Case Study
Probing the Single Nucleotide Polymorphism (Snp) of Swine PPAR Delta Gene
Department of Biology Science, Fuyang Normal College, 236237, Fuyang, People`s Republic of China
PPARs (Peroxisome proliferator-activated receptors) are a kind of transcription factors belonging to the Nuclear Receptors (NRs) super-family. When PPARs are activated, they will affect and/or regulate the transcription of many genes controlling vital physiological processes. Whole body metabolism related tissues, such as skeletal muscle, adipose tissue, heart and liver, are prone to get inflammation in metabolic disturbance. It was pointed out that PPARs had diverse functions and wide distributions serving as important links between lipids, metabolites and innate immunity (Hutter et al., 2013; Kruger et al., 2010). Herein, PPARs were considered as effective targets of drug remedy blocking obesity, diabetes and other metabolic disease.
PPARs play key roles in the metabolic syndrome and overall health of organisms including regeneration of tissues, adipocyte differentiation, lipid and glucose metabolism and immune response (Nagy et al., 2012). From a nutritional viewpoint, the PPARs are of importance because of their ability to be activated by long chain fatty acids and their metabolites (Kruger et al., 2010). In addition, several evidences showed the important role of PPARs in reproductive organs (Martinez et al., 2011; Shalom-Barak et al., 2004). Therefore, PPARs are recognized as candidates in order to improve metabolism and health and pregnancy through suitable diet.Up to date, there are three well-known PPARs subtypes, i.e., PPARa, PPARα (alias PPARβ) and PPARγ (classified as three isoforms in human, i.e., PPARγ1, PPARγ2, PPARγ3) (Asami-Miyagishi et al., 2004; Hutter et al., 2013; Matsuda et al., 2013; Schoonjans et al., 1996). Each subtype of PPARs is a product of a separate gene with a distinct tissue specific distribution and distinct functions. Among these subtypes, PPARα is predominantly expressed in the heart, kidney and liver, etc., mainly involving in fatty acid oxidation; PPARγ is mainly associated with adipose tissue,in which it controls adipocyte differentiation and insulin sensitivity; PPARδ is the only subtype currently untapped as candidate genes in metabolism and health and pregnancy research (Barak et al., 1999; Lehrke and Lazar, 2005; Lockyer et al., 2010; Martinez et al., 2011; Reilly and Lee, 2008; Robinson and Grieve, 2009; Wagner and Wagner, 2010). PPARδ is regarded as a member of nuclear hormone receptors super-family intimately regulating the expression of myriad genes involving in cell differentiation, apoptosis, energy metabolism and inflammation; It was found expressed in most metabolically active tissues, such as adipose and skeletal muscle tissues (Castillero et al., 2013; Kruger et al., 2010; Reilly and Lee, 2008; Wang et al., 2003). However, its fuction has not been clearly defined yet. On the other hand, improvement in reproductive traits (e.g., litter size), are of interest to swine producers and breeders (Johnson et al., 1999; Rothschild et al., 1996; Wang et al., 2013a). However it is frequently difficult to make genetic selection to improve the quality and quantity of animal reproductive traits due to low heritability and/or sex-linked inheritance pattern (Johnson et al., 1999; Rothschild et al., 1996). PPARs may play key roles in linking lipid and glucose metabolism and reproduction systems. Previous researches revealed that PPARγ is associated with body conditions, reproduction hormones and their receptor expression. The PPARγ/RXRα signaling was proved important in placenta, cytotrophoblast and cell fusions (Asami-Miyagishi et al., 2004; Batista et al., 2012; Hutter et al., 2013; Matsuda et al., 2013; Wang et al., 2013b). It is now clear that PPARs are important in the control of placental development. Nevertheless, unlike PPARα and PPARγ, little is known about the detailed roles of PPARδ gene in placental development and pregnancy nutrient regulation. In view of PPARδ as an important nuclear transcription factor implicated in adipocyte and myocyte differentiation, lipid and glucose metabolism, skeletal muscle wasting remedy and pregnancy in domesticated animals, we had designed animal experiments to investigate the function of PPARδ gene. In this study, a Single Nucleotide Polymorphism (SNP) corresponding to C→T substitution was detected and reported in the second intron of swine PPARδ gene locus in two swine strains, Yarkshire and Landrace.
Sampling and PCR-SSCP amplification: In total, 30 sows and 30 boars of Yarkshire strains, 22 sows and 20 boars of Landrace strains, were used in the study. Pieces of ear tissues were sampled and genomic DNA was extracted according to the manufacturers’ protocol. PCR primers were designed to detect SNP of porcine PPARδ gene locus (Table 1). The following SSCP analysis was employed based on the complete coding region and the reported 5'-regulator region of two published porcine PPARδ mRNA sequences (GenBank accession No. DQ437886, AY188501.1).
PCR amplifications were carried out in a eppendorf tube with a designed reaction system. The reaction system was a mixture being composed of multiple components (100-500 ng of genomic DNA, 2.5 μL of 10xPCR buffer, 200 μM of each dNTP, 10 pM of each primers, 2U of Taq DNA polymerase and sterile double-distilled water) with a total volume of 25 μL reaction mixture. The 10xPCR buffer contains 100 mM Tris-HCl (pH = 8.0), 500 mM KCl, 10 mM of MgCl2 and 0.1% glutin. Following an initial denaturation at 95°C for 5 min, 35 cycles of 1min denature at 94°C, 30 sec annealing at annealing temperature, 30 sec synthesis at 72°C, with a final cycle of 7 min extension at 72°C (Table 1). The amplified mixture was denatured 10 min at 98°C and then cooled on ice for 5 min with 2 μL of the PCR product and 8 μL of the loading buffer. Genetic polymorphism was detected by Single Strand Conformation Polymorphism (SSCP) in agarose gel electrophoresis and polyacrylamide gel electrophoresis, respectively. Final DNA band patterns were detected by silver staining.
Polyacrylamide gel electrophoresis: A total volume of 1.5 μL PCR product was transferred in the eppendorf tube, mixed with 6 μL gel loading solution containing 98% formamide, 0.025% bromophenol blue, 0.025% xylene cyanol, 20 mmol L-1 EDTA (pH = 8.0) and 10% glycerol. The reaction mixture was centrifuged and denatured at 98°C for 10 min, followed with a chill on ice for 5 min and loaded on 10-12% neutral polyacrylamide gels (acrylamide: bisacrylamide = 29:1). Polyacrylamide gel electrophoresis was performed in 1xTris borate-EDTA buffer (pH = 8.3) at 9-15 V cm-1 for 14-16 h at 4°C. Finally, polyacrylamide gels were stained with silver nitrate to identify SNP mutation.
Cloning and sequencing: After accomplishing the runs of polyacrylamide gel electrophoresis, PCR amplifications of different homozygous genotypes were separated on 0.7-1.0% agarose gels and recovered using geneclean II kit (Promega). Each DNA fragment was ligated into the pGEM-T easy vector (Promega) according to the manufacturer’s protocol at 4°C overnight. The ligation reactions were carried out in a 5 μL reaction mixture containing 1.5 μL of PCR product, 0.5 μL of pGEM-T vector (50 ng μL), 0.5 μL of T4 ligase (3 U μL-1 and 2.5 μL of 2xligation buffer, according to the protocol's instruction. Then recombinant plasmids were transformed into Escherichia coli DH5α competing cells. Positive clones of transformed cells were identified by restriction enzyme digestion. Final target clones of each homozygous genotype were sequenced from both directions by Shanghai Invitrogen Biotechnology Co. Ltd.
| Table 1: | Information of primer sequences |
 | |
RESULTS AND DISCUSSION
Genomic DNA extracted: Genomic DNA extracted from ear tissues was dissolved at room temperature for 24 h. Before carrying out PCR-SSCP analysis, the quality of genomic DNA should be checked with 1 μL sample on the 0.7-1.0% agarose gel electrophoresis. Figure 1 showed that the majority of genomic DNA had clear bands and was suitable for PCR-SSCP amplification.
SNP mutation detected in porcine PPARδ gene: Two pairs of primers were designed for PCR-based SSCP analysis of PPARδ gene. There was a mutation C→T at 107 bp of the PCR amplified fragment of PPARδ gene 5'-regulator region (GenBank accession No: AY188501.1). Allele gene A corresponded to base ‘C’; allele gene B corresponded to base ‘T’. The representative SNP sequencing output for homozygote AA or BB, heterozygote AB individuals is shown in Fig. 2b.
After reading and distinguishing heterozygotes from homozygotes of PPARδ genotypes in the PCR-SSCP analysis it was found that there was little polymorphism in the PCR-SSCP product of primer pair P1 but primer pair P2. A mutation C→T was identified in the second intron of PPARδ gene with primer pair P2. Therefore, all the following analysis proceeded with the result of primer pair P2. According to international research reports and their naming rules (Komatsu et al., 2010; Msalya et al., 2009) the SSCP patterns in primer pair P2 were read as AA, AB and BB genotypes respectively (Fig. 2). Every individual was tested in the sampled populations of Yarkshire strains and Landrace strains and we found both heterozygotes and homozygotes with PCR-SSCP analysis. Though the distribution of genotype frequencies was not strictly following the rules of Hardy-Weinberg equilibrium, we counted the numbers of genotypes and calculated corresponding gene frequency. There were 8 individuals genotyped as BB and 7 individuals genotyped as AB in Yarkshire strains while there were 5 individuals genotyped as BB and 9 individuals genotyped as AB in Landrace strains. The gene frequency of B was calculated as 0.1333) in Yarkshire strains and 0.3450 in Landrace strains, whereas the gene frequency of A was calculated as 0.8667 in Yarkshire strains and 0.6550 in Landrace strains according Hardy-Weinberg equilibrium.
The genetic polymorphisms of swine PPARδ gene and its Untranslated Region (UTR) were rarely reported. On the contrary, polymorphisms at the PPARα gene loci have been frequently identified (Bener et al., 2013; Domenici et al., 2013; Maciejewska-Karlowska et al., 2013; Motavallian et al., 2013; Sahmani et al., 2013; Wang et al., 2011, 2013a, b; Yang et al., 2013; Zhang et al., 2013).
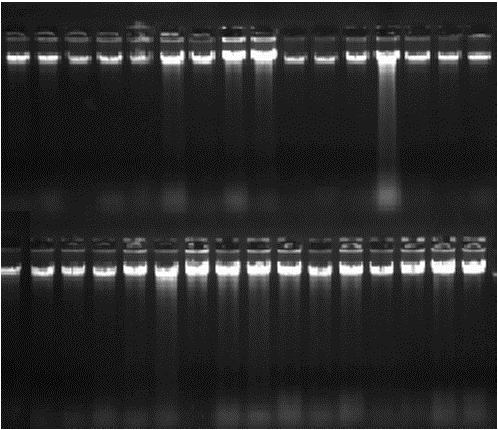 | |
| Fig. 1: | Genomic DNA extracted |
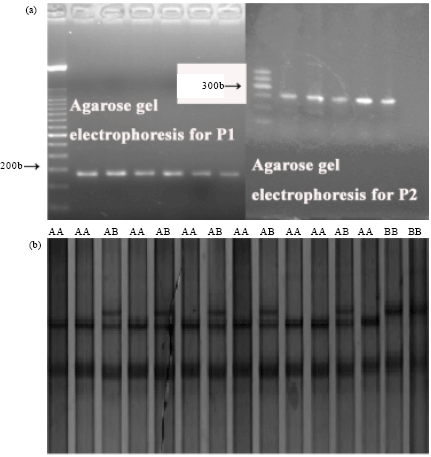 | |
| Fig. 2(a-b): | PCR-SSCP analysis for the swine PPARδ gene (a) Agarose gel electrophoresis for primers P1 and P2 and (b), Polyacrylamide gel electrophoresis for primers P2 (i.e., genotypes of the SSCP products for primers P2) |
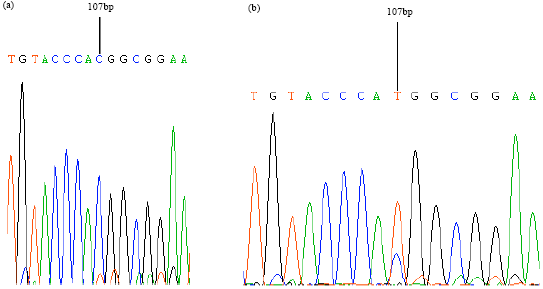 | |
| Fig. 3(a-b): | Sequencing result of SNP (single nucleotide polymorphism) in PPARδ gene (a) Sequencing for genotype AA and (b) Sequencing for genotype BB |
Among these reports, Wang et al. (2011) found two SNP mutations in swine PPARβ gene were polymorphic and significantly correlated with litter sizes (Wang et al., 2011) (Fig. 3). They also performed a few experiments to test whether other candidate genes were correlated with swine litter sizes (personal communications). In the present study, for the first time, a SNP mutation was identified in the second intron of PPARδ gene and no mutation in the coding region was detected. The deficiency in the present study was mainly the lack of the correlation analysis of PPARδ gene mutation genotype frequencies associated with the corresponding data of swine reproductive traits. As the PPARδ gene was considered as a potential genetic marker associated with growth and reproductive traits in domesticated animals, further enlarged experiments are required for the investigation on the specific SNP mutation of PPARδ gene and its association with specific phenotype traits.
It was revealed that the role of PPARs serving as vital metabolic sensors controlling cell functions including many essential cellular and physiological processes in organisms ranging from yeast to mammals. PPARδ is the only subtype of PPARs that is currently not a candidate or target gene in metabolism and health and pregnancy research. In the study, we designed two pairs of primers for Single Strand Conformation Polymorphism (SSCP) analysis based on complete coding region and the 5’- flanking region of PPARδ gene and found a SNP mutation C→T in the second intron. This mutation could provide a potential genetic marker for swine breeding. The SNP mutation and polymorphic information from this experiment would be useful for further research in medicinal genetics and animal breeding of PPARδ gene. Future research (both in vivo and in vitro) should be designed to identify novel markers and dissect the functions of PPARδ gene for growth and reproductive traits, as well as the progression of lipid and glucose metabolism and metabolic disorders. In our next experiment it will be designed and carried out to detect whether positive associations of PPARδ polymorphisms are associated with lipid and/or glucose content and relevant genetic data. According to the present study and previous reports, PPARδ could be considered as a candidate or target gene for swine reproductive traits.
We are grateful to the anonymous reviewers for the suggestions. The partial financial support of the following programs is also greatly acknowledged. This study is supported by National Natural Science Foundations of China (No. 31301965), Anhui Provincial Natural Science Foundations (No. 1308085QC63), Programs of Anhui Provincial Educational Commission Natural Science Foundation (No. KJ2012A216, No. KJ2013B198), National Statistical Scientific Research Project (No. 2013LY051) and the Fuyang Normal College Educational Project (No. 2012JYXM71).