ABSTRACT
Turkey is homeland to many species geophyte also some of them are endemic. This study aimed to do comparison of different geophyte species DNA quantity and purity collected from the flora of Turkey. The isolation of pure and high yields of genomic DNA is essential for many molecular biology studies. In addition, one of the objective projects was preservation of endemic species for pure DNA required for the future. Genomic DNA was extracted from Colchicum, Lilium, Polygonatum, Iris and Nectaroscordum species using modified QIAGEN DNeasy Plant Mini Kit. The purity of isolated genomic DNA was confirmed by spectrophotometric and agarose gel analyses. The results showed that the modified protocol almost successfully produced a sufficient amount of DNA with high quality. Iris, Lilium, Polygonatum and Nectaroscordum species of high purity and sufficient amounts of DNA were obtained with the kit. However, from some species of Colchicum, sufficient purity and quantity of DNA could not be obtained.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajps.2014.98.110
URL: https://scialert.net/abstract/?doi=ajps.2014.98.110
INTRODUCTION
Ecology of Turkey is very favorable for geophytes. They have an important place in history of the country. Unfortunately this wealth could not be evaluated fully for many years. Geophytes (bulbs, tubers and rhizomes plants) have a very important place in flora of Turkey. In recent years, a comprehensive projects was generated on geophytes of Turkey. Turkey exports geophytes for horticultural purposes mainly collected from the wild. The amount of material exported from Turkey has increased each year. Several flowering plant families contain species which have underground storage organs such as bulbs, corms and tubers, these plants are used in gardening, cut-flower and flowering pot plant production as geophytes. Geophytes of Turkey are usually used to ornamental properties and very small part of them used for phytochemical content (Sener et al., 1998). The culture and production of corm and bulb plants has been progressed in parallel with the developments in biotechnology research in recent years (Ziv, 1997).
Genetic analyses of plant rely on high yields of pure DNA samples. Some plant species, DNA extraction is difficult because of contents of polyphenols and other secondary metabolites (Aljanabi et al., 1999; Zhang and Stewart, 2000; Dehestani and Tabar, 2007; Sahasrabudhe and Deodhar, 2010; Srivastava et al., 2010; Okpodu and Abdullah-Israel, 2011). All plants have different content and a single method of DNA isolation is not possible to do for all plants (Vural, 2009). Young leaves and partly opened leaves are preferable during the isolation of DNA. Because young leaves contain lesser amounts of polyphenols and other components which prevents pure DNA synthesis (Shankar et al., 2011).
During the DNA isolation protocols, especially higher quality and yield are two important factors. The DNA isolation protocols include some similar steps. The main purpose is to separate DNA from other compounds especially proteins, carbohydrates, phenols and other polysaccharides. After isolation of DNA it is important to determine purity and yield. Generally gel electrophoresis and spectrophotometer are preferred to determine purity of DNA. The DNA isolated in pure form is used in different molecular studies and different purposes (Channarayappa, 2007; Varma et al., 2007; Tiwari et al., 2012).
Isolation of plant DNA is basic requirement for genome identification, mapping, marker assisted selection and genetic engineering. The degree of purity and quality can vary from plant and isolation protocols. Many different methods and technologies are available for the isolation of genomic DNA. These techniques are based on the selection of different factors. It was asked to be especially high purity of the DNA. It is important for a short time and result in the desired manner. The DNA is usually recovered by precipitation using ethanol or isopropanol (Khan, 2003; Da Silva, 2005).
Many researchers began to use DNA isolation kit for easy application and quick results in recent years (Salem et al., 2006; Al-Saghir, 2009; Mirmomeni et al., 2010). Particularly, these kits are preferred if samples are many and time is limited. The QIAGEN DNeasy Plant Kit, one of which was used successfully in many plant species but the first time in this study were tested over a number of different geophyte genus and species.
In this study, it was tried to isolate pure DNA from Colchicum, Lilium, Polygonatum, Iris and Nectaroscordum species by using modified QIAGEN DNeasy Plant Mini Kit. Also the purity of isolated genomic DNA was confirmed by spectrophotometric and agarose gel analyses.
MATERIALS AND METHODS
Plant materials for DNA isolation: Turkey flora Iris, Colchicum, Lilium, Polygonatum and Nectaspordum species collected from different locations were used as material in the study. The 322 Iris, 252 Colchicum, 42 Polygonatum, 39 Lilium and 3 Nectaroscordum types were collected from flora of Turkey. Fresh leaves of these geophytes were used for pure DNA isolation.
Genomic DNA extraction: The basic steps of DNA isolation are disruption of the cellular structure to create a lysate, separation of the soluble DNA from cell debris and other insoluble material and purification of the DNA of interest from soluble proteins and other nucleic acids. The majority of DNA isolation systems are based on purification by silica now a days. Regardless of the method used to create a cleared lysate, the DNA of interest can be isolated by virtue of its ability to bind silica in the presence of high concentrations of chaotropic salts (Chen and Thomas Jr., 1980; Marko et al., 1982; Boom et al., 1990). These salts are then removed with an alcohol-based wash and the DNA eluted in a low-ionic-strength solution such as TE buffer or water. The binding of DNA to silica seems to be driven by dehydration and hydrogen bond formation which competes against weak electrostatic repulsion (Melzak et al., 1996). Hence, a high concentration of salt will help to drive DNA adsorption onto silica and a low concentration will release the DNA.
The separation of DNA from cellular components divided into four stages (Fig. 1):
| • | Disruption |
| • | Lysis |
| • | Removal of proteins and contaminants |
| • | Recovery of DNA |
 | |
| Fig. 1: | DNA isolation stages with QIAGEN DNeasy plant kit |
Distruption stage: The 1 g of fresh leaf was homogenized in a mortar using liquid nitrogen. The 100 mg powdered tissue was transferred to a 2 mL capacity of tubes.
Lysis stage: Four hundred microliter Buffer AP1 and 4 μL RNase A was added into the mixture. Vortex and incubate for 20 min at 65°C. Invert tube 4-5 times during incubation (do not mix Buffer AP1 and RNase A before use). Add 130 μL Buffer AP2. Mix and incubate for 5 min on ice.
Removal of proteins and contaminants: Centrifuge the lysate for 5 min at 20,000xg (14,000 rpm). Pipet the lysate into a QIAshredder Mini spin column in a 2 mL collection tube. Centrifuge for 2 min at 20,000xg (14,000 rpm). Transfer the flow-through fraction into a new tube without disturbing the pellet. Add 1.5 volumes of Buffer AP3/E and mix by pipetting. Transfer 650 μL of the mixture into a DNeasy Mini spin column in a 2 mL collection tube. Centrifuge for 1 min at 6000xg (8000 rpm). Discard flow-through. Repeat this step with the remaining sample. Place the spin column into a new 2 mL collection tube. Add 500 μL Buffer AW and centrifuge for 1 min at 6000xg. Discard flow-through. Add another 500 μL Buffer AW. Centrifuge for 2 min at 20,000xg (remove the spin column from the collection tube carefully so, the column does not come into contact with the flow-through).
Recovery of DNA: Transfer the spin column to a new 1.5 or 2 mL microcentrifuge tube and add 100 μL Buffer AE for elution. Incubate for 5 min at room temperature. Centrifuge for 1 min at 6000xg.
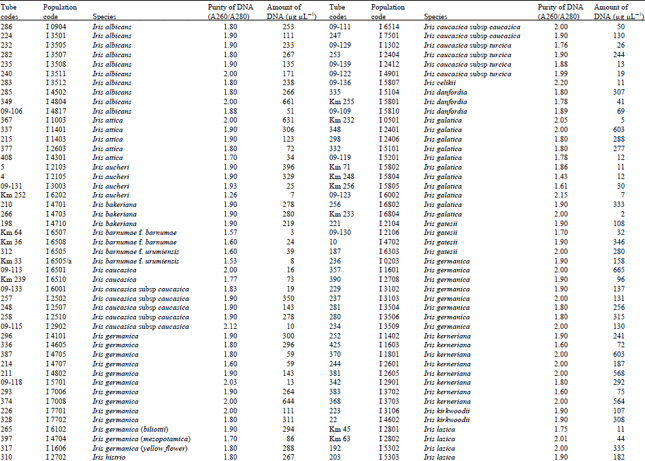
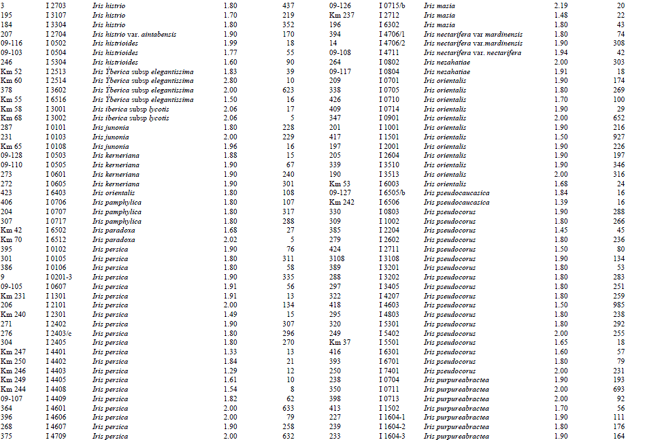
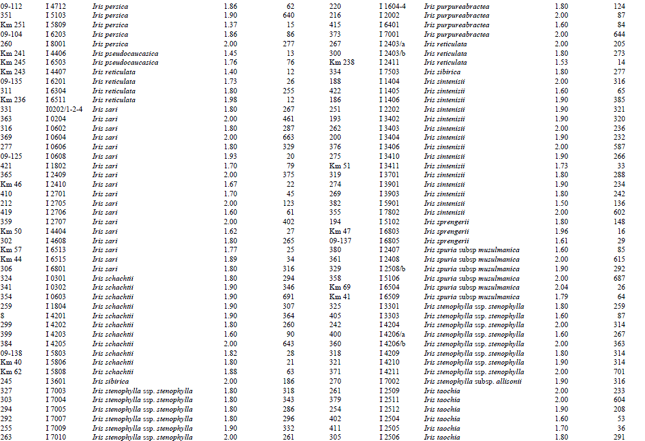
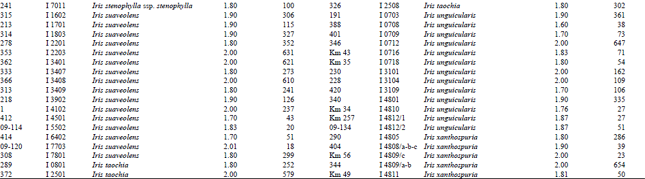
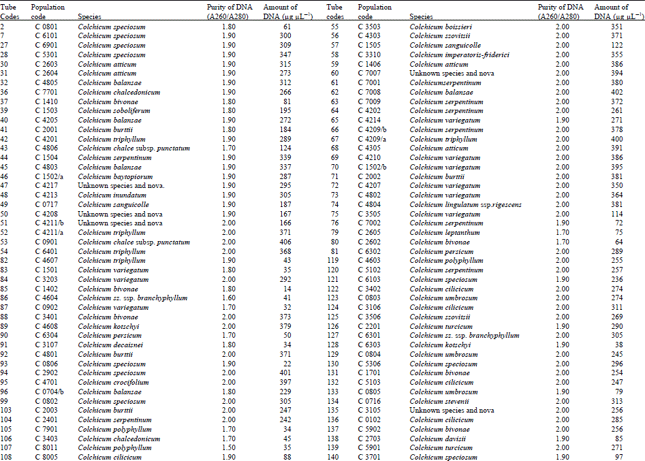
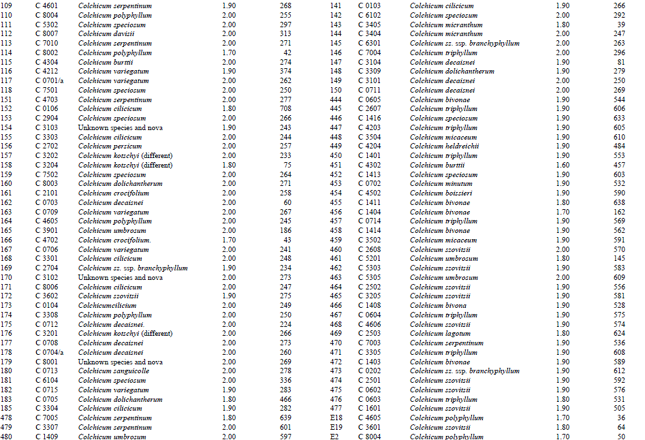
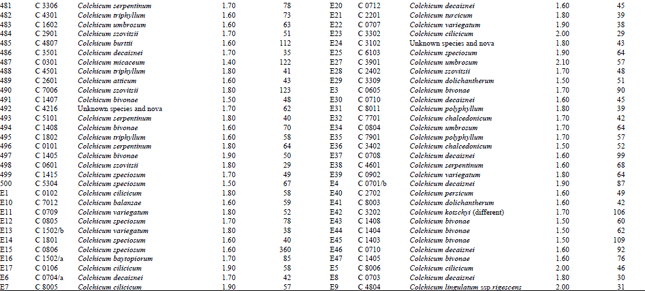
Measurement of amount and purity of DNA: The yield of DNA per gram of leaf tissue extracted was measured using a UV Spectrophotometer (Shimadzu Biospec-mini-Japan) at 260 nm. The purity of DNA was determined by calculating the ratio of absorbance at 260 nm to that of 280 nm (Qiagen Sample and Assay Technologies, 2006) (Table 1-4). DNA concentration and purity was also determined by running the samples on 1% agarose gel (Fig. 2).
RESULTS AND DISCUSSION
A reasonable amount of clean DNA from most of the geophytes with modified protocol of DNeasy Plant mini kit was yielded. The initial protocol was based on “DNeasy Plant Hand Book” (Qiagen Sample and Assay Technologies, 2006). When tables (Table 1-4) and agarose gel image (Fig. 2) analysed it was clearly seen that especially Iris species DNA were isolated in pure form and sufficient amounts. DNA isolation of 322 Iris populations were done successfully. Iris population included 40 species and 45 taxon. The highest amounts of DNA were obtained from the Iris pseudocorus (population code I 4603) 985 μg μL-1. On the other hand the minimum amount of DNA were obtained from the Iris galactica (population code I 6804) 2 μg μL-1. DNA purity values were as follows, the highest value of 2.80 (Iris iberica ssp. elegantissima population code I 2,514 population) and the lowest 1.26 (Iris aucheri population code I 6202). However, DNA purity of most Iris species was in the range 1.7-1.9.
DNA isolation of 250 Colchicum populations was done successfully. Colchicum population included 36 known Colchicum spp. and 10 unkown Colchicum spp. The highest amounts of DNA were obtained from the Colchicum cilicicum Dammer (population code C 0106) 708 μg μL-1. On the other hand the minimum amount of DNA was obtained from the Colchicum bivonae (Population code C 1402) 14 μg μL-1. DNA purity values were as follows: the highest value of 2.10 from Colchicum umbrosum (population code C 3901) and the lowest 1.40 from Colchicum micaceum (population code C 0301). However DNA purity of most Colchicum species was in the range 1.7-2.0. When agarose gel and spectrophotometric results were examined it was realized that some of Colchicum species quantity and purity of DNA was not at the desired levels. While it was isolated sufficient quantities and purity of DNA from majority of Colchicum species but in some Colchicum species could not be isolated in sufficient quantities and purity of DNA.
 | |
| Fig. 2: | Electrophoresis of isolated DNA of geophytes on 1% agorose gel |
| Table 1: | DNA purity and amount of different Iris species |
    | |
| Table 2: | DNA purity and amount of different Colchicum species |
   | |
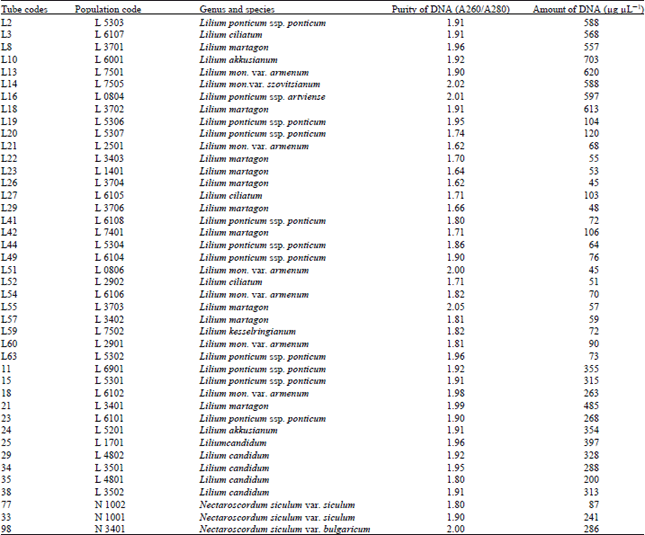
| Table 3: | DNA purity and amount of different Lilium and Nectaroscordum species |
 | |
It thought to be related with high level of antioxidant content and phenolic compounds of some Colchicum species. Sambrook and Russell (2001) mentioned that phenols and polysaccharides are compounds that prevent the DNA purity. Contamination by phenol which is commonly used in nucleic acid purification, can significantly throw off quantification estimates. Phenol absorbs with a peak at 270 nm and a A260/280 of 2. Nucleic acid preparations uncontaminated by phenol should have a A260/280 of around 1.2. Contamination by phenol can significantly contribute to overestimation of DNA concentration. Also Ahmad (2011) and Ebrahimzadeh et al. (2010) analysed some Colchicum species antioxidant activity, free radical scavenging activity and phenolics compounds. Colchicum luteum and Colchicum speciosum species reported in these studies that they have a high content.
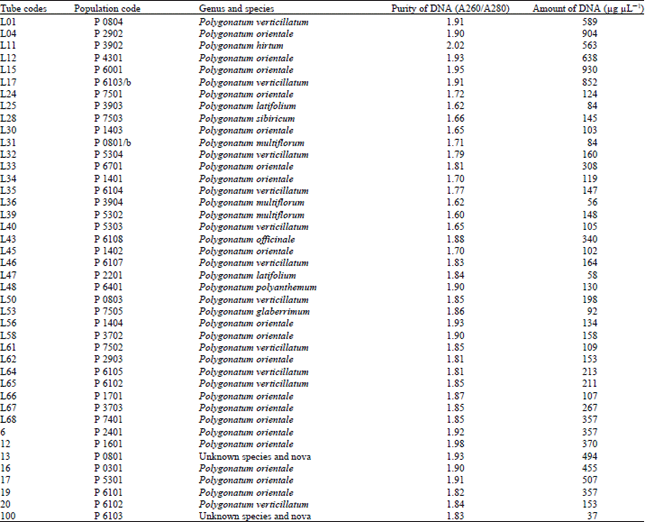
Similar results were obtained with Polygonatum, Lilium and Nectaroscordum species. The DNA isolation of 42 Polygonatum populations was done successfully. This population included 9 species. The purity of DNA values were obtained ranging between 1.60-2.02
The amounts of DNA values were obtained ranging between 37-904 μg μL-1 for Polygonatum species. The DNA isolation of 39 Lilium populations including 8 species was done successfully. The purity of DNA values were obtained ranging between 1.62-2.05 also the amounts of DNA values were obtained ranging between 45-620 μg μL-1 for Lilium species.
Van Tuyl and Boon (1996), reported the similar results in their study with different liliums. They reported that differences in DNA content between varieties and species.
| Table 4: | DNA purity and amount of different Polygonatum species |
 | |
DNA isolation of 3 including 2 species were also done successfully. The purity of DNA, respectively, 1.80, 1.90 and 2.00 also the amounts of DNA 87, 241 and 286 were obtained.
This study showed that commercial DNA isolation kit can be used successfully for geophytes DNA isolation. Mirmomeni et al. (2010) compared three different techniques for optimizing DNA extraction from Paraffin-Embedded Tissues (PETs) in order to generate DNA suitable for PCR. They mentioned that comparison of the electrophoresis pattern in all three methods including commercial DNA isolation kit (Qiagen) indicated that no significant differences were detected and all three methods showed equally efficient results.
CONCLUSION
This study is one of the most comprehensive studies with different species, cultivars and types of geophytes. In recent years, advances in genetics will help to provid the introduction of methods are needed. The isolation of pure and high yields of genomic DNA from different sources is essential for many molecular biology studies. Especially important to use methods common to different species also conclusion of these studies can help rapidly. In this study, it was compared different geophyte species DNA quantity and purity collected from the flora of Turkey.
Genomic DNA was extracted from Colchicum, Lilium, Polygonatum, Iris and Nectaroscordum species with the most widely used method of DNA isolation was modified and used successfully.
ACKNOWLEDGMENTS
The authors gratefully acknowledge the support of the researchers of the Atatürk Horticultural Central Research Institute. This study supported by grant (No.105G068) from the National Scientific and Technological Research Council of Turkey.
REFERENCES
- Aljanabi, S.M., L. Forget and A. Dookun, 1999. An improved and rapid protocol for the isolation of polysaccharide- and polyphenol-free sugarcane DNA. Plant Mol. Biol. Rep., 17: 1-8.
CrossRefDirect Link - Al-Saghir, M.G., 2009. Rapid and efficient method of genomic DNA extraction from pistachio trees (Pistacia vera L.). Res. J. Bot., 4: 70-73.
CrossRefDirect Link - Ahmad, B., 2011. Antioxidant activity and phenolic compounds from Colchicum luteum Baker (Liliaceae). Afr. J. Biotechnol., 9: 5762-5766.
Direct Link - Boom, R., C.J. Sol, M.M. Salimans, C.L. Jansen, P.M.E. Wertheim-van Dillen and J. van der Noordaa, 1990. Rapid and simple method for purification of nucleic acids. J. Clin. Microbiol., 28: 495-503.
PubMedDirect Link - Chen, C.W. and C.A. Thomas Jr., 1980. Recovery of DNA segments from agarose gels. Anal. Biochem., 101: 339-341.
CrossRef - Dehestani, A. and S.K.K. Tabar, 2007. A rapid efficient method for DNA isolation from plants with high levels of secondary metabolites. Asian J. Plant Sci., 6: 977-981.
CrossRefDirect Link - Ebrahimzadeh, M.A., S.M. Nabavi, S.F. Nabavi, F. Bahramian and A.R. Bekhradnia, 2009. Antioxidant and free radical scavenging activity of H. officinalis, L. angustifolius, V. odorata, B. hyrcana and C. speciosum. Pak. J. Pharm. Sci., 23: 29-34.
Direct Link - Marko, M.A., R. Chipperfield and H.C. Birnboim, 1982. A procedure for the large-scale isolation of highly purified plasmid DNA using alkaline extraction and binding to glass powder. Anal. Biochem., 121: 382-387.
CrossRef - Melzak, K.A., C.S. Sherwood, R.F. Turner and C.A. Haynes, 1996. Driving forces for DNA adsorption to silica in perchlorate solutions. J. Colloid Interf. Sci., 181: 635-644.
CrossRef - Mirmomeni, M.H., S.S. Majd, S. Sisakhtnezhad and F. Doranegard, 2010. Comparison of the three methods for DNA extraction from paraffin-embedded tissues. J. Biol. Sci., 10: 261-266.
CrossRefDirect Link - Khan, M.F., 2003. Evaluation of hexaploid wheat genotypes by using DNA isolation and gel-electrophoresis. Asian J. Plant Sci., 2: 212-215.
CrossRefDirect Link - Okpodu, C.M. and M. Abdullah-Israel, 2011. A DNA isolation protocol suitable for RAPD analysis from fresh or herbarium-stored leaves of a historic Quercus virginiana L. J. Plant Sci., 6: 77-87.
CrossRef - Shankar, K., L. Chavan, S. Shinde and B. Patil, 2011. An improved DNA extraction protocol from four in vitro banana cultivars. Asian J. Biotechnol., 3: 84-90.
CrossRefDirect Link - Sahasrabudhe, A. and M. Deodhar, 2010. Standardization of DNA extraction and optimization of RAPD-PCR conditions in Garcinia indica. Int. J. Bot., 6: 293-298.
CrossRefDirect Link - Salem, H.H., T.H. Huang, B.A. Ali and Q.D. Xie, 2006. Differentiation of Bacillus thuringiensis and Escherichia coli by the randomly amplified polymorphic DNA analysis. J. Applied Sci., 6: 1540-1546.
CrossRefDirect Link - Sambrook, J. and D.W. Russell, 2001. Molecular Cloning: A Laboratory Manual. 3rd Edn., Cold Spring Harbor Laboratory Press, New York, USA., ISBN-13: 9780879695774, Pages: 2344.
Direct Link - Srivastava, N., V. Sharma, B. Kamal, A.K. Dobriyal and V.S. Jadon, 2010. Polyphenolics free DNA isolation from different types of tissues of Aconitum heterophyllum wall-endangered medicinal species. J. Plant Sci., 5: 414-419.
CrossRef - Da Silva, J.A.T., 2005. Effectiveness of DNA extraction protocols for horticultural and physiological model plant analyses. Int. J. Bot., 1: 93-99.
CrossRefDirect Link - Sener, B., M. Koyuncu, F. Bingol and F. Muhtar, 1998. Production of bioactive alkaloids from Turkish geophytes. Pure Applied Chem., Vol. 70, No. 11.
Direct Link - Tiwari, K.L., S.K. Jadhav and S. Gupta, 2012. Modified CTAB technique for isolation of DNA from some medicinal plants. Res. J. Med. Plant, 6: 65-73.
CrossRefDirect Link - Varma, A., H. Padh and N. Shrivastava, 2007. Plant genomic DNA isolation: An art or a science. Biotechnol. J. Healthcare Nutr. Technol., 2: 386-392.
CrossRefPubMedDirect Link - Vural, H.C., 2009. Genomic DNA isolation from aromatic and medicinal plants growing in Turkey. Sci. Res. Essays, 4: 59-64.
Direct Link - Zhang, J. and J.M. Stewart, 2000. Economical and rapid method for extracting cotton genomic DNA. J. Cotton Sci., 4: 193-201.
Direct Link








