Research Article
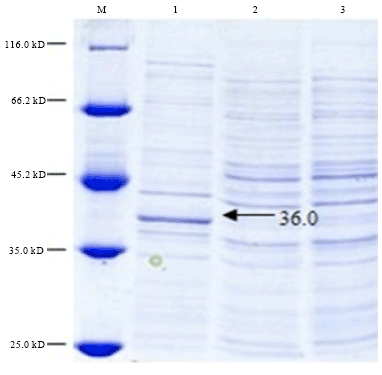
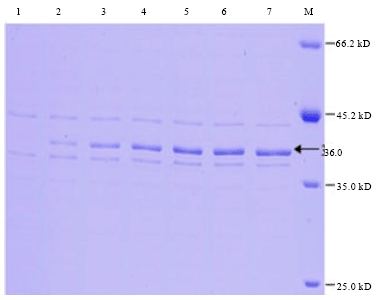
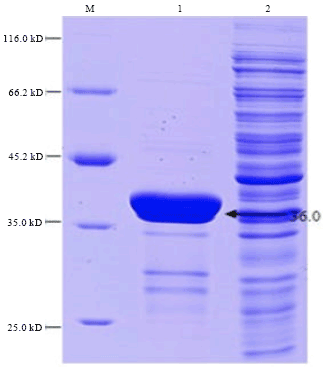
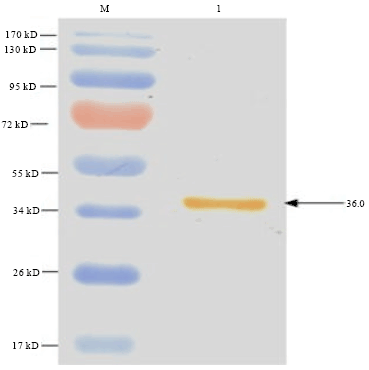
Cloning, Expression and Purification of Recombinant Envelope Protein VP36A of White Spot Syndrome Virus
Department of Microbiology, College of Veterinary Medicine, Nanjing Agricultural University, Nanjing 210095, Jiangsu, People`s Republic of China
Rongmei Fei
Department of Microbiology, College of Veterinary Medicine, Nanjing Agricultural University, Nanjing 210095, Jiangsu, People`s Republic of China