Research Article
Prevalence of Arcobacter spp. in Humans, Animals and Foods of Animal Origin in India Based on Cultural Isolation, Antibiogram, PCR and Multiplex PCR Detection
Division of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), India
Ramswaroop Singh Rathore
Division of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), India
Kuldeep Dhama
Avian Diseases Section, Division of Pathology, Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), India
Thadiyam Puram Ramees
Division of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), India
Anil Patya
Avian Diseases Section, Division of Pathology, Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), India
Prashanth Suresh Bagalko
Division of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), IndiaDivision of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), IndiaDivision of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), IndiaDivision of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), IndiavDivision of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), India
Mohd. Yaqoob Wani
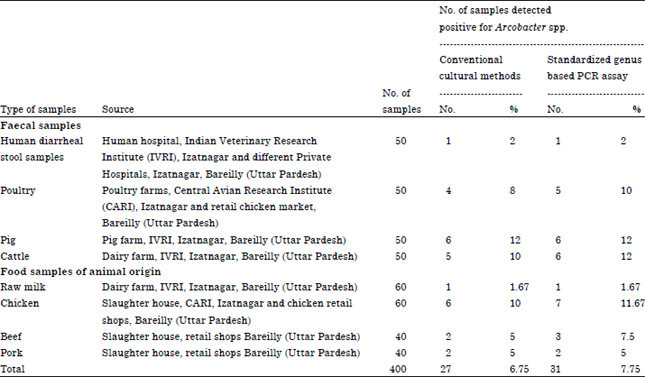
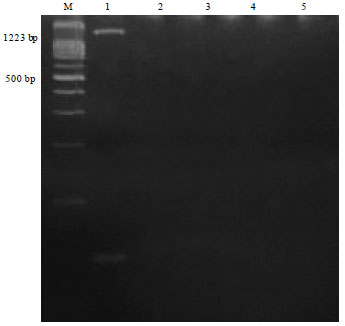
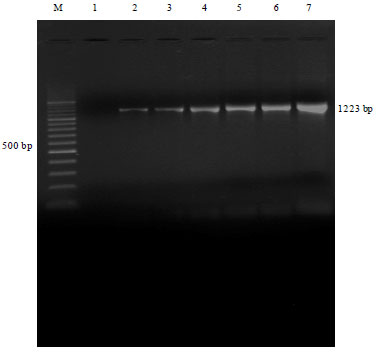
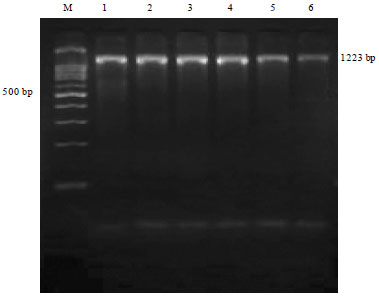
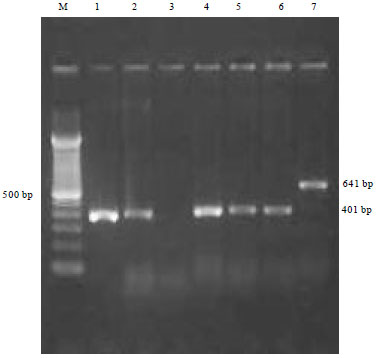
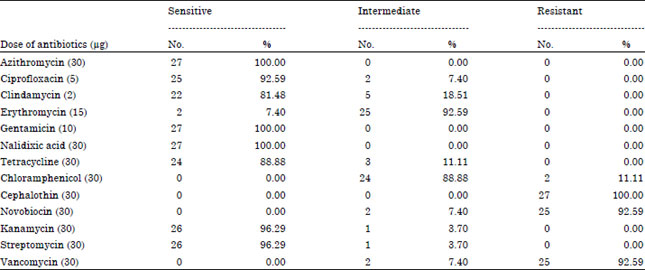
Arcobacter is an important emerging food and water borne pathogen having worldwide public health concern. The present study reports the prevalence of Arcobacter spp. in humans, animals and foods of animal origin based on cultural isolation, antibiogram, Polymerase Chain Reaction (PCR) and multiplex PCR detection. A total of 400 samples were collected as human diarrheal stool (50), faecal swabs of poultry (50), pig (50), cattle (50) and foods of animal origin [Raw milk (60), chicken meat (60), beef (40) and pork (40)]. The overall prevalence rate of Arcobacter spp. was found to be 6.75% (27/400) by cultural isolation with highest prevalence in pig faeces (12%), followed by cattle faeces (10%), chicken meat (10%), poultry faeces (8%), beef (5%), pork (5%), human diarrheal stools (2%) and milk (1.67%). PCR screening revealed prevalence of Arcobacter spp. to be 7.75% (31/400) with highest in pig faeces (12%), followed by cattle faeces (12%), chicken meat (11.67%), poultry (10%), beef (7.5%), pork (5%), human stools (2.00%) and raw milk (1.67%). Multiplex PCR assay enabled detection of A. butzleri (21/27) and A. skirrowii (6/27). In vitro antibiotic sensitivity profile of 27 Arcobacter isolates revealed most of these to be sensitive to azithromycin, gentamycin, nalidixic acid, kanamycin, streptomycin, ciprofloxacin and tetracycline. Higher resistance was observed for cephalothin, novobiocin and vancomycin with notable intermediately resistance against erythromycin and chloramphenicol. The present study demonstrated high prevalence of Arcobacter spp. in pig, cattle and poultry faecal samples which may play important role in contamination of environment, water and human food chain, thus could be of public health concerns. The PCR was found to be more rapid, sensitive, specific and efficient than cultural methods for detection of Arcobacter spp.
Kiran Narayan Bhilegaonkar
Division of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), India
Ashok Kumar
Division of Veterinary Publich Health,Indian Veterinary Research Institute, Izatnagar, 243122, Bareilly (Uttar Pardesh), India