Research Article
Phylogenetic Analysis of the Genus Pistacia L. (Anacardiaceae) Based on Morphological Data
Department of Environmental and Plant Biology, Ohio University Zanesville, 1425, Newark Road, Zanesville, Ohio 43701, USA
Pistacia L. belongs to the family Anacardiaceae (cashew family), order Sapindales (Stevens, 2008). It contains nine species and five subspecies according to the current study. Species are xerophytic, dioecious trees or shrubs (rarely monoecious) with resins. Leaves are alternate, deciduous or evergreen, usually pinnately compound, rarely trifoliate or simple, membranous or leathery, stipules are absent. Flowers are in panicles or racemes, small, unisexual, apetalous, subtended by 1-3 small bracts and 2-7 bracteoles and wind pollinated. Staminate flowers with 4-5 stamens, filaments inserted on a disc. Pistillate flowers with a compound pistil of 1-5 united carpels with 1 locule, ovules usually are 1 per locule and borne on an axile placenta; the ovary is superior, with a short 3-parted style; the stigma is capitate. Fruit is a 1-seeded drupe (according to the current study).
Pistacia is believed to have originated in Central Asia 80 million years ago (Parfitt and Badenes,1997; Kafkas and Perl-Treves, 2001; AL-Saghir, 2009). Two centers of diversity have been described. One comprises the Mediterranean region of Southern Europe, Northern Africa and the Middle East. The second comprises West and Central Asia (Kafkas and Perl-Treves, 2002; Kafkas et al., 2002). The species of the genus occur naturally from North Africa to the Philippines and from Texas to Honduras. Few systematic studies have been published on this important genus. The first complete classification of the genus was published by Zohary (1952).
Today, there are many questions about Zohary’s taxonomic treatment, the status of many of his species and the accuracy of his dividing the genus into four sections. This disagreement is mainly because of little information being available on the phylogeny of the genus. Previous phylogenetic studies were hampered by the small sampling size, weak representation of the species and poor resolution of the methods used. Consequently, these studies failed to resolve the relationships between species within the genus. Moreover, species of Pistacia easily form interspecific hybrids, suggesting close relationships, which limit the previous studies and make the actual level of speciation and relationships within the genus unclear. A better understanding of these relationships is also needed to make the species more useful for plant improvement or genetic studies.
AL-Yafi (1978) divided P. atlantica into four subspecies based on their leaf morphologies. Kokwaro described a new species from East Africa, P. aethiopica, in Kokwaro and Gillett (1980) based on leaf morphology and tree size. Lin et al. (1984) characterized leaf morphology, photosynthesis and leaf conductance of nine Pistacia species (P. atlantica, P. chinensis, P. integerrima, P. khinjuk, P. lentiscus, P. mexicana, P. mutica, P. terebinthus, P. texana, P. vera and P. weinmannifolia). EL-Oqlah (1996) described Pistacia species in Jordan morphologically and anatomically (P. atlantica, P. lentiscus,and P. palaestina). Kafkas and Perl-Treves (2001) addressed the taxonomic relationships and genetic variation of wild Pistacia germplasm in Turkey using morphological data and Random Amplified Polymorphic DNA (RAPD). Pistacia atlantica, P. eurycarpa and P. terebinthus, the common wild species in the flora of Turkey, were included in this study. In addition, many varieties of P. vera were added for comparison. The morphological analysis revealed that among the species studied the closest species to P. vera is P. eurycarpa. Kafkas et al. (2002) characterized morphologically and surveyed the germplasm of three wild Pistacia species in Turkey (P. atlantica, P. eurycarpa and P. terebinthus). All genotypes displayed high diversity at the inter- and intra-specific level. Barazani et al. (2003) assessed the genetic variability of Mediterranean P. lentiscus genotypes by morphology, composition of essential oils and RAPD. High polymorphism in morphological parameters was found among accessions, with no significant differences in relation to geographical origin or gender.
Morphological characters provide a source of data for taxonomic research on different plant taxa and may help determine the direction and the course of evolution and provide the distinctions between species (Benson, 1962). The objective of the present study was to clarify and provide additional insight into understanding the taxonomic and phylogenetic relationships among Pistacia species using morphological data.
Plant material and outgroups: This project started in 2004 and ends in 2009. It was conducted at Virginia Polytechnic Institute and State University and Ohio University Zanesville. The study included 15 described species of the genus (Table 1). It was based on analysis of a wide range of field and herbarium material from the Field Museum, Chicago, Illinois, USA (F); Missouri Botanical Garden, St. Louis, Missouri, USA (MO); Natural History Museum, London (BM); Royal Botanic Garden, Edinburgh, UK (E); Royal Botanic Gardens, Kew, UK (K); and Virginia Polytechnic Institute and State University, Blacksburg, Virginia, USA (VPI).
Four hundred and twenty eight specimens were examined. The broad spectrum of the collections should ensure the inclusive distribution of all taxa. Thirty three samples represent four wild Pistacia species collected by the author during his trip to Jordan in June 2004.
| Table 1: | List of Pistacia species used in this study |
 | |
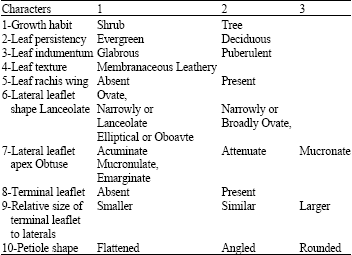
| Table 2: | Qualitative morphological characters used for phylogenetic analysis in this study |
 | |
This trip was made in order to observe the plants in the field. Various literature sources were used to pinpoint and refine the taxonomy and the localities of specimens included.
Rhus aromatica, R. copallina,R. glabra and Schinus molle were included as an out-group to the genus based on recent studies that highlight the close relationship of Rhus and Schinus to Pistacia (Miller et al., 2001).
Description of Pistacia species and scoring morphological characters.
Thirty characters were examined. Only 19 (1 habitat and 18 leaf characters) were used for phylogenetic analysis because these were the only characters found to be polymorphic among Pistacia species. The species were described with respect to tree and leaf morphology based on Benson (1979) using Pistacia descriptors developed by the International Plant Genetic Resources Institute (IPGRI, 1998). Ten characters of 19 were qualitative (Table 2) and 9 were quantitative (Table 3). Each character was scored for up to 84 samples of each species.
| Table 3: | Quantitative morphological characters used for phylogenetic analysis in this study |
 | |
Phylogenetic analysis: Neighbor joining and parsimony analyses were performed on the morphological data using PAUP 4.0 b10 (Swofford, 2003). A heuristic search was performed by RANDOM stepwise addition (10,000 replicates) with Tree Bisection-Reconstruction (TBR) branch swapping to find the most parsimonious tree. From the resulting trees, the majority rule consensus was calculated. Robustness and topology of the trees were evaluated using the Consistency Index (CI) to measure the level of homoplasy in the data (how cleanly the data fit the tree). The Retention Index (RI) was measured to determine the amount of synapomorphy on the tree. Robustness and reliability of the resulting phylogenies were measured by bootstrap (1,000 replicates with full heuristic searches). The stepwise distance matrix which measures the relatedness between the species (Kimura, 1980) was computed between all species. The data were analyzed with a neighbor joining approach.
To investigate the relationships between the Pistacia species on the basis of morphological data, 19 phenotypic characters were coded as discrete states. All characters used were polymorphic and very useful in distinguishing between and separating the different species.
All the morphological characters used were parsimony informative characters. Parsimony analysis produced one maximally parsimonious tree of 49 steps, with a Consistency Index (CI) of 0.69 and a Retention Index (RI) of 0.75 (Fig. 1). This tree was consistent with the tree produced by neighbor joining analysis (Fig. 2, 3). However, there were differences between the bootstrap support values of the different branches of the two trees and the relationships among species within each clade.
The morphological data used in the two analysis strongly supported the monophyly of Pistacia. The genus divided into two monophyletic groups: One group (Section Pistacia) contains P. atlantica, P. chinensis, P. eurycarpa, P. falcata, P. integerrima, P. khinjuk, P. mutica, P. palaestina, P. terebinthus and P. vera while the other group (Section Lentiscella) contains P. aethiopica, P. lentiscus, P. mexicana, P. texana and P. weinmannifolia (Fig. 1- 3).
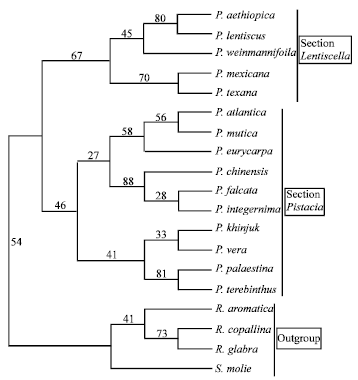 | |
| Fig. 1: | The majority rule consensus tree of 232 equally parsimonious trees generated with morphological data using parsimony analysis showing relationships of the Pistacia species with CI = 0.69, RI = 0.75 and tree length 49 steps. Numbers above the branches indicate the bootstrap values |
Both analysis show that section Pistacia contains three sister groups, the first group with P. atlantica, P. mutica and P. eurycarpa, the second with P. chinensis, P. falcata and P. integerrima and the third group contains P. khinjuk, P.palaestina, P. terebinthus and P. vera. The first group was monophyletic with the second group which had weak bootstrap support of 27% in the parsimony analysis and 35% in the neighbor joining analysis. Bootstrap support of the first group was 58% in the parsimony analysis and 88% in the neighbor joining analysis. Pistacia atlantica and P. mutica show a close relationship (56% in the parsimony analysis, 95% in the neighbor joining analysis). A branch appearing in 88% of the parsimony analysis bootstrap replicates separates P. chinensis from P. falcata and P. integerrima within the second group. Pistacia falcata and P. integerrima show a relationship with bootstrap support of 28%. A branch appearing in 91% of the neighbor joining analysis bootstrap replicates separates P. integerrima from P. chinensis and P. falcata. Pistacia chinensis and P. falcata show a relationship with 40% support.
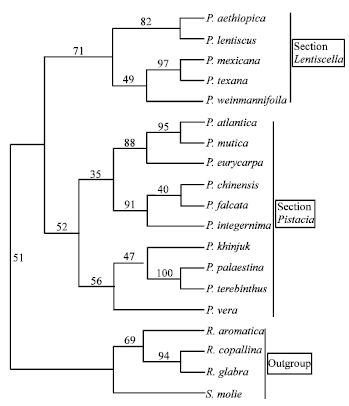 | |
| Fig. 2: | The majority rule consensus tree generated with morphological data using neighbor joining analysis showing relationships of the Pistacia species. Numbers above the branches indicate the bootstrap values |
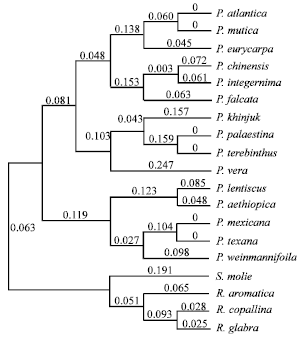 | |
| Fig. 3: | The majority rule consensus tree generated with morphological data using neighbor joining analysis showing relationships of the Pistacia species. Numbers above the branches indicate the branch lengths |
Both analysis support the branch that separates the first group and the second group from the third group with bootstrap support of 46% in the parsimony analysis and 52% in the neighbor joining analysis. A branch appearing in 41% of the parsimony analysis bootstrap replicates separates P. vera and P. khinjuk (which are related with 33% support) from P. terebinthus and P. palaestina (which are closely related with 81% support). A branch appearing in 56% of the neighbor joining bootstrap replicates separates P. vera from P. khinjuk, P. terebinthus and P. palaestina. A branch appearing in 47% of the neighbor joining bootstrap replicates separates P. khinjuk from P. terebinthus and P. palaestina. The latter two species are very closely related with 100 % support in the neighbor joining analysis. The only difference between the two analysis is the placement of P. khinjuk within the third group.
Section Lentiscella, as indicated by the two analysis, was also monophyletic. In the parsimony analysis, a branch appearing in 67% of the bootstrap replicates separates this section into two monophyletic groups, the first group contains P. aethiopica, P. lentiscus (both show a close relationship with bootstrap support of 80%) and P. weinmannifolia. The first two species were sister to P. weinmannifolia with moderate bootstrap support of 45%. The second group contains P. mexicana and P. texana which show a close relationship with bootstrap support of 70%.
In the neighbor joining analysis, a branch appearing in 71% of the bootstrap replicates separates the section into monophyletic groups. The first group contains P. aethiopica and P. lentiscus (which show a close relationship with bootstrap support of 82%). The second group contains P.mexicana and P. texana (which show a close relationship with bootstrap support of 97%) and P. weinmannifolia, which is sister to the first two species with bootstrap support of 49%. The only difference between the two analysis is the placement of P. weinmannifolia between the two groups.
This is the first comprehensive study to include this number of Pistacia species and a broad spectrum of herbarium specimens that represent each species. Both the maximally parsimonious tree and the neighbor-joining tree resolve the relationships among Pistacia species based on the current morphological data. The two trees support the monophyly of the genus as well as resolving the two sections of the genus, Pistacia and Lentiscella. Section Pistacia species are large deciduous trees (rarely shrubs), while section Lentiscella species are evergreen shrubs or small trees. Zohary (1952) divided the genus into four sections based on leaf and fruit morphology. In this study, posit a two sectional division of the genus based on leaf morphology. The current results agree with Parfitt and Badenes (1997) classification, which suggests the same two sectional division of the genus and with the results of cluster analyses made by Kafkas and Perl-Treves (2002), Katsiotis et al. (2003) and Golan-Goldhirsh et al. (2004). Moreover, the current study clearly defines the species status and the taxonomic relationships among the members of each section based on both analysis including the pairwise distances between all pairwise combinations of the species and the thorough morphological observation of a good number of specimens of each species.
Within section Pistacia, P. atlantica was close and sister to P. eurycarpa (0.11 unit genetic distance) and P. mutica. While there was no genetic distance between P. atlantica and P. mutica (0.00 unit genetic distance), the two species are found on the same branch (Fig. 1-3). Little morphological variation occurs in the specimens of the two species examined from different geographical areas. The measurements of all characters used were almost identical and found it difficult to distinguish between the two species. investigated the original description of P. mutica by Fischer and Meyer (1838). It matches the description of P. atlantica (Desfontaine, 1779) and the current analysis supports Zohary’s (1952) determination that P. atlantica and P. mutica are the same species. Given the branch support in both parsimony and neighbor joining analysis (including genetic distance value) (Fig. 1-3), P. mutica is treated as a synonym of P. atlantica. In addition, both have wide, odd-pinnate, membranaceous leaves with flattened petioles; the leaflets are 7-11, lanceolate, obtuse and puberulent with a ciliated margin; the terminal leaflet is similar or smaller than the lateral ones and the rachis is narrowly winged.
Zohary (1952) classified P. eurycarpa as a synonym of P. atlantica var. kurdica because of the presence of a leaf rachis wing. Yaltirik (1967a, b) on the other hand, treated this plant as a different species based on leaf characters. In the current analysis, P. atlantica was close and sister to P. eurycarpa (0.11 unit genetic distance) (Fig. 1, 2). This result agrees with Yaltirik’s (1967a, b) classification and observation of P. eurycarpa as a separate species (Fig. 1-3) and not as a variety of P. atlantica as suggested by Zohary (1952) because the leaves are light-green on both sides (instead of being dark green above and pale below as in P. atlantica) and the fruits are depressed and larger (Yaltirik, 1967a, b) which are confirmed by the current study. Furthermore, the leaflets (4.5-6.5 cm long, 1.5-3.8 cm wide) are usually wider and thicker than and are never as numerous as in P. atlantica (3-7 cm long, 5-20 mm wide) and the rachis wing is narrower or even absent. This treatment is supported by the phenotypic cluster analysis made by Kafkas and Perl-Treves (2001) in which the taxonomic placement of P. eurycarpa was identical to the current study’s results.
In both the parsimony analysis and the neighbor joining analysis (Fig. 1, 2), the closest relative of P. vera is P. khinjuk. According to the average genetic distances between the different species, P. khinjuk and P. vera were separated by 0.42 unit genetic distance. Pistaciakhinjuk was the closet relative to P. vera but a separate species. Pistacia khinjuk hasodd-pinnatemembrananceous leaves, the leaflets are 1-9, distinctly pinnately veined with an acuminate apex, while leaves in P. vera aretrifoliolate or odd-pinnate and leathery and the leaflets are 3-5, the lower often subopposite, ovate to broadly ovate, obtuse or mucronulate, the mid-veins and margins are minutely pubescent.
According to both analysis, P. palaestina and P. terebinthus are closely related. They form a close pair and have been regarded as a single species (0.00 unit genetic distance). Engler (1883), the first to monograph Pistacia, considered P. palaestina as a variety of P. terebinthus. However, Zohary (1952) considered P. palaestina as a distinct species, because of two main distinctive characteristics: P. palaestina has mostly even-pinnate leaves and acuminate leaflets, while P. terebinthus has odd-pinnate leaves and obtuse or acute leaflets. Yaltirik (1967a) described two subspecies within P. terebinthus. The first, P. terebinthus subsp. terebinthus had odd-pinnate leaves with the terminal leaflet of the median leaves often as large as the lateral ones and obtuse or ovate-oblong lateral leaflets. The second P. terebinthus subspecies had either even-pinnate and/or odd-pinnate leaves; the terminal leaflet of the median leaves was always smaller than the laterals or reduced to a bristle and it had acuminate or oblong-lanceolate lateral leaflets. The results of this study support Engler’s and Yaltirik’s classifications. The two entities are morphologically, ecologically and genetically similar (Zohary, 2000; Kafkas and Perl-Treves, 2002). Unique morphological variation was not observed in either species. It was difficult to distinguish between P. palasetina and P. terebinthus specimens and noticed that problem as well in Yaltirik’s annotations of specimens. Leaves in both species are odd-pinnate or even-pinnate and membranaceous; the leaflets are 6-11, ovate to narrowly ovate with an acuminate or mucronate apex; the terminal leaflet (if present) is smaller than the laterals or reduced. Therefore, posit that P. palaestina should be merged with P. terebinthus and become a synonym of the latter name.Pistaciachinensis is the most heterogeneous and complex species in the genus. Both analysis reveal that P. chinensis, P. falcata and P. integerrima are close sister taxa. In the parsimony analysis (Fig. 1), P. chinensis was sister to P. falcata and P. integerrima. In the neighbor joining analysis (Fig. 2), P. integerrima was sister to P. chinensis (0.13 unit genetic distance) and P. falcata (0.13 unit genetic distance). My results support Zohary’s (1952) conclusion that theses names represent the same species. If one compares the specimens of the three species, they would not fail to observe the close relationships between these taxa. The three species have large deciduous leaves that are 13.0-21.0 cm long and 11.4-18.2 cm wide, odd-pinnate or even-pinnate, membranaceous, with a flattened petiole; the leaflets are 10-14, 5.2-9.1 cm long and 1.0-2.0 cm wide, narrowly lanceolate, falcate, attenuate and glabrous; the terminal leaflet (if present) is 2.0-4.0 cm long and 0.5-1.1 cm wide, smaller than the laterals or reduced; the rachis is not winged. I propose that P. falcata should be treated as a subspecies of P. chinensis not as a variety, because their populations are geographically separated. Leaves in P. falcata (13.0-21.0 cm long and 11.4-18.2 cm wide) arelarger than in P. chinensis (8.2-23 cm long and 8.0-20.0 cm wide). In P. chinensis, leaflets are 8-16 and lanceolate, while leaflets in P. falcata are 10-14 and narrowly lanceolate. Moreover, P. falcata leaflets have an elongated apex, abundant veins and distinct narrow lanceolate leaflets.
Pistacia falcata is found in East Africa. Its ancestors may have been transported to Madagascar and East Africa from India or China by migratory birds as hypothesized by Renner (2004) for Melastomataceae. Anacardiaceae, including Pistacia, have fruits that are adapted for bird dispersal. The fruits are single-seeded. These fruits have a fleshy mesocarp, pseudarils and seeds are high in fat and protein, they have thick endocarps that protect the seed from being crushed (Jordano, 1989). Birds probably discard the endocarp immediately after eating the mesocarp or pseudaril and/or void some of the endocarps intact later. This mode of dispersal has been suggested, based on these fruit adaptations, by Weeks et al. (2005) as the major mode of Burseraceae fruit dispersal and Burseraceae is the sister family of Anacardiaceae. East African Pistacia could have been transported by fruits that may have been blown by the monsoon wind system via occasional storms between Africa, the Chagos Archipelago (half way between Africa and Indonesia) and India. Alternatively, fruits could have become lodged in floating vegetation as hypothesized by Renner (2004) for Melastomataceae.
This study agrees with Zohary’s (1952) classification of P. integerrima as a subspecies of P. chinensis also. P. integerrima has large leaves, 11.0-22.0 cm long and 12.0-23.0 cm wide; the leaflets are 6-14, 6.0-11.5 cm long and 2.0-4.0 cm wide and broadly lanceolate. In P. chinensis the leaves are 8.2-23.0 cm long and 8.0-20.0 cm wide, the leaflets are 8-16, 4.0-10.0 cm long and 0.9-2.4 cm wide and lanceolate. Moreover, the distinct geographical distribution of this taxon (native to the western Himalayas) from P. chinensis subsp. chinensis is another piece of evidence to consider this plant as a subspecies, not as a variety.
Within section Lentiscella, in both analysis P. lentiscus and P. aethiopica form a close pair. Kokwaro and Gillett (1980) described P. aethiopica from East Africa based on leaf morphology and tree size. My results support Zohary’s classification of it as a variety of P. lentiscus (P. lentiscus var. emarginata). Pistacia aethiopica has a distinct leafy narrow rachis wing, abundant large veins and an emarginate or mucronulate leaflet apex. Ancestors of this taxon may have reached East Africa in the same way as was hypothesized for P. falcata above. Therefore, I suggest that this plant should be treated as a subspecies of P. lentiscus, not as a variety, given the above morphological variation and the distinct geographical distribution from P. lentiscus subsp. lentiscus.
In both analysis, P. mexicana and P. texana form a close pair and they are regarded as one species (0.00 unit genetic distance). Zohary (1952) and Parfitt and Badenes (1997) classified them as separate species. I posit that P. texana and P. mexicana are the same species; P. texana should be merged with P. mexicana and the name becomes a synonym of the latter. Leaves in both species are evergreen, clustered towards ends of branches, usually odd-pinnate, 5.1-15.1 cm long, 1.8-5.3 cm wide, membranaceous; petiole flattened; rachis narrowly winged. Leaflets 10-30, 1-2.5 cm long, 3-10 mm wide, average ratio 2.7-1, narrowly ovate, slightly falcate, mucronate, glabrous; terminal leaflet 5-25 mm long, 3-10 mm wide, same size or smaller than laterals. Pistacia mexicana is the only New World species in the genus. It is perhaps descended from an ancestor like the Asian P. weinmannifolia and its ancestors may have reached the New World from China via a long distance dispersal or migration through a probable trans-Atlantic Tertiary Landbridge (Renner, 2004; Weeks et al., 2005).
Pistacia weinmannifolia is sister to P. mexicana and P. texana (3.0 unit genetic distance) in the parsimony analysis, while in the neighbor joining analysis it is sister to P. lentiscus (5.0 unit genetic distance) and P. aethiopica (3.0 unit genetic distance). Morphologically, Pistacia weinmannifolia is closer to P. mexicana than to P. lentiscus and P. aethiopica. Both have small, membranaceous leaves; the leaflets are small, ovate to narrowly ovate (rarely obovate), emarginate and are often alternate.This supports (but not strongly) the hypothesis that the ancestor of P. mexicana resembled P. weinmannifolia.
In comparison between the neighbor joining tree and the parsimony tree, there was a disagreement in placing a number of taxa such as P. vera and P. integerrima (Fig. 1, 2). This may be because of the different philosophies and assumptions of each method and treatment of the data (distance matrix in the neighbor joining method and discrete matrix in the parsimony method). The difference in the resolution of the morphological characters may account for this variation. The following characters could contribute to the difference: leaflet number, leaflet shape, leaflet apex shape and leaflet size.
Yi et al. (2008) assessed the phylogeny of Pistacia using five molecular sets, sequences of nuclear ribosomal ITS, the third intron of the nuclear nitrate reductase gene (NIA-i3) and the plastid ndhF, trnL-F,and trnC-trnD. Their molecular data were largely consistent with our independently derived intrageneric classification based on morphology. They recognized the following species: P. atlantica, P. chinensis, P. integerrima, P. khinjuk, P. lentiscus, P.mexicana, P. palaestina, P. saportae, P. terebinthus, P. texana, P. vera and P. weinmannifolia. Except for P. palaestina and P. texana, our study recognized these taxa (P. integerrima as a subspecies of P. chinensis), plus P. chinensis subsp. falcata, P. eurycarpa and P. lentiscus subsp. emarginata. Pistacia was shown to be monophyletic in all analyses. The two accessions of P. vera formed a clade with P. khinjuk in all molecular data sets. Some of the ITS and NIA-i3 sequences of these two species were identical, suggesting a close relationship. Earlier molecular results also suggested a close relationship between them (Parfitt and Badenes, 1997; Kafkas and Perl-Treves, 2001, 2002; Golan-Goldhirsh et al., 2004). Pistacia palaestina was not well separated from P. terebinthus in either the plastid or nuclear DNA data sets and Yi et al. (2008) stated that, Pistacia palaestina may need to be merged into P. terebinthus. Close relationships between these two species were also suggested by the AFLP and the RAPD results (Golan-Goldhirsh et al., 2004 ; Kafkas et al., 2002; Al-Saghir and Porter, 2006). These results are consistent with Engler (1936) and Yaltirik (1967a), along with our classification, who considered P. palaestina to be a synonym of P. terebinthus. Pistacia mexicana and P. texana were not distinguishable in the plastid restriction analysis (Parfitt and Badenes, 1997). The ITS data suggest that P. mexicana and P. texana are sister taxa and the sequence divergence between these two species is low. Our morphological data indicate that there is too little variation to warrant the recognition of two species. Pistacia saportae was shown to be a hybrid between P. lentiscus (maternal) and P. terebinthus (paternal), as others had hypothesized (Zohary, 1952).
In conclusion, the study resolves the phylogenetic and taxonomic relationships among Pistacia species. In addition, it shows that the genus is monophyletic and is divided into two sections: section Pistacia and section Lentiscella.
This study provides an additional insight into taxonomy and phylogeny of the genus Pistacia.
This study results from studies undertaken for a PhD dissertation in the Department of Biological Sciences at Virginia Polytechnic Institute and State University, Blacksburg, Virginia. I thank Professors Brent Opell, Duncan M. Porter, Stephen Scheckler and M. A. Saghai-Maroof for their supervision and valuable comments. I also thank the curators of the Field Museum, Chicago, Illinois (F); Missouri Botanical Garden, St. Louis (MO); Natural History Museum, London (BM); Royal Botanic Garden, Edinburgh (E); Royal Botanic Gardens, Kew (K); and University of Texas, Austin (UT) for the loan of specimens; Mr. Thomas Wieboldt (VPI) for arranging this; and Professor Paul Brazhnik for translation of Russian herbarium labels.