ABSTRACT
The variation of response of genotypes to environments has made multi-environment study a necessity in plant breeding. Twelve Okra genotypes were tested over three environments (locations) Ibadan, Abeokuta and Ayetoro to study the effects of Genotype by Environment Interaction (GEI) on seed yield. Seed yield data were subjected to Yield stability index (YSi) and Genotype plus GenotypexEnvironment Interaction (GGE) biplot in order to select desirable genotypes for yield and stability and to compare the relative discriminatory ability of the two techniques. The GGE biplot and YSi techniques jointly identified NHGB/09/009A as the genotype that best combined high yield and stability; and FUNAAB-11-4 and FUNAAB-11-6 as having good potential in this regard. The YSi in addition, also selected FUNAAB-11-8, FUNAAB-11-3 and LD-88. The three genotypes solely selected by YSi did not consistently produce above average yield across the three locations indicating that YSi was less reliable in the selection of genotypes for high yield and stability. Also, the mean yield of the selections by the two techniques were statistically similar indicating that the selection of high yielding but unstable genotypes would not be of any practical advantage in a selection programme. The GGE biplot identified two mega-environments viz Ibadan and Ayetoro where FUNAAB-11-8 was outstanding and Abeokuta where FUNAAB-11-3 was the best. It identified Ayetoro as the best location for the selection of superior genotypes. Okra genotypes NHGB/09/009A, FUNAAB-11-4 and FUNAAB-11-6 would therefore be desirable for cultivation across the three environments.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ijpbg.2013.146.158
URL: https://scialert.net/abstract/?doi=ijpbg.2013.146.158
INTRODUCTION
Okra (Abelmoschus esculentus L. Moench) formerly known as Hibiscus esculentus belongs to Malvaceae (Jute) family and a close relative of one of the most important fibre crops, the Kenaf. It is a short duration, day neutral and autogamous vegetable crop grown for its tender fruits (Ariyo, 1993; Osekita et al., 2000). Okra plant is of high economic importance to man and its use ranges from biochemical, food and industrial purposes (Oyelade et al., 2003; Kumar et al., 2010).
All of these uses not withstanding the decline in Okra yield in Nigeria is well documented (Kumar et al., 2010; Alake et al., 2012; FAOSTAT, 2011), a record of Okra yield in Nigeria from the year 2000 is presented in Table 1. This was as a result of the minor crop status conferred on the crop which ensured that little attention has been given to its breeding (Abdelmageed, 2010; Kumar et al., 2010). This has restricted growers to the use of old varieties that need to be replaced by new (superior) genotypes.
| Table 1: | Trends in Okra production from year 2000-2010 in Nigeria |
 | |
| FAOSTAT, 2012 | © FAO Statistics division 2012 | 24 August 2012 | |
The growing awareness of the importance of this vegetable implies that more genetic study need be done to enhance the development of superior genotypes that would yield more. Stability studies are important in the process of developing improved varieties in any crop. GenotypexEnvironment Interaction (GEI) affects breeding programs by its influence on genotype performance and therefore often complicates the evaluation and selection of superior genotypes. The knowledge of GEI processes and genotype adaptation can be used to reduce this constraint. Yan and Kang (2003) advanced that to detect GEI statistically at least two distinct genotypes must be evaluated in at least two distinct environments. GEI analysis is particularly important when the performance of genotypes changes in different environments (crossover GEI). The consequence of GEI is the differential response of genotypes to varied environmental conditions. Ceccarelli (1996) opined that genotypes that had good performance under improved environmental conditions could be worse off when the environment is poor. According to Becker and Leon (1988) a desirable cultivar/genotype must show high and stable yield when evaluated in varying growing conditions. It is therefore of great necessity that promising Okra genotypes be subjected to different environments (location) so that only genotypes that combine relative stability and high yields are selected for further improvement. GEI is of high importance to plant breeders and consequently different models have been used in its study. These include the simple classical analysis of variance by Fisher (1918) and the Joint Regression Analysis first proposed by Yates and Cochran (1938) and later modified by Finlay and Wilkinson (1963) and Eberhart and Russell (1966). Others are Ecovalence by Wricke (1962), stability variance proposed by Shukla (1972), Kang Rank Sum (KRS) by Kang (1988). KRS was later modified by Kang (1991) to Kang Modified Rank Sum (KMRS). The more popular techniques currently in use are the Additive Main Effect and Multiplicative Interaction (AMMI) by Gauch and Zobel (1988), Yield Stability index by Kang and Magari (1995), Genotype+GenotypexEnvironment (GGE) biplot by Yan (2001).The GGE biplot is the most sophisticated of the techniques that utilize biplot (Yan et al., 2007) while YSi ranks top amongst techniques that integrate yield ranks with stability rating (Waldron et al., 2002). This study in addition to identifying stable and high yielding Okra genotypes would compare the two GEI analytical techniques with a view to identifying which is more efficient for this purpose.
MATERIALS AND METHODS
Field location: The study was conducted at the Federal University of Agriculture Abeokuta, Olabisi Onabanjo University Ago-Iwoye (Ayetoro Campus) and the National Centre for Genetic Resources and Biotechnology (NACGRAB), Ibadan. Abeokuta is on 7°38’N, 3°88’E with a humid tropical climate and mean annual rainfall of 1200 mm. Ayetoro is on 7°12’N, 3°30’E with a sub-humid climate and average annual rainfall of 1000 mm. Ibadan the third location is on 7°37’N, 3°89’E with a tropical climate and mean annual rainfall of 1350 mm. The agro-meteorological data of the three locations are presented in Table 2.
Source of materials and experimental design: Twelve Okra genotypes were sourced from Federal University of Agriculture Abeokuta (FUNAAB), National Centre for Genetic Resources and Biotechnology (NACGRAB), Ibadan and University of Ibadan (UI), Ibadan as stated in Table 3. The collections were evaluated in single-row plots at the three locations. In each location, soil preparation was done by ploughing, harrowing and levelling and the experimental design was the Randomized Complete Block with three replicates. A block, 1.5 m apart, consisted of 12 rows of all the genotypes and each row was 4.5 m long. The rows were 0.75 m apart while plant-to-plant distance in each row was 0.45 m.
| Table 2: | Source and characteristics of the Okra genotypes evaluated |
 | |
| Table 3: | Agrometeorological data of the test locations for the duration of the experiment |
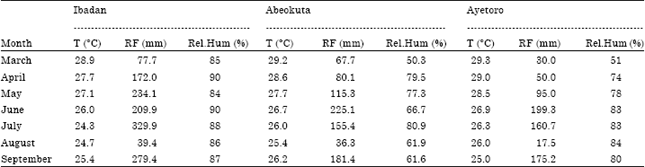 | |
| Department of water resources management and agrometeorology, FUNAAB: National Cereals Research Institute (NCRI), Ibadan: Internship weather station, CASOOU: Ayetoro, T (°C): Temperature in degree celsius, RF (mm): Rainfall in millimetres, Rel.Hum (%): Percentage relative humidity | |
Cultural practices: Two seeds were sown per hole and later thinned to one plant per stand at 2 weeks after sowing. Weeds were controlled manually as required while insect pests were controlled using Lambda-cyhalothrin+Dimethoate at 3.5 mL to 1 L of water. NPK fertilizer 15:15:15 was applied at the rate of 150 kg ha-1 at 2 weeks after sowing and at flowering. Data on seed yield were collected from eight inner plants in each row. Harvesting was done when the pods had dried up after which they were threshed to collect the seeds.
Data collection and analysis: Seed yield was determined as the average of the bulked weight of seeds of eight inner competitive plants.
Plot means for seed yield were subjected to Analysis of Variance (ANOVA) and means were separated using the Duncan Multiple Range Test of SAS Institute (2002). The yield data were then subjected to YSi statistic of Kang and Magari (1995) and Genotype plus GenotypexEnvironment Interaction (GGE) according to Yan (2001), Yan and Kang (2003) and Yan et al. (2007) using the edition to investigate the response of the genotypes to different environments and to select the best environment and genotype. The GGE biplot analysis was used to decompose the GEI by its visuals, mega-environments and their best performers were identified. Mean performance and stability of genotypes were determined and the discriminating ability and representativeness of the test environment (location evaluation) was revealed (Yan, 2001).
The Yield stability index (YSi) is a GenotypexEnvironment analytical technique designed by Kang and Magari (1995) for simultaneous selection for desirable genotypes based on yield and stability. It is an advancement of the Kang Rank Sum (KRS) by Kang (1988) and the Kang Modified Rank Sum (KMRS) by Kang (1991). It is the summation of the adjusted yield rank and the stability variance of Shukla (1972) to determine high yielding and stable genotypes.
RESULTS
The mean seed yield of the genotypes is presented in Table 4. Seed yield ranged from 6.49 for F3 45x72-11(V11) to 22.39 for FUNAAB-11-8 (V2) across the locations. At Ibadan FUNAAB-11-6 (V3), FUNAAB-11-4 (V5), NHGB/09/009A (V7), UI 53-139 (V9), NH47-4 (V12) all yielded above average with the mean yield of 9.76, 9.78, 13.80, 10.98 and 12.58, respectively.
| Table 4: | Mean seed yield (g plant-1) of 12 Okra genotypes evaluated at three locations |
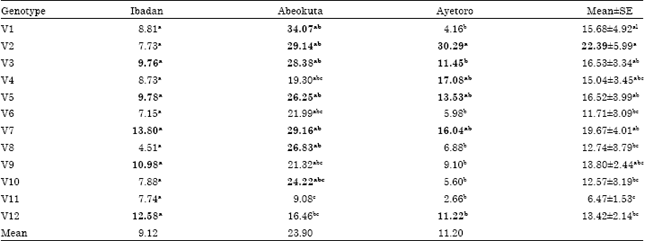 | |
| *Means with similar alphabet on the same column are not significantly different from one another at 5% probability level using, DMRT: *Means in bold indicate above average performance at each location, Mean±SD: Mean plus or minus standard error | |
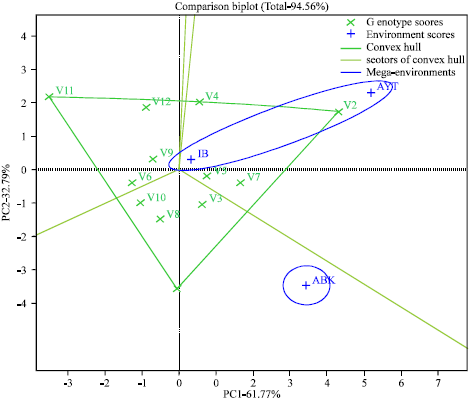 | |
| Fig. 1: | GGE biplot displaying the mega-environment and the “which-won-where” pattren Transform = 0, Scaling = 0, Centering = 2, SVP = 1 |
V7 was the best performer while UI 4-30 (V8) had the lowest yield. Similarly at Abeokuta FUNAAB-11-3 (V1), FUNAAB-11-8 (V2), FUNAAB-11-6 (V3), FUNAAB-11-4 (V5), NHGB/09/009A (V7), UI 4-30 (V8) and F3 313x104 (V10) performed above average with the mean yield of 34.07, 29.14, 28.38, 26.25, 29.16, 26.83 and 24.22, respectively. FUNAAB-11-3 (V1) was thus the best performer while F3 45x72-11(V11) had the lowest yield. In the case of Ayetoro FUNAAB-11-8 (V2), FUNAAB-11-6 (V3), LD-88(V4), FUNAAB-11-4 (V5), NHGB/09/009A (V7) and NH47-4 (V12) also yielded above average with mean yield of 30.29, 11.45, 17.08, 13.53, 16.04 and 16.46, respectively. FUNAAB-11-8 (V2) yielded than others while F3 45x72-11(V11) had the lowest yield. Among the locations, Abeokuta recorded the highest mean of 23.9 g plant followed by Ayetoro (11.2) and then Ibadan (9.12).
Figure 1 clearly showed the polygon view of the GGE biplot with the “Which-won-where” pattern. The first and second principal components accounted for 61.77 and 32.79%, respectively accounting for 94.56% of the GGE variation. The projecting lines from the origin divide the biplot into sectors, identify the mega-environments (environments in the same sector) and reveal which genotype was superior (the genotype at the vertex of the sector) in a mega-environment. The polygon view showed that all the test environments fell into two of the four sectors outlined. Ibadan and Ayetoro formed a mega-environment while Abeokuta was another mega-environment on its own. The biplot identified FUNAAB-11-8 (V2) as outstanding at Ibadan and Ayetoro mega-environment to which LD-88 (V4), FUNAAB-11-4 (V5) and NHGB/09/009A (V7) also belong. FUNAAB-11-3 (V1) was outstanding at Abeokuta where it out-performed F3 313x104 (V10), UI 4-30(V8) and FUNAAB-11-6(V3) that were shown to be associated with that mega-environment. F3 45x72-11 (V11), NH47-4 (V12), UI 53-139 (V9) and NG/TO/JUN/09/007 (V6) all fell into sector that contained none of the locations tested.
 | |
| Fig. 2: | GGE biplot of the mean yield performance and stability of 12 Okra genotypes in the tested locations (Genotype evaluation) |
Figure 2 contains the evaluation of the genotypes for mean yield performance and stability. The biplot was based on genotype focused singular value partitioning (“SVP = 1”) and therefore appropriate for visualizing the similarities among genotypes. It explained 94.56% of the total variation due to GGE. The two lines that passed through the origin are the ordinate (line with double arrows) and the abscissa (single arrowed) of the Average Environment Coordinate (AEC). The AEC itself represented by the small circle close to the abscissa is the mean PC1 and PC2 scores of the environment (Yan and Kang, 2003). The ordinate divides the genotypes into those that yielded above average (genotypes on the right) and those that yielded below average (genotypes on the left). Thus, the abscissa arrow points in direction of increasing yield performance. The best performer across three locations based on yield is FUNAAB-11-8 (V2) followed by NHGB/09/009A (V7), FUNAAB-11-4 (V5), FUNAAB-11-6(V3), FUNAAB-11-3 (V1) and LD-88 (V4). In the bottom half in descending order of performance are UI 4-30 (V8), UI 53-139 (V9), F3 313x104 (V10), NH47-4(V12), NG/TO/JUN/09/007 and F3 45x72-11(V11). The projections on to the ordinate are measures of variability or instability of the genotypes. The longer the vector irrespective of the direction, the more unstable is the associated genotype. Thus, short vector implies high stability (Yan and Kang, 2003). FUNAAB-11-4 (V5) has the shortest vector and therefore identified as the most stable. It is closely followed by UI 53-139 (V9), NHGB/09/009A (V7), NG/TO/JUN/09/007 and then FUNAAB-11-6 (V3). UI 4-30(V8) and F3 313x104 (V10) are fairly stable while NH47-4 (V12), LD-88 (V4), F3 45x72-11 (V11) and FUNAAB-11-3 (V1) are unstable. The small circle close to the arrow of the AEC abscissa delineates the ideal genotype and the genotype closest to it is adjudged the best (Yan and Kang, 2003), in this case it was NHGB/09/009A (V7).
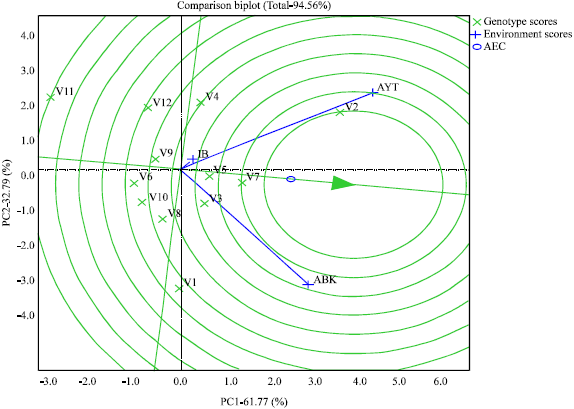 | |
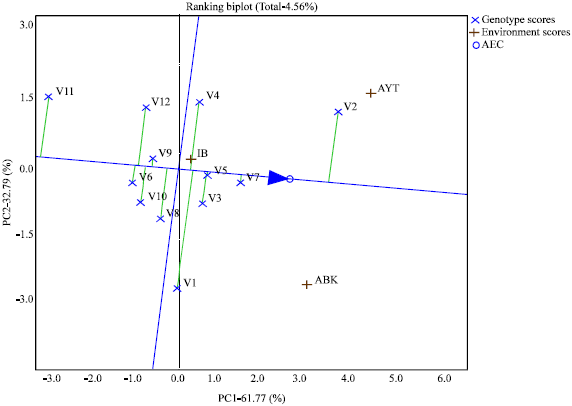
| Fig. 3: | GGE biplot showing the discriminatory ability and representativeness of the test environments |
Figure 3 shows the discriminatory ability and representativeness of the test locations. The length of the environment vectors (which approximates the standard deviation within each environment) from the biplot origin and the angle formed with the abscissa of the AEC reveals the discriminatory ability and the representativeness of the test locations (Yan and Kang, 2003). The longer the vector the higher the discriminatory ability of the associated environment and the shorter the angle formed the more the representative the associated environment (Yan et al., 2007). The biplot identified Ayetoro as the most representative since its vector formed the shortest angle with the AEC abscissa. It was followed by Abeokuta while Ibadan with the largest angle is the least representative. Ayetoro also has the highest discriminatory power due to its possession of the longest vector, followed by Abeokuta and then Ibadan with the least. The small circle close to the arrow of the AEC abscissa delineates the ideal environment and the location closest to it is adjudged the best (Yan and Kang, 2003). From the biplot Ayetoro was the closest to ideal environment and therefore the best.
Table 5 shows the means, stability variance, yield rank adjustment, stability ratings and the yield stability index (YSi) for 12 Okra genotypes. The yield stability index (YSi) by Kang and Magari (1995) employs the stability variance (Shukla, 1972) to determine the stability or instability of a genotype. The genotypes were ranked based on yield such that V2 which had the highest yield got the highest rank of 12, V7 that was second got a rank of 11 and the rank kept decreasing with reduced yield so that V11 associated with the lowest yield got the lowest rank of 1 (Kang and Magari, 1995). Genotypes with non-significant stability variance were adjudged to be stable (Shukla, 1972) as was the case with all genotypes except V2 that had a stability variance significant at 1% probability level (Kang and Magari, 1995).
| Table 5: | Mean yield, stability variance and the simultaneous selection of genotypes based on yield and stability (YSi) by Kang and Magari (1995) |
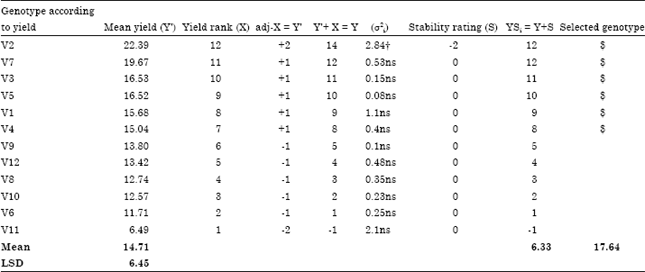 | |
| *adj-: Adjustment to, σ2i: Stability variance, †: Significant at p<0.1, $: Selection symbol according to Kang and Magari (1995) | |
| Table 6: | Comparison between the yield stability statistic and GGE biplot techniques showing mean yield of the selected genotypes in parenthesis |
 | |
The YSi labels a genotype as being high yielding and stable if it YSi is above average (6.33); thus V2, V7, V3, V5, V1 and V4 that met the requirement were selected ($) (Kang and Magari, 1995). Though V2 was identified as an unstable genotype, it was still selected by the YSi. The mean of the genotypes selected by the YSi was 17.64.
Table 6 shows the comparison between selections made by the YSi and the GGE Biplot. Both techniques selected V7, V5 and V3 while V2, V1 and V4 were only selected by the yield stability index. Though the two techniques made different number of selections, the means of their selections were statistically similar, 17.57 for GGE biplot and 17.64 for YSi.
DISCUSSION
It is a usual practice for breeders to evaluate different genotypes in multi-environment trials to measure stability of performance. The variation in the response of genotypes to different environments has made such trial a necessity. This phenomenon called GEI notoriously hinders progress from selection and superior varietal release. This is so because a genotype with good performance in an environment may be bad in another. It implies that the desirability of a genotype is not yield-based alone but in relation to stability to ensure repeatable performance. In this study the YSi and the GGE biplot techniques comparatively performed this function to evaluate twelve Okra genotypes for stability of performance and high yield.
The highest mean seed yield associated with FUNAAB -11-8 implies that this genotype is high yielding and had the highest average yield over the test locations. It is therefore desirable for selection based on yield alone. F3 45x72-11 with lowest mean seed yield is a poor genotype and therefore not desirable in terms of yield. Across the locations the rankings of genotypes differed. NHGB/09/009A that was the best at Ibadan turned out to be second and third at Abeokuta and Ayetoro respectively. FUNAAB-11-3 was the highest yielder at Abeokuta but came fifth and last at Ibadan and Ayetoro respectively. Also FUNAAB-11-8 that was the most superior at Ayetoro was tenth and third at Ibadan and Abeokuta respectively. The inconsistency in the ranking of the genotypes from one location to another is an indication of the presence of GenotypexEnvironment Interaction (GEI)-crossover type. According to Fernandez (1991), GEI refers to the differential ranking of genotype among locations or years. Crossover interaction is due to an imperfect correlation between genotype performance and environment (Ariyo, 2009). Haldane (1946) had earlier posited that when superior genotypes differ with environments, then GEI is suspected. It would therefore be technically wrong to base superiority judgement among these genotypes on yield alone. This justifies the employment of techniques that incorporated both stability and yield (GGE biplot and YSi) in this study.
The GGE biplot methodology (Yan et al., 2000; Yan, 2001, 2002; Yan and Kang, 2003; Yan and Tinker, 2006) allows for the identification of the best genotype in each mega-environment (“which-won-where”) designated as the genotype at the vertex position in the sector (Yan and Kang, 2003). In this study the high percentage of total variation explained by the first and second principal component axes (PC1 and PC2) indicated that the biplot of the two axes adequately approximated the environment-centered data. Subjecting the environment-centered yield means for each environment to singular value partitioning i.e., genotype-focused (SVP = 1) and environment-focused (SVP = 2) singular value decomposition made the biplot appropriate for visualizing the relationships among genotypes and environments respectively (Yan, 2001). The GGE biplot showed that the tested locations fell into two sectors, an indication that different genotypes won in different sectors. Thus, two mega-environments were identified-Ibadan and Ayetoro on one hand and Abeokuta on the other. The Ibadan and Ayetoro mega-environment contained LD-88, FUNAAB-11-4, NHGB/09/009A and FUNAAB-11-8 but the vertex genotype was FUNAAB-11-8 which was the most outstanding and therefore the most responsive genotype in the Ibadan and Ayetoro mega-environment. The Abeokuta mega-environment had FUNAAB-11-3 as the vertex genotype with F3 313x104, UI 4-30 and FUNAAB-11-6 as members. This also means that FUNAAB-11-3 was the most outstanding and therefore the most responsive genotype at the Abeokuta mega-environment. F3 45x72-11, UI 53-139, NH47-4 and NG/TO/JUN/09/007 fell into sector that contained none of the tested location. This means that these genotypes were not the best in any of the locations and probably the poorest in some or all of the locations (Yan, 2001). Indeed, the four genotypes had low yields in all the locations. GGE biplot ranks genotypes on both average yield and stability. NHGB/09/009A, FUNAAB-11-4 and FUNAAB-11-6 combined high yield with stability indicating that these genotypes possess the ability to avoid substantial fluctuation in yield over a range of environmental conditions (Heinrich et al., 1983). These three genotypes performed above average in the three tested locations. Though FUNAAB-11-8, FUNAAB-11-3 and LD-88 were high-yielding, they were not stable. This means that they had highly inconsistent rankings across the environments and therefore are unpredictable. UI 4-30 and F3 313x104 were fairly stable but were low yielding which makes them undesirable for selection. The case was similar with NG/TO/JUN/09/007 and UI 53-139 that though more stable were low yielding. Even poorer were NH47-4 and F3 45x72-11 that were both unstable and low yielding. Yan (2001) has defined an “ideal” genotype on the basis of mean performance and stability and represented it with a small circle on the biplot. This “ideal” genotype rarely exists in nature as a result the closest genotype to that point is adjudged the ideal (Yan and Kang, 2003). Based on this information the ideal genotype in this study was NHGB/09/009A as it was the closest to the ideal point though FUNAAB-11-4 and FUNAAB-11-6 are worthy of a mention. The GGE biplot evaluates the test locations through their “discriminatory power and representativeness”. The environment with the highest discriminatory power is that which gives the best information on the genotypic differences among the tested genotype while ability to represent the average environment confers representativeness (Yan, 2001; Yan and Kang, 2003; Yan et al., 2007). Yan (2001) defined an “ideal” test environment as one which is a virtual environment that has the longest vector of all test environments (most discriminatory) and located on the AEC abscissa (most representative). This “ideal” environment rarely exist consequently environments can be visually ranked for their usefulness in identifying superior genotypes based on the distances i.e. the number of rings (on the GGE biplot) between their markers and the marker of the ideal test environment.
Ibadan is the least representative and discriminatory among the locations. This means that it is farthest from the average (“ideal”) environment and offers little or no information on the differences among the genotypes. This is particularly true as the genotype means at Ibadan were not significantly different from one another. It is therefore not suitable to test these genotypes as the result may be misleading. According to Yan and Tinker (2006) such environments give little or no encouragement to good genotypes with concomitant potential to upgrade poor genotypes. Ayetoro with the longest vector (most discriminating) and the smallest angle with the AEC abscissa (the most representative) is ideal for selecting superior genotype. This is true of the test location as it’s the closest to the “ideal” environment with just a ring separating it from the ideal point. Abeokuta which was another ring away was second best in this regard as it formed a larger angle with the AEC abscissa than Ayetoro but possessed a shorter vector than Ayetoro. Yan et al. (2007) posited that such environment may not be ideal for selecting superior genotypes but could be useful in culling unstable genotypes. The yield stability index (YSi) by Kang and Magari (1995) selects superior genotypes based on yield and stability. It is an improvement on the Kang Rank Sum, KRS (Kang, 1988) that sums up the yield ranking of genotypes and their stability rank and Kang Modified Rank Sum, KMRS Kang (1991) that sums up the yield ranking and the stability rating. The YSi which is the summation of the adjusted Yield and stability rating therefore compensates or punishes genotypes for high or low yielding ability and for stability or instability. This technique employs the Shukla stability variance as a tool for determining the stability of a genotype. All genotypes except FUNAAB-11-8 had a non-significant stability variance an indication that all the genotypes were stable except FUNAAB-11-8 (Shukla, 1972; Kang and Magari, 1995). However a genotype is only adjudged to be stable and high yielding if its YSi value is more than the overall YSi average (Kang and Magari, 1995; Waldron et al., 2002; Nassir and Ariyo, 2011). Based on this criterion FUNAAB-11-8, NHGB/09/009A, FUNAAB-11-6, FUNAAB-11-4, FUNAAB-11-3 and LD-88 were selected. Also FUNAAB-11-8 and NHGB/09/009A that had the joint highest YSi may be considered as the best combiners of high yield and stability. GGE biplot and YSi were in agreement in the yield rank for all the genotypes but not for stability where only FUNAAB-11-8 was jointly identified to be unstable by the two techniques. For the YSi technique, all other genotypes were stable while the GGE biplot, in addition, identified FUNAAB-11-3, LD-88, NH47-4, F3 45x72-11 to be unstable. The two techniques were in agreement in selecting NHGB/09/009A, FUNAAB-11-6 and FUNAAB-11-4 as being high yielding and stable. It is noteworthy that these three genotypes had above average performances in the three test locations. Also GGE biplot identified NHGB/09/009A as the best combiner of high yield and stability-due to its proximity to the ideal genotype-just as Ysi selected NHGB/09/009A and FUNAAB-11-8 as the joint best in this regard as a result of their joint highest YSi values. Yan and Kang (2003) and Nassir and Ariyo (2011) mentioned that the YSi and GGE biplot do make similar simultaneous selections for stable and high yielding genotypes. FUNAAB-11-8, FUNAAB-11-3 and LD-88 that made the YSi list as being high yielding and stable were conspicuously missing in GGE biplot selection. These three genotypes did not consistently produce above average yield in all locations as seen at Ibadan and Ayetoro for FUNAAB-11-3, Ibadan for FUNAAB-11-8 and Ibadan and Abeokuta for LD-88. This is an indication that their abilities to avoid substantial fluctuations in yield across the location were not as strong as those that were jointly selected by the two techniques. Shukla stability variance which determines the stability or not of a genotype under the YSi technique adjudged FUNAAB-11-3 and LD-88 to be stable whereas GGE biplot labelled them as unstable. This may have paved the way for their selection by the YSi. The selection of these genotypes by the Ysi indicates that YSi may not be as effective as the GGE biplot in simultaneous selection of genotypes for yield and stability. Bachireddy et al. (1992) and Waldron et al. (2002) observed that in comparing two GEI techniques one may select more unstable genotype than the other. Though the YSi had more selections than the GGE biplot, the means of selection by the two techniques were not significantly different from each other. This implies that the additions of unstable but high yielding genotypes are of no advantage to a selection programme as such may be devastating due to crop failures (Kang, 1993). According to Yan and Kang (2003) although GGE biplot and YSi provide similar information, the GGE biplot is more comprehensive and its visual effects are unparalleled. The presence of GEI in this study necessitates that the Okra genotypes be tested in a wider range of environment including years and location to give more impetus to the stability observed and the identification of a best environment.
NHGB/09/009A adjudged to be the best genotype along with FUNAAB-11-4 and FUNAAB-11-6 should be considered in future Okra breeding programme particularly in hybridization.
REFERENCES
- Abdelmageed, A.H.A., 2010. Inheritance studies of some economic characters in okra (Abelmoschus esculentus (L.) Moench). Trop. Sub-Trop. Agroecosyst., 12: 619-627.
Direct Link - Alake, C.O., O.J. Ariyo and O.B. Kehinde, 2012. A quantitative analysis of the genetics of yield and yield components in west african okra, Abelmoschus caillei (A. chev) stevels. Int. J. Plant Breed. Genet., 6: 94-104.
CrossRef - Ariyo, O.J., 1993. Genetic diversity in West African okra (Abelmoschus caillei) (A. Chev.) Stevels-Multivariate analysis of morphological and agronomic characteristics. Genet. Res. Crop. Evol., 40: 25-32.
CrossRefDirect Link - Bachireddy, V.R., R. Payne Jr., K.L. Chin and M.S. Kang, 1992. Conventional selection versus methods that use genotype X environment interaction in sweet corn trials. HortScience, 27: 436-438.
Direct Link - Becker, H.C. and J. Leon, 1988. Stability analysis in plant breeding. Plant Breed, 101: 1-23.
CrossRef - Eberhart, S.A. and W.A. Russell, 1966. Stability parameters for comparing varieties. Crop Sci., 6: 36-40.
CrossRefDirect Link - Finlay, K.W. and G.N. Wilkinson, 1963. The analysis of adaptation in a plant-breeding programme. Aust. J. Agric. Res., 14: 742-754.
CrossRefDirect Link - Fisher, R.A., 1918. The correlation between relatives on the supposition of Mendelian inheritance. Trans. R. Soc. Edinburgh, 52: 399-433.
Direct Link - Gauch, Jr. H.G. and R.W. Zobel, 1988. Predictive and postdictive success of statistical analyses of yield trials. Theoret. Applied Genet., 76: 1-10.
CrossRefDirect Link - Heinrich, G.M., C.A. Francis and J.D. Eastin, 1983. Stability of grain sorghum yield components across diverse environments. Crop Sci., 23: 209-212.
Direct Link - Kang, M.S., 1993. Simultaneous selection for yield and stability in crop performance trials: Consequences for growers. Agron. J., 85: 754-757.
Direct Link - Kang, M.S. and R. Magari, 1995. STABLE: A basic program for calculating stability and yield-stability statistics. Agron. J., 87: 276-277.
Direct Link - Sanjeet, K., D. Sokona, H. Adamou, R. Alain, P. Nicolas and K. Christophe, 2010. Okra (Abelmoschus spp.) in West and Central Africa: Potential and progress on its improvement. Afr. J. Agric. Res., 5: 3590-3598.
CrossRefDirect Link - Nassir, A.L. and O.J. Ariyo, 2011. Genotype x environment interaction and Yield-stability analyses of rice grown in tropical Inland Swamp. Notulae Botanicae Horti Agrobotanici Cluj-Napoca, 39: 220-225.
Direct Link - Osekita, O.S., O.J. Ariyo and O.B. Kehinde, 2000. Variation and inter-character association in the segregating F3 populations arising from two crosses of okra (Abelmoschus esculentus (L.) moench). Moor J. Agric. Res., 1: 32-36.
Direct Link - Oyelade, O.J., B.I.O. Ade-Omowaye and V.F. Adeomi, 2003. Influence of variety on protein, fat contents and some physical characteristics of okra seeds. J. Food Eng., 57: 111-114.
CrossRefDirect Link - Shukla, G.K., 1972. Some statistical aspects of partitioning genotype-environmental components of variability. Heredity, 29: 237-245.
CrossRefDirect Link - Waldron, B.L., K.H. Asay and K.B. Jensen, 2002. Stability and yield of cool-season pasture grass species grown at five irrigation levels. Crop Sci., 42: 890-896.
Direct Link - Yan, W., 2001. GGEbiplot-A windows application for graphical analysis of multi environment trial data and other types of two-way data. Agron. J., 93: 1111-1118.
CrossRefDirect Link - Yan, W., 2002. Singular-value partitioning in biplot analysis of multienvironment trial data. Agron. J., 94: 990-996.
Direct Link - Yan, W. and N.A. Tinker, 2006. Biplot analysis of multi-environment trial data: Principles and applications. Can. J. Plant Sci., 86: 623-645.
CrossRefDirect Link - Yan, W., L.A. Hunt, Q. Sheng and Z. Szlavnics, 2000. Cultivar evaluation and mega-environment investigation based on the GGE biplot. Crop Sci., 40: 597-605.
CrossRefDirect Link - Yan, W., M.S. Kang, B. Ma, S. Woods and P.L. Cornelius, 2007. GGE biplot vs. AMMI analysis of genotype-by-environment data. Crop Sci., 47: 643-653.
CrossRefDirect Link - Yates, F. and W.G. Cochran, 1938. The analysis of groups of experiments. J. Agric. Sci., 28: 556-580.
Direct Link








