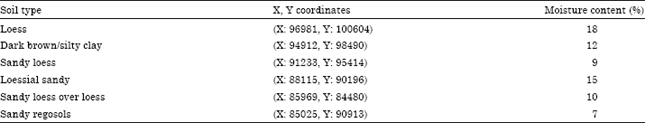
Samples from six soil types are were randomly collected regarding geographical locations of site determined in X and Y coordinate (Fig. 2). The moisture content of soil was determined for each soil type and was expressed as a percentage moisture content (Table 1).
The soil samples were collected in the winter season (December 2011-January 2012), of average temperature 10-15°C. Soil samples were taken from the top 0±10 cm for the open and grass soils, 0±20 cm for the vegetable soils. To obtain the required amount of soil, a long metal corer (25x100 mm I.D.) was used. The corer was transported to laboratory in sealed polyethylene bags and stored at 4°C for 2 h before cultivation experiments, preparations for total bacterial counts, sporulation titer and morphological characterization of bacteria were initiated.
| Table 1: | Soil types, geographical coordinates and percentage moisture content of collected samples |
 | |
For dry-weight determinations, 1 g of freshly sieved soil was accurately weighed and then dried at 105°C for 3 days. The samples were then reweighed after cooling to room temperature. The factor for conversion of fresh weight to dry weight of soil was calculated and all results were expressed per gram (dry weight) of soil.
Cell extraction: The soil samples were sieved through (1.7 mm aperture size) to remove large particles, roots, stones and vegetation, which was found in some samples. One gram of soil was transferred to plastic tube and suspended with a sodium cholate solution (9 mL, 0.1%) to disrupt the adhesive polymers the causes the cells to bind to each other and to the soil surface as well (Macdonald, 1986). Then, a mixture of chelating sorbent (0.2 g chelating resin) and polyethylene glycol (0.25 g, PEG 6000) was added followed by centrifuging for 1 h (220 rpm, temp. 37°C). After 1 min sedimentation of the mixture, the supernatant was transformed to another test tube.
Cultivation experiments and plate count: Serial liquid dilution culture technique was used in this study to improve culturability and isolation of bacteria in anoxic soil systems (Chin et al., 1999), 1 mL aliquot of soil supernatant was suspended with 9 mL Phosphate Buffered Saline (PBS) for determination of bacterial number. This solution was decimally diluted (10-2 to 10-6) and 100 μL aliquots of each dilution were spread on dilute nutrient broth, solidified with agar, DNBA (Janssen et al., 2002), containing cycloheximide (end concentration 100 μg mL-1) to inhibit fungal growth and incubated at 25°C for 3 days. Bacterial colony counts were determined as Colony-Forming Units (CFU) per gram dry weight. For determination of bacterial spore titer, 1 mL aliquots of each dilution was heated at 80°C for 30 min and then plated on nutrient agar containing cycloheximide.
Physicochemical identification of bacteria: Gram stain technique was used to get general information about the distribution of Gram positive and Gram negative bacteria in the different soil types found in Gaza. The 1:104 dilution of all soil types cultured on 1/10 diluted nutrient agar were representative according to morphological characteristics such as colony appearance, size and color to the lower dilution series and therefore were preserved. Because of the limited research facilities, the genus identification of bacterial isolates was done according to morphological characteristics and some biochemical tests.
Colonies that developed on the plates were subculture repeatedly to obtain pure single colony, which was maintained on agar slants for further characterization and identification. The identification of isolated bacteria was performed by Gram stain, morphological characteristics and ability to form endospores, followed by biochemical genera identification tests such as starch hydrolysis, catalase, oxidase, indol and glucose fermentation.
Statistical analysis: The average values of the number of CFU g-1 dry weight of soil and standard deviation were statistically analyzed by SPSS 17 program.
Total bacterial count and sporulation titer in the different soil types: Bacterial diversity was detected in all soil types of Gaza Strip. The number of bacteria in 1 g dry soil was determined as colony-forming units (CFU g-1 dry soil). Three collects were taken as samples of each type of soil from different areas of the Gaza Strip with given GPS coordinates and specified below, where the total bacterial count and sporulation each collect was determined.
Collects from loessial sandy soil, which has a flat/rolling interdune areas as land form, from Deir Al Balah at X, Y coordinates (X: 88115, Y: 90196), demonstrated a total bacterial count of 6.91±106 CFU g-1 and sporulation titer of 5.1x105 CFU g-1. About 7.38% of the total bacterial population was found as spores, whereas collects of sandy loess soil, which has flat/rolling plains or depressions as land form, from northern part of Deir Al Balah at X, Y coordinates of (X: 91233, Y: 95414) showed a lower-total bacterial count of 1.78x106 CFU g-1 dried sandy loess and a lower sporulation titer of 2.1x105 CFU g-1. About 11.8% of the total bacterial population was found as spores. For collects of sandy loess over loess, which has Gently rolling plains as land form, from Khan Yunis at X, Y coordinates of (X: 85969, Y: 84480) the total bacterial count was almost half of that of loessial sandy soil collects and double that of sandy loess soil collects (3.13x106 CFU g-1). However, The sporulation titer was lower than both (1.02x105 CFU g-1). The ability of total bacterial population in sandy loess over loess soil to produce spores was about 3.26% .
Moreover, in sandy regosols soil, which has a Active steep dunes and Calcareous ridges as land form, the collects from the southern part of Deir Al Balah at X, Y coordinates of (X: 85025, Y: 90913) showed a total bacterial count close to that of loessial sandy soil collects (5.6x105 CFU g-1) and the sporulation titer was the lowest (8.2x104 CFU g-1). About 14.64% of the bacteria found in sandy regosols soil was exist as spores. In collects of loess soil, which compose of rolling plains as land form, from Gaza city (X: 96981, Y: 100604), the total bacterial count dropped to 2.82x106 CFU g-1 and the sporulation titer increased to 4.9x105 CFU g-1. 17.37% of the bacterial population in loess soil was found as spores.
Samples of dark brown/silty clay, which has depressions and slopes as land form, were collected from Wadi Gaza at X, Y coordinates (X: 94912, Y: 98490). The total bacterial count was the highest (9.36x106 CFU g-1) and the sporulation titer was 2.3x105 CFU g-1. The lowest ability of bacterial population to form spores was detect in loess soil, about 2.46%. The total bacterial count and sporulation titer of the different soil types in the Gaza Strip are stated in Table 2 and the percentage of sporulation titer in all soil types is listed in Table 3. The mean of bacterial count is all soil types in the Gaza Strip estimated in this study was 4.09x106 CFU g-1 which is about one hundredth and one thousandth the values estimated in previous studies (Janssen et al., 2002; Schoenborn et al., 2004).
From the pervious results we can conclude that the bacterial population was found in all soil types in Gaza Strip. The bacterial communities were able to form spores in all soil types, however in different percentages.
Contribution of G+ and G- bacteria in the different soil types: About 80 bacterial isolates were randomly selected from the different soil types and sub-cultured in order to get pure bacterial colony and then preserved for staining and characterization via biochemical tests.
| Table 2: | Total bacterial count and sporulation titer in different soil types |
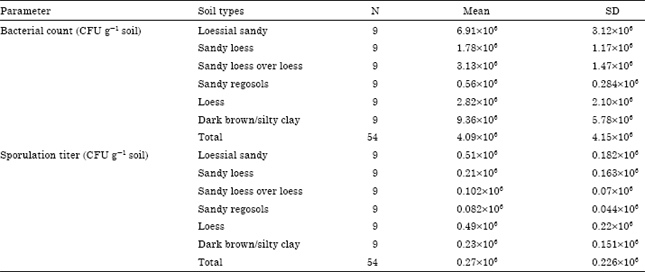 | |
| N: Number of soil samples tested for estimation of total bacterial count and sporulation titer | |
| Table 3: | Percentages of sporulation in the different soil types |
 | |
| Table 4: | Number and percentage of G+ and G- bacteria in the different soil types |
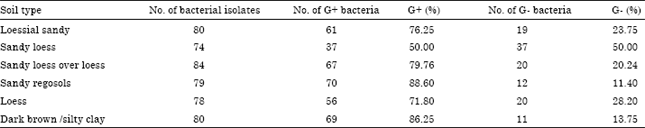 | |
| G+: Gram positive bacteria, G-: Gram negative bacteria | |
The distribution of Gram positive bacteria (G+) in all soil types was more than those of G negative bacteria (G-), except for sandy loessial soil, where the distribution of both G+ and G- bacteria was equal (Table 4). In loessial sandy soil, it was found that the percentages of Gram positive bacteria was about three times higher than Gram negative bacteria 76.25 and 23.75%, respectively. Similar distribution, with slight variation, of Gram positive bacteria was observed in sandy loess over loess and loess soil. The percentage distribution of Gram positive bacteria in sandy loess over loess soil was 79.97, while in loess soil was 71.8. Sandy regosols and dark brown/silty clay soil showed almost the same content of Gram positive bacteria.
| Table 5: | Identification of Gram positive bacteria |
 | |
| +: Positive test result, -: Negative test result, +w: Weak positive result | |
| Table 6: | Identification of gram negative bacteria |
 | |
| +: Positive test result, -: Negative test result, +w: Weak positive result | |
The percentage distribution of Gram positive bacteria in sandy regosols was 88.6 of total bacterial isolates, while the percentage distribution of Gram positive bacteria in dark brown/silty clay soil reached 86.25. Only in sandy loess soil were the percentage distribution of Gram positive bacteria equal to that of Gram negative bacteria.
The most abundant shape of stained bacteria an all soil types was the bacilli. Coccobacilli and filamentous bacteria were more common in sandy loess over loess soil.
Biochemical identification of bacteria: Recently, the identification of soil bacteria is based on molecular 16S rRNA nucleotide sequence analysis. The molecular genetic methods used for the analysis of soil bacterial diversity showed a large number of members of the phylum Acidobacteria, Actinobacteria, Bacteroidetes, “Deinococcus-Thermus”, Firmicutes, Gemmatimonadetes, Planctomycetes, Proteobacteria, Verrucomicrobia (Joseph et al., 2003).
Because of the limited scientific and research resources and facilities in Gaza Strip, the genus identification of soil bacteria in this study was done by means of biochemical tests described in the identification flow chart of Bergey’s manual (Holt et al., 1994), such physicobiochemical characterization and identification of the bacterial isolates were used in various studies (Qazi et al., 2009).
A total of 100 bacterial isolates were randomly selected for identification, 60 bacterial isolates were Gram positive, while 40 were Gram negative bacteria. Identification of Gram positive and Gram negative bacteria were done by means of some biochemical tests. The results of bacterial identification and their percentage distribution in the different soil types were shown in Table 5 and 6. The Gram positive bacterial isolates were grouped according their shape into bacilli, Cocci and Filamentous. The identification of bacterial genera using biochemical tests like endospores formation, oxidase, catalase, glucose fermentation, indol and starch hydrolysis resulted in expected genera as Bacillus, Micrococcus, Staphylococcus and Streptomyces (Table 5).
The Gram negative bacterial isolates were grouped according their shape into bacilli and coccobacilli. The identification of bacterial genera using biochemical tests resulted expected genera as Escherichia, Pseudomonas, Aeromonas, Xanthomonas, Acinetobacter and Moraxella (Table 6).
Physiological identification of bacteria (phenotype) have shown that various genera are present in the different soil types.
The contribution of bacterial population in the soil of Gaza Strip was the main aim of this research. Studying different samples in different areas of Gaza showed that the total bacterial count range is 1-9x106 CFU g-1 as the maxima count was detected in the Dark brown/Silty clay soil type. It should be noted that this type of soil was collected from Wadi Gaza, which is considered a heavily polluted region because untreated sewage, agricultural and industrial effluents are discharged there (Environmental Assessment of the Gaza Strip, following the escalation of hostilities in December 2008-January 2009 (UNEP, 2009; Abed et al., 2002). The minimum bacterial count was detected in the sandy regosols soil that covers the costal area of Gaza Strip.
The total bacterial count estimated in this study was lower than bacterial count described in other studies. Chin et al. (1999) measured a total bacterial count up to 2.5x108 cells per gram dry weight. The first study employed diluted nutrient agar method yielded a viable bacteria counts of 1.96x106 to 5.56x107 using most probable number method and 2.39-10.01x107 CFU g-1 using plate count experiments. (Janssen et al., 2002). Other study applying fluorescence microscopy after staining with 4,6-diamidino-2-phenylindole (DAPI) showed a bacterial count of 1.2-1.6x109 cells per gram dry soil (Schoenborn et al., 2004).
The ability of bacteria to resist environmental stresses by formation of endospores was characteristics an all types of soil, however in different percentages. The distribution of Gram positive and Gram negative bacteria was investigated and varying among the different types of soil. Gram positive bacteria was generally dominating the Gram negative bacteria which is probably due to the ability of Gram positive bacteria to form endospores and develop other stress response answers (Hecker et al., 2007). The dominance of Gram positive bacteria over Gram negative in soil was also observed in other studies (Da Silva and Nahas, 2002). Consequently, it survives environmental changes better than its counterpart Gram negative bacteria (Brigulla et al., 2003; Hecker and Volker, 2001; Joseph et al., 2003).
A morphological and biochemical identification of bacterial isolates were done because of the limited scientific and research resources and facilities in Gaza Strip. The genus identification of soil bacteria via morphological and biochemical test was used and employed in many studies (Da Silva and Nahas, 2002; Phillips et al., 2012). The identified bacterial isolates in this research project belonged to families Bacillaceae, Micrococcaceae, Staphylococcaceae, Streptomycetaceae, Aeromonadaceae, Pseudomonadaceae, Moraxellaceae, Enterobacteriaceae and Xanthomonadaceae. The most recently research projects depend on genotypic and molecular identification of bacterial isolates in their natural habitat such as air, water and soil. For example, Soil DNA extraction and sequencing PCR for bacterial tag-encoded FLX amplicon pyrosequencing of the 16S rDNA gene can be used to evaluate bacterial diversity from soil (Acosta-Martinez et al., 2010). The use of molecular biology methods in bacterial identification allows phyla level identification of bacterial diversity.
This study is considered as a promising start for identification and distribution of various bacterial families in the different soil types existed in the Gaza Strip. For more specific and genotypic identification of bacterial isolates in soil habitat of Gaza Strip, a the molecular biological methods should applied in future studies.