ABSTRACT
Many DNA repair genes have been identified by current genome sequencing projects. X-ray cross-complementing group 3 (XRCC3) assists to RAD51 complex for DNA repair and recombination in humans. A gene for XRCC3 (AtXRCC3) was isolated and characterized from Arabidopsis. The AtXRCC3 gene consists of single exon in chromosome 5 in Arabidopsis and contains 915bp open reading frame encoding 304 amino acids. The AtXRCC3 amino acid sequence contains characteristic P-loop for ATP/GTP binding site motif A. It is similar to other RecA/RAD51 family sequences and highly homologous to human XRCC3 with 46% similarity. Yeast cells expressing AtXRCC3 showed resistance to methylmethane sulfonate. These results suggest that AtXRCC3 serves a role for DNA repair and recombination in plants.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ijb.2006.23.28
URL: https://scialert.net/abstract/?doi=ijb.2006.23.28
INTRODUCTION
DNA is incessantly damaged by ultraviolet, X-rays or mutagen throughout the entire lives of organisms. Damaged DNA can lead to the disruption of cellular processes, such as DNA replication and recombination and cause to lethal to cells. To counteract this damage, the cell has evolved tolerance mechanisms in which damaged DNA is rapidly repaired through the precise and concerted activities of a variety of repair proteins. DNA repair mechanisms also maintain genetic stability and suppress the formation of aberrant aging events that can cripple the cell. The damages are occurred not only on single strand of DNA but also on the breaks of double strands (DSB) (Britt, 1996). The DSB can be generated by physical stress on chromosomes or crossover during the process of meiosis or by treatment of radioactive rays or radiomimetic chemicals, which have the similar effects like ionization in cells. There are two well-known repair mechanisms for DSB (Ray and Langer, 2002). One is the Non-homologous End Joining (NHEJ) which attaches the damaged double-strands areas. The other is the Homologous Recombination (HR) mechanism, which replaces the damaged areas with the same gene using gene conversion. The problems in inversion, deletion, translocation and partial duplication occurred in case of the NHEJ because the damaged ends of DNA are randomly connected (Lees-Miller and Meek, 2003). Contrary to this, there are no such problems occurred in HR repair because it uses the same gene in homologous chromosome. Although the repair mechanism of DSB has been revealed to exist in all living organisms, they differ in which to apply first above two mechanisms. In general, creatures with small genome such as bacteria and yeasts use HR mechanism when similar DNA sequence to the damaged area exists. On the other hands, higher organisms with large genome such as plants and animals frequently use NHEJ to repair damages on (Valerie and Povirk, 2003). HR mechanism has been well studied in yeasts and occurs throughout four stages: finding homologous sequence, pairing DNA, strand invasion and synthesis of DNA to finish repairing (Shinohara and Ogawa, 1995). RAD52 epistasis group (RAD50-57, RAD59, MRE11, XRS2) is known as genes contributed to HR repair. These genes play important roles in repairing on DSB and the process of recombination in meiosis. Especially, RAD51 plays central roles in the process of HR as a homologue of RecA recombinase (Sung, 1994; Gupta et al., 1997). Several genes related to HR were reported in human based on RAD51 homology. Especially, five RAD51 paralogs (RAD51B/RAD51L1, RAD51C/RAD51L2 and RAD51D/RAD51L3, XRCC2, XRCC3) were reported and there are 20-30% identities between RAD51 and other protein sequences. Moreover, it has been confirmed by yeast two-hybrid assay that RAD51 paralogs interacts with one or more of the others and combines mutually to serve function (Schild et al., 2000; Masson et al., 2001). XRCC3 is a DNA repair protein needed to repair DNA damage caused by ionizing radiation and interacts with RAD51B and RAD51C (Wiese et al., 2002). RAD51 and RecA homologs have been reported as their engaging in HR in Arabidopsis plants (Doutriaux et al., 1998). Until now, the genes related in HR are not completely reported in plants but the researches have been actively conducted in this area after the determination of whole genome sequences of Arabidopsis thaliana which provides fertile ground for isolating important genes that regulate repair events.
We characterized a gene encoding XRCC3 with the immediate goal of elucidating the function of the XRCC3 protein in plants. Here, the full-length cloning of the Arabidopsis gene encoding a DNA repair protein was reported, XRCC3, in detail, predict its putative protein domain and examine its activity in yeast strains with a heightened sensitivity to DNA damaging agents.
MATERIALS AND METHODS
DNA sequence analysis: DNA sequencing was performed by the dideoxynucleotide chain termination method with an automatic sequencer (AlF express DNA sequencer, Pharmacia Biotech, Inc.). Nucleotide sequences and amino acid sequences were compared with sequences present in the GenBank and EMBL databases and analyzed using BLAST and PILEUP of the GCG Wisconsin Package or Biology WorkBench 3.2 (http://workbench.sdsc.edu; San Diego Super computer Center; University of California San Diego). Comparison of sequences with published animal XRCC3 sequences was performed at the nucleotide and amino acid level. PSORT program (http://psort.nibb.ac.jp) was used to predict localization site into cellular organelles and protein sorting signals on the XRCC3 amino acid sequence.
Polymerase Chain Reaction (PCR): To isolate the XRCC3 gene, which is consisted of single exon in Arabidopsis, PCR was performed with the primers indicated below. The plant materials were prepared from leaves of mature plants on soil. XRCC3 gene was amplified using primers derived from the genomic sequence of XRCC3 in GenBank (Acc. No. AB011482): X3-F primer (5'-TGGTACCTTTTGGCGGTGAAATG-3') and X3-R primer (5'-ATCTCCGGATCCTTCTGCAAT-3'). The underlined regions in the X3-F and X3-R primers are sites for restriction enzymes KpnI and BamHI, respectively, introduced to facilitate cloning after PCR. The PCR was performed using BiothermTM DNA polymerase (Genecraft Ltd, Munster, Germany) in Perkin-Elmer GeneAmp PCR System 9700(Norwalk, CT). PCR conditions: 5 min at 95°C, followed by 30 cycles: 60 sec at 94°C, 30 sec at 55°C and 60 sec at 72°C, followed by 7 min at 72°C.
Over expression of XRCC3 in yeasts and MMS resistance test: To investigate the function of At XRCC3, the gene was subcloned into the yeast expression vector pYES2/CT (Invitrogen Corp., Carlsbad, CA) by KpnI and BamHI sites to generate plasmid pYES::XRCC3; expression of XRCC3 is under the control of the inducible GAL1 promoter. The pYES::XRCC3 or pYES2/CT (as a negative control) were transformed into a yeast strain lacking RAD51 (LSY402; Liu et al., 2002). Transformants were selected and maintained on agar containing SD supplemented media without uracil (SD-Ura, Invitrogen Corp, Liu et al., 2002).
The resistance to DNA damage of yeast strains conferred by plasmid pYES:: XRCC3 was assessed using a traditional radiomimetic drug-sensitivity assay as follows: A single yeast colony was inoculated and grown in SD-Ura liquid media to OD600 = 0.7. XRCC3 was then induced by replacing the SD-Ura media with a second SD-Ura media containing 2% Galactose (Gal), which induces the GAL1 promoter in the plasmid. The culture was grown to OD600 = 0.5-0.9 over a period of 9 to 24 h. The culture was diluted 1/10-5 and plated on SD-Ura+Gal media containing various concentrations of the Mutagen Methylmethane Sulfonate (MMS) ranging from 0-0.05.
Intracellular localization of XRCC3::GFP in plants: To investigate the intracellular localization of AtXRCC3, the 5’ partial gene of 0.44kb was subcloned into the GFP expression vector pCAMBIA1302 by KpnI and EcoRI sites to generate plasmid XRCC3::GFP. The fusion construct was bombarded into onion epidermis at a pressure of 500 psi by using a Bio-Rad Helios Gene Gun (Bio-Rad) with 0.1 μg of DNA and 1 mg gold particles. Bombarded tissues were analyzed in the confocal microscope (Zeiss LSM510).
RESULTS
Identification of XRCC3 gene and sequence analysis: DNA repair genes typically exhibit sequence similarity, even among widely differing species. To identify genomic sequences from Arabidopsis plants based on sequence similarity to known DNA repair genes, we performed a database search with various gene fragments known to encode conserved regions of DNA repair proteins. Queries to the GenBank database using human XRCC3 protein sequences (GenBank Acc. No. AF035586) by TBLASTN revealed an Arabidopsis gene with similar sequence (GenBank Acc. No. AB011482) and I identified it as a putative XRCC3 gene. The sequence analysis in the genome of Arabidopsis was predicted that At XRCC3 is consisted of single exon existed as single copy in Arabidopsis.
The XRCC3 gene was directly isolated by PCR method with X3-F and X3-R primers.
 | |
| Fig. 1: | Amino acid sequence alignment of the XRCC3s from Arabidopsis and human ClustalW was used to generate the alignment. Dashes indicate gaps introduced to optimize the alignment. The identical or similar amino acids were aligned or represented by dots, respectively and the conserved motif was underlined with bold. The GenBank accession numbers for each sequence are as follows; Arabidopsis thaliana (AtXRCC3), AY032998, this study; Homo sapiens (HsXRCC3), AF035586 |
The gene contains an Open Reading Frame (ORF) of 915 bp encoded a 304 amino acid polypeptide. Inspection revealed that amino acids 46-53 are characteristic P-loop for ATP/GTP binding site motif A present in XRCC3 homolog from H. sapiens (Fig. 1). The consensus for the P-loop is A or G with any four nucleotide base interval followed by GKT.
The deduced amino acid sequence of At XRCC3 (304 amino acids) is most similar to the human counterparts (Fig. 2). XRCC3 amino acid sequence between Arabidopsis and human shows that there are 35% identity and 46% of similarity with similar size. Clustering the RAD51 related genes using ClustalW in Arabidopsis, human and yeast, there are high affinities between AtXRCC3 and HsXRCC3 on the phylogenic tree (Fig. 2).
MMS resistance: Methylmethane Sulfonate (MMS) induces DNA damage through DSB. Cells do not survive if the damage is not properly or efficiently repaired. We can, therefore, predict that overexpression of DNA repair proteins, such as XRCC3, would result in an increased resistance to DNA damaging agents, such as MMS. The effect of expressing the XRCC3 gene on the resistance to DNA damage was investigated in a mutant yeast strain (LSY402) lacking RAD51. The results, shown in Fig. 3, indicated that the yeast strain containing pYES::XRCC3 exhibit enhanced resistance to MMS relative to yeast cells containing the control plasmid pYES2/CT.
Intracellular localization of XRCC3 in plants: A search for protein localization sites by PSORT program with XRCC3 amino acid sequence was resulted that XRCC3 would be localized in nucleus where is the site for DNA repair and recombination. Amino terminal region of XRCC3 protein contains high frequency (30%) of K and R residues, 6 over total 20 amino acids, which is a feature for higher possibility of being nuclear than cytoplasmic (Fig. 1). The classical Nucleus Localization Signal (NLS) is consisted of 4 basic amino acids or 3 basic amino acids with a H or P residue. The XRCC3 sequence does not contain the classical NLS but is rich in basic residues.
To verify the intracellular localization of XRCC3, A translational XRCC3::GFP fusion construct was produced with 5’ portions of the XRCC3 gene containing possible NLS region. The fusion construct was introduced into onion epidermis by particle bombardment. The results revealed that XRCC3 is localized into nucleus in plants. The GFP fluorescence is visible only in the nucleus after bombardment of XRCC3::GFP fusion (not shown). The control tissues bombarded with a control vector without fusion showed no green fluorescence.
 | |
| Fig. 2: | Phylogenetic tree Phylogenetic analysis of RAD51 family genes using ClustalW. Accession numbers for 15 deduced amino acids sequences used in this analysis are as follows: AtRAD51(NM122092), AtRAD51B(AB194809), AtRAD51C(AJ296174), AtXRCC2(AJ421043), AtXRCC3(AY032998), HsRAD51(BT019705), HsRAD51B(U84138), HsRAD51C(AY623112), HsRAD51D(AF034956), HsXRCC2(AF035587), HsXRCC3(AF035586), ScRAD51(M88470), ScRAD52(M10249), ScRAD55(U01144), ScRAD59(V53668) |
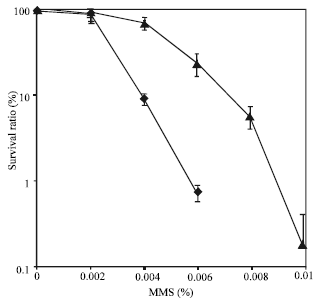 | |
| Fig. 3: | MMS sensitivity test after overexpression of XRCC3 in yeasts Each transformants with pYES::XRCC3 (closed triangle) and with pYES2/CT (closed circle) were measured MMS sensitivity by survival tests in log scale of Y-axis. The mean points from three independent experiments were presented |
DISCUSSION
I have sequenced and characterized a gene encoding the XRCC3 from A. thaliana, a flowering plant. The AtXRCC3 sequence displays significant homology with other RecA/RAD51 sequences from many organisms, including E. coli, S. cerevisiae and H. sapiens. Also the deduced amino acid sequence of AtXRCC3 is highly similar to other XRCC3 proteins in animals. The genomic sequence of XRCC3 is identical to a previously reported sequence by Arabidopsis Genome Project (Arabidopsis Genome Initiative, 2000). AtXRCC3 shows complementation to MMS sensitivity in the mutant cells of yeasts lacking RAD51. The yeast strains containing pYES::XRCC3 exhibit enhanced resistance to MMS relative to the yeasts bearing only the control plasmid pYES2/CT. XRCC3 functions in recombination (Lio et al., 2004) and in the DSB repair pathway by interacting to DNA repair proteins such as RAD51C to DNA damage sites (Brenneman et al., 2002; Kurumizaka et al., 2003). The damaged DNA sites are repaired by recombinational repair with the damaged termini being converted to 3'-OH or 5'-P residue by RAD51 recombinase (Henry-Mowatt et al., 2003). In the second phase of the reaction, the gap is filled by DNA ligase. The function interacting the key proteins such as RAD51, XRCC3 or XRCC4 would enhances efficiency in DSB repair. These reasons would explain why AtXRCC3 complements to a RAD51 mutant in yeasts. The genes related to HR repair mechanism such as XRCC3 would have significant roles in cell division in flowers which are the site occurred recombination during fertilization in plants. Recently a report presented that a knockout Arabidopsis line for AtXRCC3 showed sterile not to set seeds (Bleuyard and White, 2004).
ACKNOWLEDGMENTS
I wish to thank Dr. Eric B. Kmiec’s laboratory in Delaware Biotechnology Institute, DE, USA and Dr. Sang-Gu Kim’s laboratory in Seoul National University, Seoul, Korea for providing many helpful suggestions and strains. This study was supported by a grant from Korea Research Foundation (KRF-2002-C00275) Republic of Korea.
REFERENCES
- Britt, A.B., 1996. DNA damage and repair in plants. Ann. Rev. Plant Physiol. Plant Mol. Biol., 47: 75-100.
PubMed - Henry-Mowatt, J., D. Jackson, J.Y. Masson, P.A. Johnson and P.M. Clements et al., 2003. XRCC3 and Rad51 modulate replication fork progression on damaged vertebrate chromosomes. Mol. Cell, 11: 1109-1117.
Direct Link - Liu, L, S. Cheng, A.J. van Brabant and E.B. Kmiec, 2002. Rad51p and Rad54p, but not Rad52p, elevate gene repair in Saccharomyces cerevisiae directed by modified single-stranded oligonucleotide vectors. Nucleic Acids Res., 30: 2742-2750.
Direct Link - Shinohara, A. and T. Ogawn, 1995. Homologous recombination and the roles of double-strand breaks. Trends Biochem. Sci., 20: 387-391.
Direct Link - Arabidopsis Genome Initiative, 2000. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature, 408: 796-815.
CrossRefPubMedDirect Link








