Research Article
Variation in the Population of Alternaria solani by Using Sequencing of ITS1 Isolated from Tomato Plants from Jordan Valley
Department of Biology, College of Haqel, University of Tabuk, Saudi Arabia
LiveDNA: 962.15505
The species of Alternaria solani (Ellis and G. Martin) Sorauer causing early blight disease on tomato plants become very important pathogen. Early blight is an important and widely distributed disease throughout the world resulting economic yield losses. The yield loss caused by Alternaria solani on tomato fruits1,2 was 48-80%. Symptoms of early blight on tomato plant start on lower, old and mature leaves, which become chlorotic and abscise prematurely spots of one-half inch in diameter. These spots have concentric rings or ridges and have a target-like pattern which always surrounded by a yellow halo. Moreover, these symptoms affect stem and fruits as well3.
In most pathogens, there is a variation between the populations from different areas. The variation in the populations of plant pathogenesis well known that they directly affects disease control, especially when the method related to the resistant cultivars development and fungicide usage. Alternaria solani found to be a highly variable pathogen based on morphological, physiological and molecular characterizations in different parts of the word4-7. Lourenco et al.8 found that the Brazilian population of Alternaria solani is highly genetically variable by studying up to 150 isolates obtained from potato plants and tomato plants and analyzed with RAPD and AFLP markers to estimate the amount and distribution of genetic variability of Alternaria solani in Brazil.
Various researches have been characterized Alternaria solani in different part of the world6,7,9-12. A parts of these researchers studied the variation of the population based on morphological and physiological characterizations2,6,13,14. Other researchers looked into the variations of the populations based on molecular characterizations7,15. Different molecular methods were used to study the variation of the population of Alternaria solani. El Komy et al.15 found a variation in the population of the Alternaria solani isolated from potato in Egypt by using random amplified polymorphic DNA (RAPD) and simple sequence repeats (SSR) markers. Loganathan et al.16 found that Alternaria solani recovered from diseased tomato plant in India has wide variation in the population by using the sequencing of ITS1 and ITS4. Other studies using the ITS2 sequencing technology to study the variations in the population for Aspergillus sp.17.
In Jordan, scarcity of studies have been investigated Alternaria solani. The majority of these studies were investigated the disease control18-20. A single research was characterized Alternaria solani morphologically and physiologically6. This study will be the first study designed to characterize the fungus of Alternaria solani recovery from diseased tomato plants grown in Jordan valley based on molecular methods using sequences of ITS region of rDNA.
Isolation: Twelve isolates were used and divided in 4 groups, isolated from diseased tomato plants grown in Jordan valley as described in AlHussaen6. These isolates were recovered from different farms in the Jordan valley in early 2012. The infected leaves were cut into small bits measuring about 5 mm and sterilized their surface with 1% sodium hypochlorite solution for 1 min, then washed with sterile distilled water. The 5 mm pieces were placed on the media of Potato Dextrose Agar (PDA) and incubated under 12 h light and 12 h dark at 25±1°C according to Naik et al.21. Pure culture of the fungus was obtained by hyphal tip isolation method.
Molecular methods: Molecular work was done in the period of March, 2017 to September, 2017. Mycelium of Alternaria solani isolates were used to extract the DNA and PCR was used to amplify the nuclear rDNA ITS region, including ITS1 and ITS2 and the 5.8 S ribosomal gene according to Paul22. The sequencing of ITS1 was carried out and maintained at a commercial facility (Macrogen Inc., Seoul, South Korea) following the standard methods. BLAST search (http://blast. ncbi.nlm.nih.gov) was used to analyze the sequences and ClustalW (http://www.ebi.ac.uk) was drown the phylogram of the isolate and its relative.
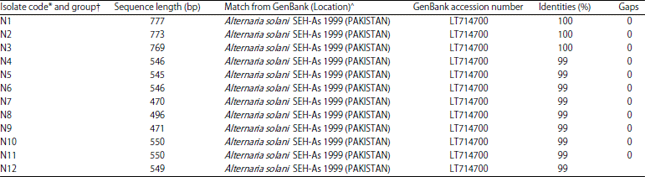
Sequence of ITS region of rDNA: Sequences of 12 isolates obtained in this study matched Alternaria solani (LT714700) from GenBank (Table 1). The sequence of isolate N1 was 777 bp in length and was 100% identical to the corresponding sequence from Alternaria solani (LT714700). Moreover, the sequence of isolates N2 was 773 bp in length and N3 was 769 bp in length were 100% identical to the corresponding sequence from Alternaria solani (LT714700). However, the sequence of isolate N4 was 546 bp in length was 99% identical to the corresponding sequence from Alternaria solani (LT714700). Furthermore, isolates N5 to N12 were 99% identical to the corresponding sequence from Alternaria solani (LT714700) (Table 1).
| Table 1: | Sequence length (bp) of ITS region of rDNA for 12 isolates of Alternaria solani from diseased tomato plants and comparison with sequences in GenBank |
 | |
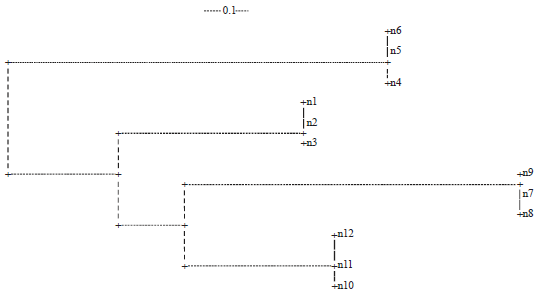 | |
| Fig. 1: | Dendrogram based on sequence of ITS region of rDNA of the Alternaria solani |
Variation observed using the sequence of ITS region of rDNA: Data described by sequence length (bp) of ITS region of rDNA were used to construct an UPGMA dendrogram (Fig. 1). The UPGMA dendrogram showed that there were 4 groups (Fig. 1) at similarity coefficient 0.1. Each group contained 3 isolates obtained from diseased tomato plants collected from different farms in the Jordan valley in early 2012. These four groups showed that the isolates obtained from the same area and showed the same morphological and physiological characters were grouped together in the UPGMA dendrogram (Fig. 1).
This is the first study to characterize the fungal of Alternaria solani by molecular methods recovery from diseased tomato plants in Jordan valley. There was a single study in Jordan described Alternaria solani based on morphological and physiological characterization6. Moreover, a scarcity of studies in Jordan on Alternaria solani on disease management and prevent were done and fungicides sensitivity18,20,23. Variation in populations of plant pathogens is important to control strategies.
Alternaria solani isolates described in this study was identified based on morphology and physiology characteristics according to the Alternaria identification manual6,24. The morphology features sometime are similar within the different groups of the same species and sometimes did not results to correct identification25. Moreover, these differences may be exacerbated by environmental factors (such as temperature) and physiological factors (such as nutrition)3. Morphological characteristics, however are still useful and often provide the basis for species identification.
The sequences of the ITS region of rDNA for the 12 Alternaria solani isolates, selected to represent the 4 groups described in Alhussaen were obtained to confirm the identification to species level6. The 12 isolates were found to be identical to Alternaria solani (LT714700) in the GenBank. The identification of fungi by using sequences of the ITS region of rDNA is very useful and give correct identification with most resent researches use it26-29.
This study demonstrated that there were variation in the population of Alternaria solani recovery from tomato plant grown in Jordan valley based on the sequences of the ITS region of rDNA. This variation could be related to different farms that use different agriculture techniques and the environmental conditions in the area of Jordan valley, which has high temperatures and humidity. Molecular techniques had been used to investigate the variation between populations of one species obtained from different areas or different environmental condition or even different host29-32. The method used in the present study, sequences of the ITS region of rDNA, more accurate to develop a clear vision of the variation in the population of plant pathogens. Several researches reported that the sequences of the ITS region of rDNA is the best technique to drew a map of the population of plant pathogen29.
The variation in the populations of plant pathogens is well known that they directly affects disease control, especially when the method related to the resistant cultivars development and fungicide usage. Moreover, this results will help researcher in finding the best control methods to Alternaria solani.
In this study, Alternaria solani isolated from diseased tomato plants from Jordan valley have a variation in the population based on molecular characteristics. This variation mean to control this fungus and need more research by using different fungicides and more controlling methods in order to keep the early blight disease under control. The results of this study will help farmers and researchers in finding the best and the effective control methods for the early blight disease and for the Alternaria solani.