Research Article
Perspective on Chromosome Numbers in the Genus Pistacia L. (Anacardiaceae)
Department of Environmental and Plant Biology, Ohio University Zanesville, 1425 Newark Road, Zanesville, Ohio 43701, USA
The genus Pistacia L. belongs to the Anacardiaceae family and includes at least 11 species (Al-Saghir, 2010). Pistacia vera L. (cultivated pistachio) is by far the most economically important species in the genus. It has edible seeds and considerable commercial importance. The other species grow in the wild habitat and their seeds are used as a rootstock seed source and sometimes are used for fruit consumption, oil extraction and soap production.
The pistachio is native to the arid zones of Central Asia; it has been cultivated for 3000-4000 years in Iran and was introduced into Mediterranean Europe by Romans at the beginning of the Christian Era (Crane, 1974). Pistachio cultivation extended Westward from its center of origin to Italy, Spain and other Mediterranean regions of Southern Europe, North Africa and the Middle East, as well as to China and more recently to the United States and Australia (Maggs, 1973). Pistacia vera L. is the only species in this genus that is successfully grown in orchards; it produces edible seeds large enough to be commercially acceptable. Pistachios are adapted to a variety of soils and are probably more tolerant of alkaline and saline soil than most tree crops (Tous and Ferguson, 1996). Moreover, Pistachios thrive in hot, dry and desert-like conditions.
Currently, Iran, the United States, Turkey and Syria are the main Pistachio producers in the world, contributing over 90% of the world production.
Cytogenetic studies addressing the genus Pistacia are rather few. Chromosome numbers of the different Pistacia species are questionable due to the fact that poor chromosome counting protocols were used (Ila et al., 2003), these protocols are hampered by the extremely small sized chromosomes of Pistacia species and frequently having a few cell divisions visible in a single root tip (Ayaz and Namli, 2009).
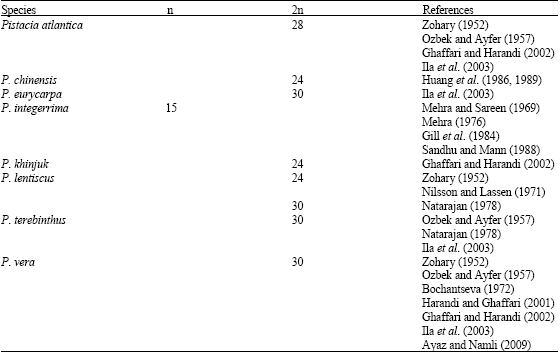
| Table 1: | Previous reports of chromosome data on Pistacia genus |
 | |
Chromosomal data have been valuable tools for cytogeneticists and breeders. They are often providing more insight into taxonomic and phylogenetic relationships (Raven, 1975; Stuessy, 1990). The aim of this perspective was to provide more insight into understanding the cytogenetics of the genus Pistacia and provide additional information on the different Pistacia species for future cytogenetic and breeding research.
Previous studies showed that all Pistacia species are diploid with chromosome numbers 2n = 24, 28 and 30 (Table 1). The 2n = 28 was reported for P. atlantica Desf. or its subspecies by Zohary, (1952), Ozbek and Ayfer (1957), Ghaffari and Harandi (2002). However, a recent study by Ila et al. (2003) reported the chromosome number for the first time as 2n = 30 for the same species.
Chromosome number of P. chinensis Bunge was reported as 2n = 24 (Huang et al., 1986, 1989). Chromosome number of P. eurycarpa Yalt. was reported as 2n = 30 (Ila et al., 2003) for the first time. Chromosome number of P. intergimma L. Stew. ex Brandis was reported as 2n = 30 (Mehra and Sareen, 1969; Mehra, 1976; Gill et al., 1984; Sandhu and Mann, 1988). Chromosome number of P. khinjuk Stocks was reported as 2n = 24 (Ghaffari and Harandi, 2002) and as 2n = 30 (Ozbek and Ayfer, 1957). The 2n = 24 was reported for P. lentiscus L. (Zohary, 1952; Nilsson and Lassen, 1971; Ghaffari and Harandi, 2002) and 2n = 30 by Natarajan (1978). Chromosome number for P. terebinthus L. as reported as 2n = 30 (Ozbek and Ayfer, 1957; Natarajan, 1978; Ila et al., 2003).
Chromosome number for P. vera L. was reportedly 2n = 30 (Zohary, 1952; Ozbek and Ayfer, 1957; Bochantseva, 1972; Harandi and Ghaffari, 2001; Ghaffari and Harandi, 2002; Ila et al., 2003; Ayaz and Namli, 2009).
I have been working on genus Pistacia for many years. The genus at morphological, anatomical and molecular level studied by Al-Saghir and Porter (2005, 2006), Al-Saghir et al. (2006). The study showed that Pistacia is monophyletic. Pistacia vera is the most primitive species according to morphological and anatomical characters (Al-Saghir, 2010).
All previous studies clearly showed that 2n = 30 is the right exact chromosome number for P. vera (Table 1). Therefore, I suggested that the primitive basic number for the genus is x = 15.
From the previous studies, the frequently reported chromosome number for most species is 2n = 30 (Table 1). Given the frequent hybridization and close genetic relationship among Pistacia species, it is likely that all species have the same basic number x = 15. If we consider the 2n = 28 and 2n = 24, it is unlikely that two reductions occurred in the genus (from 2n = 30 to 2n = 28; then from 2n = 28 to 2n = 24) given the close genetic relationship among species (instead the genus undergoes explosive radiation from an ancestral species similar to P. vera).
Pistacia khinjuk was reported as 2n = 24 and 2n = 30. Our morphological and genetic data showed, that P.khinjuk and P.vera are very closely related (clustered together) along with P. terebinthus (Al-Saghir, 2010). Therefore, it is unlikely that 2n = 24 is the right chromosome number.
Yi et al. (2008) assessed the phylogeny of Pistacia using five molecular sets, sequences of nuclear ribosomal ITS, the third intron of the nuclear nitrate reductase gene ( NIA-i3 ) and the plastid ndhF, trnL-F and trnC-trnD. Their molecular data were largely consistent with our independently derived intrageneric classification based on morphology. Pistacia was shown to be monophyletic in all analysis. The two accessions of P. vera formed a clade with P. khinjuk in all molecular data sets. Some of the ITS and NIA-i3 sequences of these two species were identical, suggesting a close relationship. Earlier molecular results also suggested a close relationship between them (Parfitt and Badenes, 1997; Kafkas and Perl-Treves, 2001, 2002; Golan-Goldhirsh et al., 2004). Pistacia palaestina was not well separated from P. terebinthus in either the plastid or nuclear DNA data sets and Yi et al. (2008) stated that, Pistacia palaestina may need to be merged into P. terebinthus. Close relationships between these two species were also suggested by the AFLP and the RAPD results (Golan-Goldhirsh et al., 2004; Kafkas, 2006; Al-Saghir and Porter, 2006). These results are consistent with Engler (1936) and Yaltirik (1967), along with our classification, who considered P. palaestina to be a synonym of P. terebinthus. Pistacia mexicana and P. texana were not distinguishable in the plastid restriction analysis (Parfitt and Badenes, 1997). The ITS data suggest that P. mexicana and P. texana are sister taxa and the sequence divergence between these two species is low. Our morphological data indicate that there is too little variation to warrant the recognition of two species. Pistacia saportae was shown to be a hybrid between P. lentiscus (maternal) and P. terebinthus (paternal), as others had hypothesized (Zohary, 1952).
In conclusion, I posit that all Pistacia species have the same basic number which is x = 15 based on all previous morphological and molecular studies, which clearly suggest a very close genetic relationship among Pistacia species and the chromosome numbers reported by various cytogenetic studies (Table 1).
The author is grateful to Ohio University Zanesville for funding this project.