Research Article
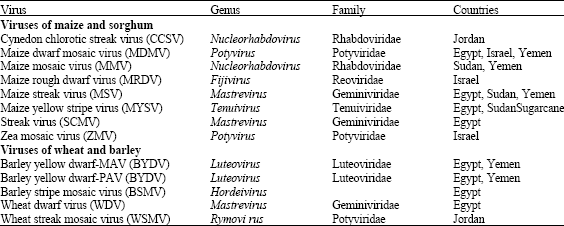
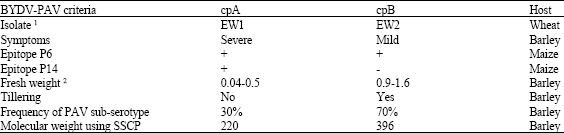
Diagnosis of Cereal Viruses in the Middle East
Plant Virus and Phytoplasma Research Section, Plant Pathology Research Institute, Agriculture Research Center, 12619 Giza, Egypt
G. Anfoka
Department of Biotechnology, Faculty of Agricultural Technology,Al-Balqa` Applied University, Al-Salt 19117, Jordan
M. Zeidan
Molecular Genetics and Virology, Al-Qassmi Research Center,Baqa El-Gharbia 30100, Israel
H. Czosnek
Institute of Plant Sciences and Genetics in Agriculture,Robert H. Smith Faculty of Agriculture, Food and Environment,The Hebrew University of Jerusalem, Rehovot 76100, Israel