ABSTRACT
Banana Bunchy Top Nanovirus (BBTV), the causal agent of Banana Bunchy Top Disease (BBTD) is the most serious viruses infecting banana allover the world as well as in Egypt. Polymerase Chain Reaction (PCR) as a molecular tool was used for detection of BBTV in banana samples showed the characteristic symptoms of BBTV as well as in symptomless banana samples. Two oligonucleotides belonging to the BBTV-component 2 resulted the full length were used as primers at 50°C annealing temperature. DNA was extracted from the tested samples and three DNA concentrations, i.e., 50, 100 and 150 ng were used as templates. A PCR fragment with a size of 1056 bp was amplified in case of virus-infected banana samples. As an interesting result, the tested samples were varied in their PCR results according to the DNA concentrations and this could be reflect the low concentration of the viral DNA with the total DNA extracted from the virus-infected samples. Furthermore, the presence of the BBTV-DNA was also detected in the total DNA extracted from the viruliferous banana aphid (Pentalonia nigronervosa Coq). The results also confirmed the use of PCR for BBTV detection in particularly the coast was not limited.
PDF Abstract XML References
How to cite this article
DOI: 10.3923/biotech.2007.53.56
URL: https://scialert.net/abstract/?doi=biotech.2007.53.56
INTRODUCTION
In Egypt, the cultivated area of bananas increased to 60000 Feddans with total production of 1032000 tons (Anonymous, 2004). Production of bananas is limited in areas with fungi, viruses or nematodes and these pathogens are increasing seriously in the banana cultivated regions of the world (Dale, 1987). BBTV as a phloem-limited virus used to occur in low concentrations in banana-infected plants (Harding et al., 1991, 2000). The early detection of BBTV, either in the symptomless young plants or in micropropagated plant materials, plays an important role in its control (Dale, 1987; Sadik et al., 1997; Harding et al., 2000). Banana bunchy top nanovirus (BBTV) is an isometric virus particles (18-20 nm) consists of at least six components of cicular ssDNA, each about 1 kb (Burns et al., 1995). The virus has a single protein with Mr of about 20-21 kDa (Dietzgen and Thomas, 1991; Harding et al., 1991; Sadik et al., 1999) and naturally transmitted by a banana aphid (Pentalonia nigronervosa, Dale, 1987). Recently, the detection of BBTV is now available via double antibody sandwich-enzyme linked immunosorbent assay (DAS-ELISA) (Othman et al., 1996; Sadik et al., 1997). The PCR was successfully used for detecting several plant viruses including BBTV (Xie et al., 1994; Harding et al., 2000; Abdel-Hamid et al., 2003). Wetzel et al. (1991) reported that the importance of PCR lies in its ability to amplify specific DNA or cDNA sequences, from as short as 50 bp to over 10000 bp in length, more than a million fold in a few hours. Therefore, this study was designed to use the PCR for detecting the BBTV in banana samples showed the characteristic symptoms of BBTV as well as in symptomless banana samples and banana aphid.
MATERIALS AND METHODS
Sample source: A number of 15 samples of banana plants showed the characteristics symptoms of BBTV and 10 symptomless banana samples were collected from the open field area at Qalubia Governorate, Egypt.
Banana aphid: Ten adult groups of banana aphid (Pentalonia nigronervosa Coq.) fed on BBTV-infected and symptomless banana suckers (five groups for each) under open field conditions at Qalubia Governorate, Egypt were collected.
DNA extraction: According to the method of Dellaporta et al. (1985) and Abdel-Hamid et al. (2003) the DNA was extracted and purified from BBTV-infected and symptomless plant samples. The final pellets were resuspended in a volume of about 50 μL of sterile TE buffer, pH 8.0 (Sambrook et al., 1989). The concentration of the purified DNA was determined as recommended by Abdel-Hamid et al. (2003). The DNA was extracted from 10 groups of adults (10 adults for each group) fed and BBTV-infected and symptomless banana plants as reported by Abdel-Hamid et al. (2003).
PCR detection: This work was done in Agriculture Genetic Engineering Research Institute (AGERI ), Cairo, Egypt (2002).
In a volume of 50 μL as described by Harding et al. (2000) and Abdel-Hamid et al. (2003) the PCR was conducted for virus detection. Two primers, i.e., P1: 5’TAC AAG ACG CTA TGA CAA ATG TAC G3’ and P2: 5’TTT CAT AGC CTG TTG TAT TTC GTT CCA3’ designed based on the nucleotide sequences of BBTV-DNA-2 (Sadik, 1994) were kindly provided by Prof. Dr. Atef S. Sadik, Faculty of Agriculture, Ain Shams University, Egypt). The reaction was performed in a Perkin-Elmer/DNA Thermal Cycler 480 (Norwalk, CT) for 40 cycles after initial denaturation for 5 min at 94°C. Each cycle consisted of denaturation at 9°C for 1 min, annealing temperature at 50°C for 1 min and extension at 72°C for 2 min. The final primer extension cycle was extended to 10 min.
DNA electrophoreses: The amplified products were resolved by electrophoresis in a 1.2% agarose at 80 volts for 1 h with TAE buffer (Sambrook et al., 1989). DNA was visualized by staining gel in ethidium bromide (0.5 μg mL-1) and photographed under UV light using a Polaroid camera.
RESULTS AND DISCUSSION
A large number of literatures described the development of PCR primer pairs and of very sensitive assays for essentially all groups of viral pathogens (Wetzel et al., 1992; Hadidi et al., 1993; Shamloul et al., 1995).
Mullis and Faloona (1987) showed that PCR is one of the most sensitive molecular methods for pathogen detection currently available. The PCR process uses a series of hot, cold and warm cycles. Hot cycles split DNA into single strands. Cold cycles allow the primers to attach. During warm cycles, the enzyme makes a copy of each piece of primed DNA. If a virus is present, PCR will produce a large quantity of a certain piece of DNA. We use a process called electrophoresis that separates pieces of DNA by size to tell if that piece of DNA is present. Several investigators (Levy and Hadidi, 1991; Smith et al., 1992; Hu et al., 1993; Sadik et al., 1997; Harding et al., 2000; Abdel-Hamid et al., 2003) reported that new molecular methods have become available for studying the plant viruses.
In this study, PCR as a molecular tool was used for detection of BBTV in banana samples showed the characteristic symptoms of BBTV as well as in symptomless banana samples. Two oligonucleotides belonging to the BBTV-component 2 resulted the full length were used as primers. Sadik (1994) used the same two oligonucleotides as primers for amplification of a fragment with a size of about 1056 bp from the BBTV-DNA-2. DNA was extracted from the tested samples and three DNA concentrations, i.e., 50, 100 and 150 ng were used as templates. Results in Table 1 and Fig. 1 the tested samples were varied in their PCR results according to the DNA concentrations. As a PCR fragment with a size of 1056 bp was amplified in case of virus-infected banana samples with 100% when the BBTV was detected using 150 ng DNA per reaction. While, 73.3 and 20% of the BBTV-infected samples gave positive PCR results when 100 and 50 ng DNA per reaction were used as templates, respectively. This result could be reflecting the low concentration of the viral DNA with the total DNA extracted from the virus-infected samples. Similar results reported by Abdel-Hamid et al. (2003), they recommended that 125 ng of banana samples were suitable enough for BBTV detection.
In case of symptomless banana plants, PCR proved the presence of BBTV in the DNA extracted from 30% of the tested samples using 150 ng DNA per reaction. This result agree with that found by Harding et al. (2000) and Abdel-Hamid et al. (2003), they pointed to the importance detection of BBTV as early step for virus control in particularly in the plants produces via tissue culture as well as symptomless plants.
| Table 1: | PCR detection of BBTV in BBTV-infected and symptomless banana samples using two primers belonging to BBTV-DNA-2 |
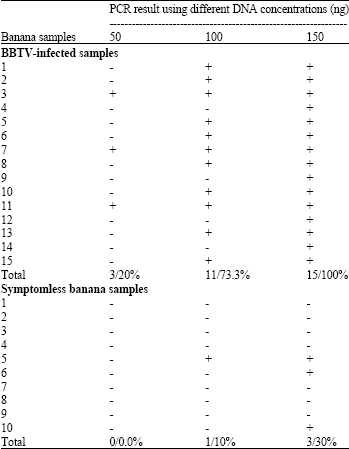 | |
| -: Negative, +: Positive | |
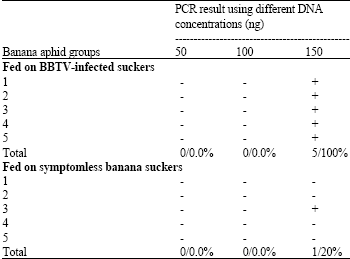
| Table 2: | PCR detection of BBTV in banana aphid (P. nigronervosa Coq.) fed on BBTV-infected and symptomless banana suckers using two primers belonging to BBTV-DNA-2 |
 | |
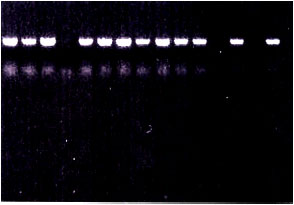 | |
| Fig. 1: | PCR detection of BBTV in 100 ng of DNA extracted from banana plants naturally exhibited BBTV characteristic symptoms using agarose gel (1.2%) in TAE buffer stained with ethidium bromide. N: negative control (PCR mixture without any DNA template). A PCR fragment with a size of about 1056 bp was amplified |
The presence of the BBTV-DNA was also detected in the total DNA extracted from the viruliferous banana aphid (P. nigronervosa Coq). Results in Table 2 and Fig. 2 and 3 showed that BBTV was successfully detected when the DNA was extracted from 10 adults of banana aphid either fed on BBTV-infected (100%) or fed on symptomless (20%) banana suckers and 150 ng per reaction were used as a template for PCR amplification. On the other hand, no PCR fragments were obtained when 50 and 100 ng DNA were used. The results confirmed the importance of the use of PCR for BBTV detection in particularly the coast was not limited. Similar results were obtained by Shamloul et al. (1995), Hu et al. (1996), Sadik and Gad-El-Karim (1997) and Abdel-Hamid et al. (2003).
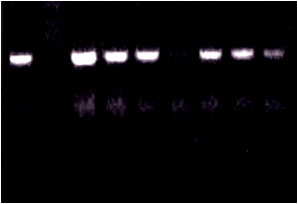 | |
| Fig. 2: | PCR detection of BBTV in 100 ng of DNA extracted from symptomless banana plants using agarose gel (1.2%) in TAE buffer stained with ethidium bromide. N: negative control (PCR mixture without any DNA template). A PCR fragment with a size of about 1056 bp was amplified |
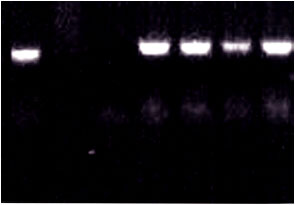 | |
| Fig. 3: | PCR detection of BBTV in 150 ng of DNA extracted from banana aphid fed on BBTV-infected banana plants (Lanes) using agarose gel (1.2%) in TAE buffer stained with ethidium bromide. N: negative control (PCR mixture without any DNA template). A PCR fragment with a size of about 1056 bp was amplified |
Furthermore, PCR tests are more sensitive than ELISA tests (Wetzel et al., 1991; Sadik et al., 1997) and are better suited to plant tissue that is likely to have lower numbers of pathogens, such as tissue culture.
ACKNOWLEDGMENTS
We would like to thank Prof. Dr. Atef S. Sadik, Department of Agric. Microbiology, Faculty of Agric., Ain Shams Univ., for his endless help to accomplish such work.
REFERENCES
- Abdel-Hamid, I.A., E.E. Wagih, H.T. Mahfouz and A.S. Sadik, 2003. Non-radioactive detection of banana bunchy top nanovirus. Arab. J. Biotechnol., 6: 329-338.
Direct Link - Burns, T.M., R.M. Harding and R.M. Dale, 1995. The genome organization of banana bunchy top virus: Analysis of six ssDNA components. J. Gen. Virol., 76: 1471-1482.
CrossRefDirect Link - Dale, J.L., 1987. Banana bunchy top: An economically important tropical plant virus disease. Adv. Virus Res., 33: 301-325.
PubMed - Hu, J.S., M. Wang, D. Sether, W. Xie and K.W. Leonhardt, 1996. Use of polymerase Chain reaction (PCR) to study transmission of banana bunchy top virus by the banana aphid (Pentalonia nigronervosa). Ann. Applied Biol., 128: 55-64.
Direct Link - Mullis, K.B. and F.A. Faloona, 1987. Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction. Methods Enzymol., 155: 335-350.
CrossRefPubMedDirect Link - Sambrook, J., E.F. Fritsch and T.A. Maniatis, 1989. Molecular Cloning: A Laboratory Manual. 2nd Edn., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA., ISBN-13: 9780879695774, Pages: 397.
Direct Link - Smith, G.R., R.V. Velde and J.L. Dale, 1992. PCR amplification of a specific double-stranded RNA region of Fiji disease virus from diseased sugarcane. J. Virol. Methods, 39: 237-246.
PubMed - Wetzel, T., T. Candresse, G. Macquaire, M. Ravelon and J. Dunez, 1992. A highly sensitive immunocapture polymerase chain reaction for plum pox potyvirus detection. J. Virol. Methods, 39: 27-37.
Direct Link








