ABSTRACT
Molecular characterization and genetic variation patterns between 18 varieties of water chestnut (Trapa sp.) were investigated through RAPD markers. PCR amplifiable DNA was isolated using the CTAB method and amplified fragments were obtained from 20 random 10-mer and 20 random 12-mer primers. Genetic distances among the different varieties were analyzed with a UPGMA-derived dendrogram. Similarity matrix showed the similarities between varieties ranged from 0.25 to 1.0. The dendrogram results suggested that the European varieties were in the same cluster group. Japanese varieties were divided into three clusters group. Indian varieties were in the separate cluster group. Korean and Japanese small were in the same cluster group. Chinese 1~7 and Japanese medium and Kobe-large were in the same group but have sub-group. Genetic distance of Kobe small is high and most devient from other varieties and made separate cluster group.
PDF Abstract XML References
How to cite this article
DOI: 10.3923/biotech.2005.144.148
URL: https://scialert.net/abstract/?doi=biotech.2005.144.148
INTRODUCTION
Water chestnut is a starch producing aquatic plant and some of them are cultivated in paddy field in place of rice in Japan[4] It is taxonomically interesting due to the mrique dicot nature of monotypic genus Trapa. Generally in plants, species relationships have been studied based on morphological, ecological, karyological and biochemical similarities. From the beginning, all species of this genus are classified according to floral, fruit and leaf morphology. Although the present classification is based solely on morphological characteristics, comprehensive phylogenetic relationships among the species of Trapa have not yet been clarified specially biochemical. Occasionally T. bispinosa formed 3 to 4 horns as same as T. quadrispinosa and researchers are faces many problems to identifying their name because of the diverse variation of fruit shape.
Now a days, many molecular techniques have been used to distinguish cultivars in several crops and this technique is solely dependent on their genetical protein levels and highly informative. Various DNA-based marker systems have been applied to several plants groups for delimiting clones and to assess their level of relatedness[5,6].One of the most efficient molecular marker methods in terms of ability to produce polymorphic markers within a comparatively short time and with a limited budget is RAPD (Random Amplified Polymorphicb DNA). Sience its introduction about a decade ago[7], RAPD has become widely used in various plant research and the potential to contribute to advances m breeding, genetics and systematics. Intra-populational and intra-specific differentiation assessed using RAPDs[8].
From these points of view, we studied and elucidating the phylogenetic relationships among the species of Trapa which also collected from the different parts of the world, based on RAPD.
MATERIALS AND METHODS
Eighteen varieties of water chestnut (Trapa sp., Table 1) from Japan, Chim, India, Italy, Israel, France, Korea were examined for RAPDs.
Total genomic DNA was extracted from fresh leaves, using CTAB method. A composite 1 g sample ofleaffrom each variety was grmmd in liquid nitrogen with a mortar and pestle and transferred powder to an appropriate centrifnge tube and immediately added 3 mL 2% CTAB solution. After completely mixing, incubated 30 to 60 min at 65°C. Extracted with an equal volume of chloroform and centrifuge 10 min at 8000 rpm, 4°C. Transferred aqueous phase to a fresh tube and extracted with equal volume of chloroform again as in previous step. Transferred aqueous phase to a fresh tube and added 9 mL 1 % CTAB solution and also mixed thoroughly as well as centrifuged 1 0 min at 8000 rpm, 4°C.
| Table 1: | Plant materials used in the RAPD analysis |
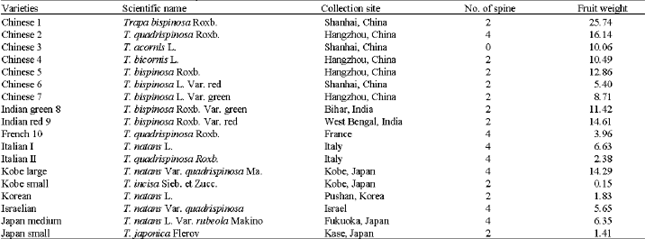 | |
| Table 2: | Nucleotide sequence, number of base pairs (size) and guanine and cytosine content (G+C%) of the DNA in the primers used to generate RAPD bands by Polymerase Chain Reaction |
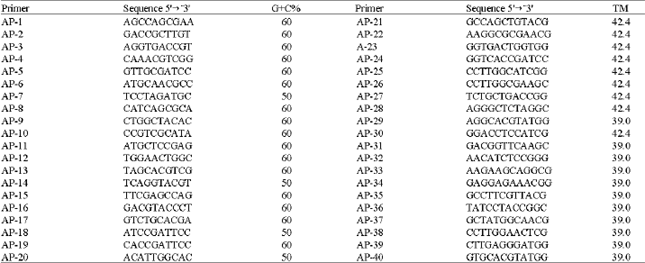 | |
Removed supernatant and resuspend pellet in 4 mL 1 M NaCI solution. Added same ammmt of isopropanol and mixed thoroughly and centrifuged 10 min at 8000 rpm, 4°C. Removed supernatant and rinse pellet with cold 70% ethanol solution. Removed 70% ethanol solution with wiping paper (Do not dry completely). Resuspend pellet with 0.5 mL TE buffer containing RnaseA (100 mg-1 ) and incubated 60 min at 37°C to remove RNA. The DNA length and concentration were determined in comparison with standard -DNA by electrophoresis in 1% agarose gel in TBE buffer and by spectrophotometry, respectively. The DNA concentration was adjusted to 20 ng μL-1 before use and the solution was stored at -20°C nntil analyzed.
Polymerase Chain Reaction (PCR) was carried out in 10μL of each reaction mixtures. The reaction mixtures contained genomic DNA (20 ng μL-1) 2 μL, I OX PCR buffer I μL, 2.5 mM of dNTPs I μL, primer (2 pmol μL-1) 5 μL, Tth DNA polymerase (Fast Start Tag) 0.1 μL and dH2 0 PCR 0.9 μL. Amplification reactions were carried out in a Quarter Bath DNA thermal cycler (I-Cycler, BIORAD, USA) using the following sequence of conditions during each of 30 cycles: 1 min at 95°C (denaturation), 2 min at 35°C (annealing) and 3 min at 72°C (extension). After the last cycle, reaction mixtures were incubated at 72°C for 8 min to ensure that the primer extension reactions were completed.
The 40 arbitrary primers used in the initial experiment are shown in Table 2. Amplification products were analyzed by electrophoresis using 1% agar gel in Iris Borate EDTA (TBE) buffer pH 8.0 at a constant voltage of I OOV, followed by staining with ethidium bromide and visualising nnder UV radiation. To determine the size of amplified fragments, a DNA marker ladder with I kb steps or 2log (New England, BioLabs) was nm in each geL Later experiments with five primers were repeated at least twice to determine the repeatability of the results.
| Table 3: | Similari!Y matrix of 18 cultivars of water chestnut |
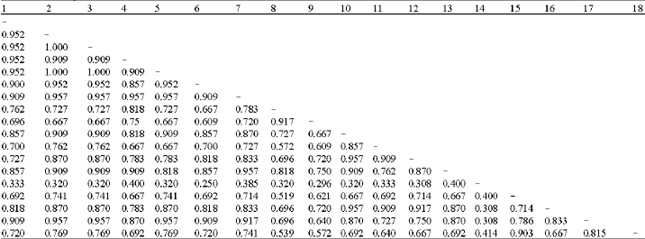 | |
The Random Amplified Polymorphic DNA (RAPD) banding profiles of each of water chestnut varieties were scored manually for the presence (1) or absence (0) of the bands. These qualitative data were analyzed using Nei similarity index (Nei andLi 1979) which excludes common negative data on the basis of the following equation: Similarity= 2 Nab/(Na + Nb), where, Nab= number of scored amplified fragments with the same molecular weight shared between genotype a and b; Na =number of scored amplified fragments in genotype a, Nb = number of scored amplified fragments in genotype b. A dendrogram was constructed on the basis of the similarity matrix data by Unweighed Pair Group Method with Aritlnnetic Average (UPGMA) cluster analysis using the software MEGA program (Pennsylvania State Uinv., PA).
RESULTS AND DISCUSSION
The strategy to obtain reproducible fragment profiles of water chestnut DNA involved reactions in which various components of the reaction mixture were varied. Large changes in concentrations of template DNA did affect the amplification, too little DNA resulted in either reduced or no amplification of small fragments. As the DNA concentration was increased the number of fragments appearing on the gels was also increased, while too much DNA produced either a smear effect or did not amplify any fragments.
The optimum concentration of dNTP was 250 FM, which gave increased specificity, high fidelity as well as satisfactory yield of PCR products. At lower dNTP concentrations fragments were faint, while higher concentrations generated W1Separated, large sized bands, which could have been because of mispriming and misincorporation of nucleotides. Since dNTPs bind magnesium ions, the magnesium chloride concentration required for maximum activation of the enzyme depends on the dNTP concentrations. At low MgC12 ion concentration, the number of DNA fragments amplified with a given primer were either decreased or absent. However, the number of fragments amplified was directly correlated with the mcrease m magnesium Ion concentration to certain extent.
Preliminary screening facilitated the selection of primers producing a higher level of polymorphism and more reproducible fragment patterns. Such screenings are essential to save time, cost and to reject primers not informative for the analysis. In the preliminary experiments, eight primers, namely AP-1, AP-4, AP-5, AP- 11, AP-16, AP-22 and AP-30 (Table 2) showed the more clear polymorphisms in RAPD banding patterns among the water chestnut varieties. The banding patterns were reproducible in repeated experiments. The banding patterns generated by the respective primers can be classified into several types based on the presence or absence of bands of various sizes in the gels (Fig. 1).
Similarity values are summarized in Table 3 and the relationships among the varieties are shown as a dendrogram in Fig. 2. Similarities between varieties ranged from 0.25 to 1.0 (Table 3). The highest similarity values were formd between the varieties of 1~7 and 17. The lowest similarity value of 0.25 was observed in the variety 14.
From the dendrogram results it is clear that the European varieties (Italian I, Italian II, French and Israelian (Fig. 2), Israelian cultivars previously collected from Italy) are in the same cluster group. Japanese varieties are divided into three clusters group. Indian varieties are in the separate cluster group. Korean and Japanese small are in the same cluster group. Chinese 1~7 and Japanese medium and Kobe-large are in the same group but have sub-group (Fig. 2).
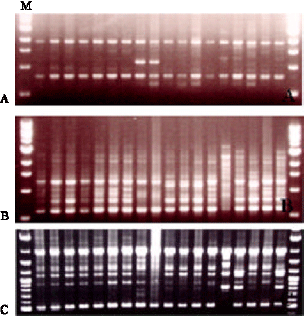 | |
| Fig. 1: | RAPD profiles obtained by amplification of DNA samples with primer AP-1 (Fig. A), AP-5 (Fig. B) and AP-30 (Fig. C). The samples from left to right represent accessions 1 to 18 and the left-right-most lane represent a molecular marker (M) |
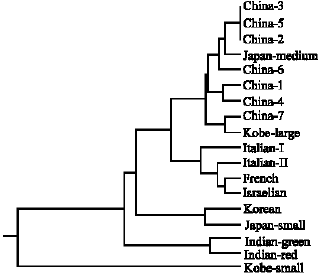 | |
| Fig. 2: | Dendrogram showing the relationship among the 18 water chestnut varieties as obtained by cluster analysis of the distance values. |
With prudent selection of primers and adherence to a particular set of DNA extraction, purification and amplification procedures, it has been shown that RAPDs provide a reliable method for the identification of 18 water chestnut varieties of diverse origin. It was found in the present experiments that the purity of template DNA, type of PCR equipment, dNTP concentration, MgCl2 reaction mixture were the most critical factors affecting the reproducibility of RAPD banding patterns. These confirm the conclusions of Ellsworth et al.[9] that the amplification profiles with specific oligonucleotide primers are highly dependent on PCR conditions and that the banding patterns may vary extensively because of inconsistencies in a number of reaction parameters. The type of thermocycler used for RAPD analysis was also reported to be a key determinant ofthe reproducibility of banding patterns among different laboratories[10].Taken together, these variations in techniques explain the occasional production of artificial DNA products[11] and the doubts raised by Weeden et al.[12] and others about the reproducibility and reliability of the RAPD technique. Nevertheless, the technique has been useful for detecting genetic polymorphisms and for variety and clonal identification in different plant species[13-15].
The present study is the first report on the use of RAPD markers to estimate the molecular diversity in water chestnut. Although study has brought out the close relationship between the genotypes, there were no duplicates. The standardized protocols and results obtained could be utilized in future analyses and improvement programmes.
REFERENCES
- Hoque, A., S.M. Rahman, S. Arima and Y. Takagi, 2001. Efficient in vitro germination and shoot proliferation of chilling treated water chesnut (Trapa japonica Flerov.) embryonal explants. In Vitro Cell. Dev. Biol., 37: 369-374.
Direct Link - Williams, J.G.K., A.R. Kubelik, K.J. Livak, J.A. Rafalski and S.V. Tingey, 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res., 18: 6531-6535.
CrossRefPubMedDirect Link - Nei, M. and W.H. Li, 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA., 76: 5269-5273.
CrossRefPubMedDirect Link - Penner, G.A., A. Bush, R. Wise, W. Kim and L. Domier et al., 1993. Reproducibility of Random Amplified Polymorphic DNA (RAPD) analysis among laboratories. PCR Methods Applied, 2: 341-345.
CrossRefDirect Link - Ellsworth, D.L., D. Rittenhouse and R.L. Honeycutt, 1993. Artifactual variation in random amplified polymorphic DNA banding patterns. Bio. Technol., 14: 214-217.
PubMed - Weeden, N.F., G.M. Timmerman, M. Hemmat, B.E. Kneen and M.A. Lodhi, 1992. Inheritance and Reliability of RAPD Markers. Proceedings of the Symposium on Application of Rapd Technology to Plant Breeding, November 1, 1992, Crop Science Society of America, Madison, WI., pp: 12-17.
Direct Link - Shigyo, M., T. Miyazaki and Y. Tashiro, 2002. Development of randomly amplified polymorphic DNA markers in cultivated and wild species of sections Cepa and Phyllodolon in Allium. J. Hortci. Sci. Biotechnol., 77: 373-377.
Direct Link - Khanuja, S.P.S., A.K. Shasany, M.P. Darokar and S. Kumar, 1999. Rapid isolation of DNA from dry and fresh samples of plants producing large amounts of secondary metabolites and essential oils. Plant Mol. Biol., Rep., 17: 1-7.
Direct Link








