Research Article
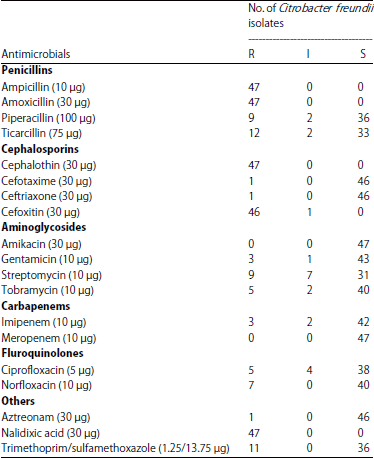
Virulence Factors and Antimicrobial Resistance Pattern of Citrobacter freundii Isolated from Healthy Pet Turtles and their Environment
Veterinary Medical Center, College of Veterinary Medicine, Chungbuk National University, 28644 Cheongju, Korea
S.H.M.P. Wimalasena
Veterinary Medical Center, College of Veterinary Medicine, Chungbuk National University, 28644 Cheongju, Korea
Gang-Joon Heo
Veterinary Medical Center, College of Veterinary Medicine, Chungbuk National University, 28644 Cheongju, Korea