Research Article
Diversity of Coral Eunicea fusca Associated Bacteria Using Culture Dependent Techniques
Department of Chemistry, University of Prince Edward Island, 550 University Avenue, C1A4P3, PEI, Canada
Coral reefs are the most biodiverse of all marine ecosystems; however, very little is known about the prokaryotic diversity in these systems. Our understanding of the role of bacteria in coral reef ecosystems is still evolving. Bacteria are known to be abundant and active in seawater around corals, in coral tissues and within their surface micro layer. Many researchers as Santavy (1995), Kushmaro et al. (1996), Koh (1997), Kushmaro et al. (1997) Gast et al. (1998), Gili and Coma (1998), Lyons et al. (1998) and Toren et al. (1998) have examined the interactions between corals and microbes. These studies have shown that there is a dynamic microbiota living on the surface and possibly within the tissue of corals and in the surrounding reef waters. However, it is still not known whether microbes play specific roles in coral biology or if the observed associations are merely opportunistic interactions of the coral animal with water-column bacteria. Early studies using culturing methods demonstrated the importanceof coral-associated bacteria in coral nutrition and inresponse to stress. During the last three decades, most of the studies on coral-associated bacteria have linked the presence of bacteria in corals to several diseases (Ben-Haim et al., 2003; Sutherland et al., 2004). It was shown that the bacterialcommunity changes when corals are bleached (Ritchie et al., 1994) or exhibited white-band disease or exposed to an environmental stress (Ritchie and Smith 1995). Our primary goal in this study was to characterize the culturable natural bacterial community associated with Enicea fusca coral using culture-based methods in comparison with that of the coral surrounding water column.
Coral Samples Collection and Preparation
Eunicea fusca was collected in sterile bags at Hillsboro ledge in Florida, August 2006 by SCUBA diving at depth of approximate 30 feet. Samples were kept fresh in seawater in and processed upon arrival to the laboratory. Control seawater was collected in the area surrounding the coral. In this case, the water sample was collected in sterile bottle that was opened adjacent to the coral in situ.
Isolation and Enumeration of Coral Associated Bacteria
The collected corals (Three samples) were washed under aseptic condition using filtered seawater (FSW) to remove the loosely attached bacteria, weighed, minced with razor blade and homogenized using a sterile blender. The homogenate was filtered through sterile cheesecloth to remove the large particles and debris. The homogenate was serially diluted up to 10-5 with FSW. A 100 μL aliquot of each dilution was plated in triplicates on different agar media. Nutrient rich and. nutrient poor media, Marine agar and 1/10th strength Marine agar media, respectively were used at different pH 5, 7 and 9 (DifcoTM, Sparks, MD). The plates were incubated at 30°C for 2-3 weeks. Bacterial isolation from the collected seawater was done at the same conditions. Representatives of each colony morphotype were serially streak-plated onto fresh media to obtain pure cultures and stored in 20% glycerol at -80°C.
Box-PCR Genomic Fingerprint
The bacterial isolates were grown overnight in 5 mL Marine broth. Total DNA was extracted with the Ultra Clean Microbial DNA Isolation Kit (Mo Bio Laboratories Inc., Carlsbad, CA) according to the manufacturer`s procedure. BOX PCR was carried out using BOX-AIR primer (5`-CTACGGCAAGGCGACGCTGACG-3`) (Nick et al., 1999). Cycling conditions included an initial denaturation step of 95°C for 2 min followed by 35 cycles of 94°C for 3 sec, 92°C for 30 sec, 50°C for 1 min, 65°C for 8 min. A final extension step at 65°C for 8 min was added. The amplified fragments were separated by electrophoresis on a 1.5% agarose gel containing ethidium bromide and ran at 60 V for 3.5 h.
Bacterial Identification by 16S rRNA Gene Sequence Analysis
The bacterial strains were identified by 16S rRNA gene sequence analysis. The bacterial isolates were grown overnight in 5 mL Marine Broth. Total DNA was extracted with the Ultra Clean Microbial DNA Isolation Kit (Mo Bio Laboratories Inc., Carlsbad, CA) according to the manufacturer`s procedure. Eubacterial-specific primers, forward primer 16F27 (5´- AGA GTT TGA TCC TGG CTC AG-3´) and. reverse primer. 16R1525 (5´- AAG GAG GTG ATC CAG CCG CA-3´) derived from E. coli 16S-rDNA sequence (Lane, 1991) were used to amplify 16S rDNA gene. The reaction mixture of 50 μL contained at least 100 ng of genomic DNA (in 10 mM Tris-HCl, pH 8), 0.2 μM of each primer and PCR Supermix High fidelity (Taq and Go, Promega, CA). PCR fragments were purified using a QIAquick Gel Extraction Kit (Qiagen, Valencia, CA) and sequenced on an ABI 377 automated sequencer using the PRISM Ready Reaction Kit (Applied Bio Systems, Foster City, CA). Sequence data were analyzed by comparison with 16S rRNA genes in the GenBank database. The nearest relatives of each organism were obtained by BLAST searches (Altschul et al., 1997).
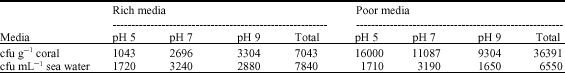
The total colony forming unit (cfu) of the coral associated bacteria in different media is shown in Table 1. The total CFU of the coral associated bacteria was found to be five fold more in nutrient poor media than in nutrient rich media. In nutrient rich media, the ration of the cfu at different pH was 14.6, 37.8. and. 47.6 at pH 5, 7, 9, respectively. In nutrient poor media, the ratio of the total cfu at different pH was 44, 30.5 and 25.5 at pH 5, 7 and 9, respectively. The total cfu g-1 coral was about 4.8 fold more than cfu mL-1 of the seawater in nutrient poor media. In case of seawater, in nutrient rich media, the ration of the cfu at different pH was 22, 41.3 and 36.7% at pH 5, 7 and 9, respectively. In nutrient poor media the ratio of the total cfu at different pH was 26.1, 48.7 and 25.2% at pH 5, 7 and 9, respectively.
| Table 1: | Colony forming unit count (cfu) of the Eunicea fusca associated bacteria and seawater on nutrient rich and nutrient poor media at different pH values |
 | |
 | |
| Fig. 1: | DNA fingerprint of the Eunicea fusca associated bacteria generated with primers BOX-A IR. Lane 1 is a standard DNA marker of 1 Kb ladder |
Box-PCR
All the isolated, were subjected to Box-PCR analysis to provide indication of the unique fingerprints (Fig. 1). A total of 56 unique patterns resulted from the BOX-PCR. All the strains with unique patterns were identified with sequence analysis of the 16S rDNA.
Phylogenetic Analysis of the Isolates
Isolates with unique patterns from the BOX-PCR were subjected to 16S rDNA sequence analysis. The 16S rDNA of 29 strains, isolated from the Eunicea fusca and 29 strains, isolated from the seawater, were sequenced. The resulted sequences were compared to the sequences of known bacteria in the GenBank. The sequences of the isolates possess 96-99% similarity compared to sequences of previously known bacteria in the GenBank (Table 2). The coral associated bacteria were dominated by Alphaproteobacteria with 65.5% of the isolates falling within this group. This was followed by 20.7% of Gammaproteobacteria, Firmicutes 6.9%, Betaproteobacteria 3.4 and 3.4% of Cytophaga-Flexibacter Bacteroids (CFB) (Table 4). Gammaproteobacteria has been reported to be the major bacterial group in many other soft coral using culturing technique (Ritchie and Smith, 1995; Rohwer et al., 2001). By comparing the composition of culturable bacterial isolates of the coral Fungia scutaria to the present study it is found that while F. scutaria isolates were dominated by an almost equal distribution of Alphproteobacteria and Gammaprotebacteria (Lampert et al., 2006), E. fusca was dominated mainly by Alphaproteoacteria.
| Table 2: | Characterization of culturable bacteria associated with E. fusca isolated in different media. |
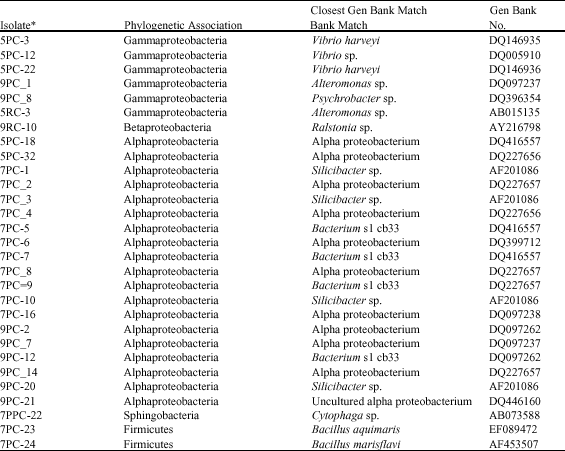 | |
| *The acronyms given to the isolates read as a follow: e.g. PC: Nutrient poor (coral), RC: Nutrient rich media (coral). The first digit is referring to the pH value | |
The seawater was dominated with Alphaproteobacteria with 82% of the isolates falling within this group. This was followed by 9% of Gammaproteobacteria and 6% of Betaproteobacteria and 3% was falling within the division Cytophaga-Flexibacter Bacteroids (Table 4).
Differences were observed in the types of culturable bacteria between seawater and coral. The three groups α, β and γ proteobacteria were represented in both samples but with different species. The CFB and Firmicutes groups were only isolated from the coral samples. The coral associated bacteria are falling within nine genera with different species, Vibrio sp., Alteromonas sp., Psychrobacter sp., Ralstonia sp., Silicibacter sp., Bacterium s1 cb33, Bacillus sp., Cytophaga sp. and uncultured alpha proteobacterium.
Vibrio sp. has been isolated from other corals Oculina patagonica and Montastraea franksi (Koren and Rosenberg, 2006; Rohwer et al., 2001). Alteromonas sp. and Silicibacter sp. have been isolated from the Red Sea coral Fungia scutaria (Lampert et al., 2006).
Karen and Rosenberg (2006) found the Bacterium slcb33 associated with mucus and tissues from the coral Oculina patagonica in summer and winter of the cost Israel. This suggests a global distribution of Bacterium slcb33 as well as a possible coral specific association.
| Table 3: | Characterization of culturable bacteria isolated from sea water samples at different pH and nutrient media |
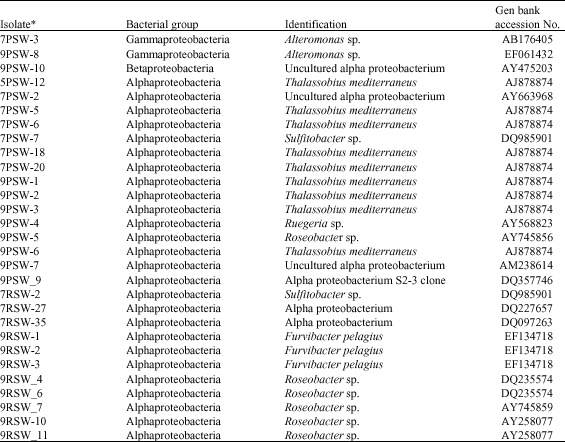 | |
| *The acronyms given to the isolates read as a follow: e.g., PSW: Nutrient poor media (seawater). PSW: Nutrient rich nutrient media (seawater) (RC). The first digit is referring to the pH values | |
| Table 4: | Different bacterial groups in coral tissues and seawater sample |
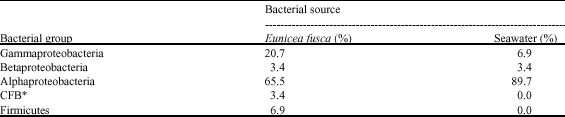 | |
| *Cytophaga-flexibacter bacteroids | |
The seawater strains, 29 isolate, were found to be more diverse that affiliated to nine genera with different species, Alteromonas sp., Ralstonia sp., Uncultured alpha proteobacterium, Thalassobius, Sulfitobacter sp., Ruegeria sp., Roseobacter sp., Furvibacter, Rhodobacter sp. (Table 3).
Most of the strains isolated from the E. fusca tissues were absent in the strains isolated from the seawater, only one strain, Alteromonas sp., has been found in both the coral and the surrounding seawater sample which provides evidence of specific bacterial association with the coral indicated a specific bacterial association with the coral tissues. Similar bacterial specific association with the coral has been reported in coral Montastraea franksi (Rohwer et al., 2001).
Analysis of The Nutrient Rich and Nutrient Poor Media Isolates
The total bacterial count (cfu) of the coral associated bacteria was found to be five fold more in nutrient poor media than in nutrient rich media. In nutrient rich media, while it was about 1.2 fold more in rich media in case of seawater. The results shown in Table 5 indicated that the nutrient poor media showed more bacterial groups diversity than nutrient rich media while both were dominated by Alphaproteobacteria. Furthermore, the number of the different strains was about two fold higher in nutrient poor media.
| Table 5: | Different bacterial groups isolated in nutrient poor and nutrient rich media |
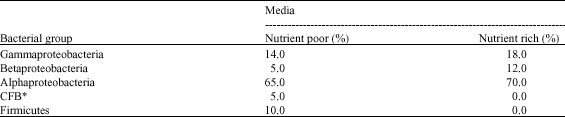 | |
*Cytophaga-flexibacter bacteroids | |
In addition to media types, the pH of the media was another important parameter. The results of bacterial isolation from the coral tissues and seawater, suggested that acidic pH was not the most optimal for bacterial growth as evidence by the many fewer strains obtained at pH 5 in comparison with pH 7 and. 9. Overall, using slightly alkaline oligotrophic conditions, nutrient, increased the number and diversity of the culturable bacteria isolated from seawater and Eunicea fusca.
The term the great plate count anomaly was coined by Staleyand Konopka in 1985 (Staley and Konopka, 1985) to describe the difference in ordersof magnitude between the numbers of cells from natural environmentsthat form colonies on agar media and the numbers countable bymicroscopic examination. Marine ecosystems are a well-studiedexample of this phenomenon: only 0.01 to 0.1% of oceanic marinebacterial cells produce colonies by standard plating techniques(Stephanie et al., 2002). There are numerous explanations for this anomaly. Forexample, species that would otherwise be culturable may failto grow because their growth state in nature, such as dormancy,prevents adjustment to conditions found in the medium used forthe plate counts (Deming and Baross, 2000). This hypothesis does not explain thesubstantial discrepancy between 16S rRNA genes recovered fromseawater directly by cloning and those of the readily culturedmarine taxa (Stephanie et al., 2002). Another explanation for the great platecount anomaly is that many of the microbial species that dominatein natural settings are not adapted for growth in media containinghigh concentrations of complex organic carbon. Many microorganismsmay need oligotrophic or other fastidious conditions to be successfullycultured. There are many examples of microbial strains thatare common in nature, but can only be cultivated by specializedtechniques (Button et al., 1998; Partensky et al., 1999; Vancanneyt et al., 2001; Wirsen et al., 2002; Stephanie et al., 2002).
In summary, Eunicea fusca associated bacteria have been isolated and characterized. Comparison of the coral associated bacterial population with the bacterial community in the coral surrounding seawater provided evidence of specific bacterial association with the coral. Furthermore, investigation of different isolation conditions demonstrated that slightly alkaline oligotrophic conditions increased the culturability and diversity of E. fusca associated bacteria.
The author would like to sincerely thank Mrs. Amira Abdelmaksoud for her valuable help in writing and revision of this manuscript