Research Article
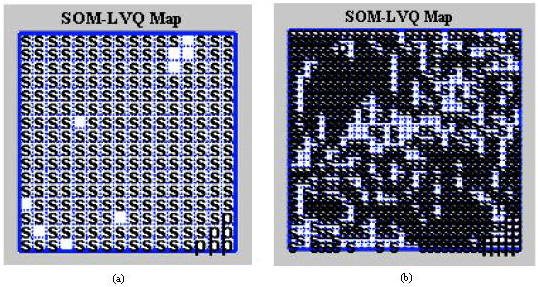
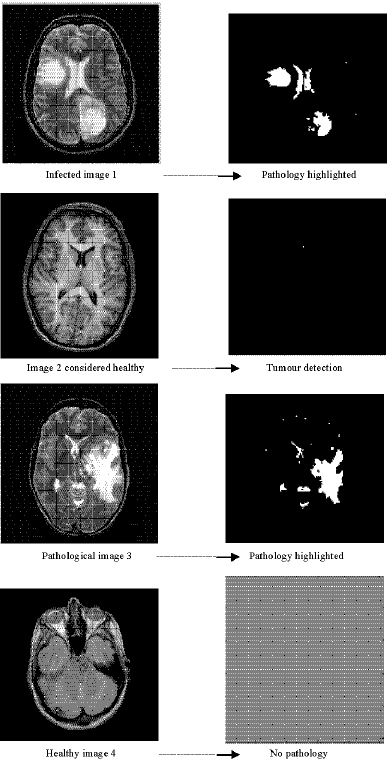
Classification of the Medical Images by the Kohonen Network SOM and LVQ
Research Laboratory in Intelligent Systems, Department of Electronics, Algeria
N. Berrached
Research Laboratory in Intelligent Systems, Department of Electronics, Algeria
N. Kharchouche
Research Laboratory in Intelligent Systems, Department of Electronics, Algeria
Y. Ghellemallah
Research Laboratory in Intelligent Systems, Department of Electronics, Algeria
M. Mansour
Laboratory of LSI-REC, University of Sciences and Technology, Mohamed Boudiaf,USTOran 31000, BP 1505, Algeria
H. Mouhadjer
Laboratory of LSI-REC, University of Sciences and Technology, Mohamed Boudiaf,USTOran 31000, BP 1505, Algeria










ankit patel Reply
sir i want python code of image classification using kohonen self organizing maps and report