ABSTRACT
The study was conducted in the organic trial field of Wageningen University and Research Centre with the objective of how diversity with the composite cross populations of winter wheat evolves over the years? And which traits show most diversification? Four different aged composite cross population, one CCP-extra population and one pure line winter wheat cultivar were evaluated using a randomized complete block design with three replications. Significant differences were observed for the traits plant height, flag leaf spike length and width of the leaf next to the first leaf. For plant height and flag leaf spike length, all CCPs have higher values for the diversity index. There were no significant differences between the CCPs in values for the Shannon-Weaver diversity index, H'. Genotypes showed highly significant differences for the SD within plots for the traits plant height (p<0.001), flag leaf spike distance (p<0.001), width of the 1st leaf next to the 1st node (p<0.008) and width of the flag leaf (p<0.001).
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajps.2014.40.45
URL: https://scialert.net/abstract/?doi=ajps.2014.40.45
INTRODUCTION
Wheat is a cereal grass of the Graminae (Poaceae) family and of the genus Triticum is the world’s largest cereal crop. It has been described as the ‘king of cereals’ due to the area covered by wheat, high productivity and the prominent position it holds in the international food grain trade. Wheat (Triticum spp.) was domesticated 8000 years ago, it is one of the first cultivated cereal crops in all over the world. South-western Asia is believed as the origin of wheat (Acquaah, 2007). It can be used as food and feed (Acquaah, 2007; Hildermann et al., 2009; Prohens-Tomas et al., 2009).
The gene pool of wheat is large and it is known by its diversity among the cereal crops (Prohens-Tomas et al., 2009). The species of Triticum are grouped into three ploidy classes, which are diploid (2n = 2x = 14), tetraploid (2n = 2x = 28) and hexaploid (2n = 6x = 42). Common wheat (T. aestivum) is an allohexaploid of genomic formula AABBDD (Prohens-Tomas et al., 2009). The two main commercial type of wheat are durum (Triticum durum L., 2n = 4x = 28) and common wheat (Triticum aestivum L., 2n = 6x = 42). In hexaploid wheat the 21 chromosomes are divided into seven homologous groups identified with numbers from 1-7. Homoeologous chromosomes are similar both in structure and gen content.
The three chromosomes within the ABD homologous group usually share some loci in common for specific trait. Tetraploid and hexaploid wheat reproduce naturally as diploid. The gene ph1 (pairing homoeologous) which is present on the long arm of 5 B chromosome makes the reproductive mechanism possible which enables diploid pairing to occur (Acquaah, 2007; Prohens-Tomas et al., 2009). The common wheat Triticum aestivum L. is the more widely grown wheat.
The difference between individuals due to their genetic makeup or the influence of environmental factor in which they are grown is known as variation. If two individuals grow in the exactly similar environment and show differences in their characters such variation is genetic variation. The information, the type and the magnitude of genetic variability has great importance for the formulation of a plant breeding program (Khodadadi et al., 2011).
Genetic variability is the primary interest of the plant breeder since the highest genetic distance between parents will result in higher heterosis in the progeny (Khodadadi et al., 2011). Phenotypic variation is the result of both the genetic and the environmental factors. Genetic distance estimation could be one of the essential tools for selection of parents in wheat hybridization program.
Organic agriculture is not only focussed the product of a certain cultivar (e.g., traits related to productivity) but it is also a production controlled system (Van Bueren et al., 2002). Organic farming systems shares priority on traits like yield and end-use quality with the conventional breeding but in addition aim at traits such as good competition with weeds, resistance to seed-borne diseases and nitrogen use efficiency (Acquaah, 2007; Van Bueren et al., 2011; Van Bueren and Myers, 2012; Loschenberger et al., 2008). Most organic farms depend on mechanical weed control. Mechanical harrowing machines may cause damage to the plant during harrowing; the ability of genotypes to tolerate the mechanical damage or to recover rapidly from the damage are good traits which are important for organic farming systems (Donner and Osman, 2006).
According to Van Bueren et al. (2002), the management of the organic farming is not only aiming at high yield under low-input condition but also at yield stability. The buffering capacity of the population is higher than the pure lines in the fluctuation of the growing environment, due to the fact that populations have a diverse genetic pool to buffer adverse conditions compared to the pure line cultivars. To evaluate the performance of the genotypes, selection in a breeding program needs many years and location. This makes selection for yield stability difficult (Tester and Langridge, 2010). Infection of cultivars by fungal diseases is one of the causes for the reduction of the yield in the areas of humid and temperate climate areas like the Netherlands (Van Bueren et al., 2002).
The advantages of genetic diversity on crops are complementation, cooperation, compensation and capacity (Doring et al., 2011). When crops grow together with diverse genotype have different resource need they will complement each other in the uptake of limiting resource (Busch et al., 1974). For example, if crops with different rooting patterns grow together or crops with different light interception strategy they will not compete each other for the same resource. Some genotypes produce volatiles which have a repellent or toxic effect for insect pests and if those genotypes grow together with other genotypes they can help to protect those genotypes from pest attack. These type of effects termed as cooperation. In many cases, when one grows a pure line cultivar he can harvest a good yield only in good growing environment. If the growing environment varies the harvest will decline, whereas if the cultivar is mixed with genotypes which can withstand different environmental conditions it is possible to compensate each other. The average yield over different growing seasons will be higher with mixed genotypes than with pure lines under high variable environmental condition due to compensation (Doring et al., 2011). The population including various genotypes have the capacity to have more characters than the pure line.
Whereas genetic diversity also has some limitations, like competition of genotypes in populations for the same resources (e.g., plants compete for light and water). In diverse population plants may cost much of the energy for competition rather than grain yield, for instance if plants compete for light and plants that have good competitive ability for light but are poor in grain yield grow with those plants which have poor competitive ability with good grain yield, plant which have good competition for light will dominate and finally the productivity of the crop will be low (Doring et al., 2011). In genetically diverse population maximized grain yield is not stable through evolution the population can be dominated by high competitiveness but low grain yield ability genotypes (Zhang et al., 1999).
Murphy et al. (2005) argued that the organic growers and low-input farmers have diverse mechanisms for maintaining the soil fertility and pest management. This diversity is a challenge for the breeder to develop a cultivar for diverse agro-climatic zones. It is not economically feasible having test plots of different practice in the breeding station. Whereas the evolutionary breeding is a cost effective method to select specific traits in large number of plants having various populations growing on different farms.
To increase sustainability and competitive ability of the cultivar by increasing the genetic diversity, the Organic Research Centre, Elm Farm in the United Kingdom developed Composite Cross Populations (CCP) of wheat from 20 parents. To evaluate the phenotypic diversity and the potentials of those composite populations’ field experiments were conducted in the organic trail field of Wageningen University and Research Centre.
MATERIALS AND METHODS
Experimental set-up: The field experiment was conducted at the organic trial farm Droevendaal of Wageningen University and Research Centre the Netherlands during 2011/12 winter cropping session. The experimental design was a randomized complete block design and had three replications. Number of plots was differ each year as the number of genotypes differed each year. The plot sizes of 6x7.5 m (45 m2) were used. Each plot consisted of four beds of 1.5x7.5 m with a small path between the beds. The soil of the experimental plot was sandy.
Data collection: Plant height was measured on 40 randomly selected plants from ground level to the top of the spike excluding the awn. Length and width of the leaf next to the first node, next to the second node and flag leaf was determined from 40 randomly selected plants per plot. Measurements for the leaf were taken in the field and the results were taken to calculate the leaf area as indicated follows (Bilgi, 2006):
Leaf area = LxWxF
Where:
| L | = | Maximum length (cm) |
| W | = | Maximum width (cm) |
| F | = | Factor (0.707 for wheat (Bilgi, 2006) |
Number of spikelets per spike, Number of fertile and unfertile spikelets was determined from mean 40 randomly selected plants per plot (The number of spikelets with grains was counted as fertile spikelets and the empty spikelets counted as unfertile spikelets). Spike length was measured from the base of the spike to the top of the last spikelet, excluding the awns from mean of 40 randomly selected plants per plot. Distance between the flag leaf and the spike data’s were collected from the mean of 40 randomly selected plants per plot. The distance between spikelets data were recorded by exploiting the data of spike length and number of spikelet per spike:
![]()
Statistical analysis: The statistical software GenStat 15th edition was used for the statistical analyses. For the analysis of variance a general treatment structure in randomized blocks was carried using a threshold p<0.05 to declare differences significant. To check the assumptions of ANOVA (normality and homogeneity of variance) were not violated the residual plots were run. When the difference was significant (p<0.05) the fishers protected least significant (LSD) test (α = 0.05) was used to study which means differed significantly.
Analysis of the genotypic variance: The genotype and phenotypic variance components and coefficient of phenotypic and genotypic variability were estimated as:
![]()
Where:
| MSg | = | Mean square due to genotype |
| MSe | = | Environmental variation (error mean square) |
| r | = | No. of replication |
Environmental variance (σ2e) = Error mean square
Phenotypic variance (σ2p) = σ2g + σ2e
Heritability in the broad sense is the ratio of the total genetic variance to the phenotypic variance (Dudley and Moll, 1969). This is calculated as:
![]()
Where:
| H2 | = | Broad sense heritability |
| σ2e | = | Environmental variance |
| σ2p | = | Phenotypic variance |
| σ2g | = | Genotypic variance |
To study the diversity in performance of the genotypes the phenotypic traits were converted to different discrete classes: Plant height to five classes but other traits were converted to three classes as low, intermediate and high. The proportions of each class for each genotype were calculated.
The phenotypic frequency data of the traits was analyzed by the Shannon-Weaver diversity index, H'. This diversity index is widely used in studies H' of germplasm collections (Bechere et al., 1996). This is calculated as:
![]()
where n is the number of phenotypic classes for the trait and pi is the proportion of the total number of entries in the i-th class. The H' was calculated for each genotype and phenotypic traits. To keep the H' value between the range of 0-1 each value of H' were divided by log (n). For the analysis variance of diversity of each trait the normalized value of H' was used.
RESULTS AND DISCUSSION
There was no significant difference (p = 0.37) among the genotypes for the grain yield, PAR at the flowering stage and area of the second leaf next to the second node (Table 1). Whereas very highly significant differences were observed among the genotypes for the thousand kernel weight, number of spikelets per spike, number of unfertile spikelets, number of productive tillers per square meter, plant height, area of the 1st leaf, length between the flag and spike and spike compactness (Table 1). Area of the flag leaf and spike length also exhibited significant and highly significant difference, respectively.
There were much more observed differences between the genotypes at the vegetative stage and not for the grain yield.
Estimation of diversity index
Shannon-Weaver diversity index: The relative values of the traits for Shannon-Weaver diversity index, H' were different. The diversity index was small for all the traits (Table 2). Significant differences were observed for the traits plant height, flag leaf spike length and width of the leaf next to the first leaf.
| Table 1: | Analysis of variance of traits measured in the winter wheat trial 2011/12, Droevendaal, Wageningen, Netherlands |
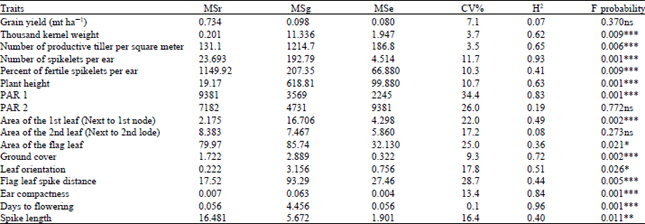 | |
| MSr: Mean square due to replication, MSg: Mean square due to genotype, MSe: Mean square due to error, CV%: Coefficient of variation, ***Very highly significant difference, **Highly significant difference, *Significant difference, ns: No significant difference | |
| Table 2: | Mean diversity index of each phenotypic characters of genotype (Shannon-Weaver diversity index, H') applied to the results of the trial 2011/2012, Droevendaal, Wageningen , Netherlands |
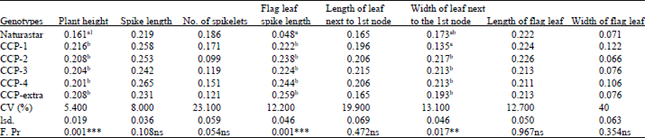 | |
| 1Genotypes with the same letter did not show significant difference (p = 0.05), ***Very highly significant difference, **Highly significant difference, ns: No significant difference was observed between genotypes | |
For plant height and flag leaf spike length, all CCPs have higher values for the diversity index than the pure line cultivar Naturastar (Table 2).
For traits of which no significant differences were found between genotypes: The pure line cultivar often showed the lowest Shannon-Weaver diversity index (H') such as for spike length and length of the leaf next to the first node (0.165). These results are plausible, as we expect lower diversity index from the pure line cultivar. However, the pure line cultivar had a high H' value for the number of spikelets per spike (0.186) which could be due to the fact that this cultivar had high tillering and the younger tillers were smaller than the earlier formed ones. There was no significant difference in number of spikelets per spike, however.
There were no significant differences between the CCPs in values for the Shannon-Weaver diversity index (H'), for the traits shown in Table 2 except for width of the leaf next to the first node. For the trait width of the leaf next to the first node this population showed a low H' index value (0.135) which is surprising as we expect a high diversity index value for the newly introduced population due to the fact that this population comes from a different environment and is likely less adapted.
The lowest diversity index was observed in width of the flag leaf with the highest value of CCP-1 (0.122) and the lowest of CCP-2 (0.066); however, no significant different was observed between the genotypes tested. This traits may be less appropriate to assess the diversity of the genotypes, since the values for the Shannon-Weaver diversity index (H') were very different.
Standard deviation within plots: The tested genotypes showed difference in their Standard Deviation (SD) within the plots. No significant difference was observed for the SD of the grain yield ton ha-1. The highest SD was observed in the CCP-4 (0.73) and the lowest SD was in the CCP-1 (0.39). Low SD in grain yield suggests a good level of stability.
Genotypes showed highly significant differences for the SD within plots for the traits plant height (p<0.001), flag leaf spike distance (p<0.001), width of the 1st next to the 1st node (p<0.008) and width of the flag leaf (p<0.001). Other traits did not show significant difference for the SD. These results are similar to the outcomes of the Shannon-Weaver diversity index (Table 3), except for the trait width of the flag leaf. For these traits, the genotype with high SD is more diverse than with lower SD.
| Table 3: | Mean comparison of standard deviations within plots genotypes tested in the 2011/12 growing season at Droevendaal, Wageningen |
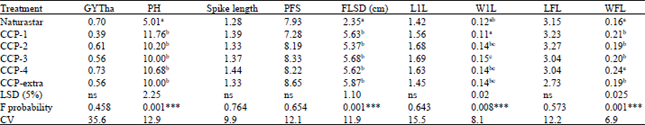 | |
| 1Genotypes having the same letter did not show significant difference (p = 0.05), GYTha = Grain yield ton per hectar FLSD: Flag leaf spike distance, PFS: Percent of fertile spikelet’s per ear, W1L: Width of the first leaf next to the 1st node, L1L: Length of the first leaf next to the 1st node, WFL: Width of the flag leaf, LFL: Length of the flag leaf. ***Very highly significant different between the genotypes, ns: No significant difference observed | |
For plant height, a significant difference was observed between the pure line cultivar and the CCP-populations and no significant difference was observed between the CCP-populations. Also for the flag leaf spike distances no significant difference was observed between the CCP-populations. The only difference was observed between the CCP-populations and the pure line cultivar Naturastar. For the width of the 1st leaf next to the 1st node the CCP-1 is significantly different from other CCP-populations and the Naturastar. The CCP-1 is not significantly different from the pure line cultivar. This result also had a similar pattern with the Shannon-weaver diversity index values.
However, for the width of the flag leaf the highest SD was observed in the CCP-4 which was significantly different from other CCP-populations and the pure line cultivar. The pure line cultivar had the lowest values and was significantly lower than all other genotypes. This result is different from the Shannon-Weaver diversity index values that were not significantly different. This difference in results suggests that it is good to use different methods.
The pure line cultivar generally had lower values for both the Shannon-Weaver diversity index and the SD within plots than the CCPs. This confirms our expectations since the CCPs have a higher level of diversity and we expect more variation from those populations. The differences in values between the CCPs, including the CCP-extra, are not significant for most of the traits measured and this shows that the CCPs maintain diversity within population for more than four generations.
CONCLUSION
The result of Shannon-Weaver diversity index and the standard deviation has similar output for the traits plant height, flag leaf spike distance, width of the 1st next to the 1st node and width of the flag leaf. There was no significant variation observed between the composite cross populations for the phenotypic diversity whereas, the variation was between the composite cross populations and the pure line cultivar. From this study we can see that the phenotypic diversity of the composite cross populations did not evolve through time.
ACKNOWLEDGMENT
I am highly thankful to my advisors Prof. Dr. Edith Lammerts van Bueren and Dr. Edwin Nuijten for the guidance, encouragement, constructive comment and supervisions. My special gratitude also goes to Nuffic Netherlands Fellowship Programme for offering me a scholarship opportunity to study at Wageningen University.
REFERENCES
- Bechere, E., G. Belay, D. Mitiku and A. Merker, 1996. Phenotypic diversity of tetraploid wheat landraces from northern and North-Central regions of Ethiopia. Hereditas, 124: 165-172.
CrossRefDirect Link - Busch, R.H., J.C. Janke and R.C. Frohberg, 1974. Evaluation of crosses among high and low yielding parents of spring wheat (Triticum aestivum L.) and bulk prediction of line performance. Crop Sci., 14: 47-50.
CrossRefDirect Link - Doring, T.F., S. Knapp, G. Kovacs, K. Murphy and M.S. Wolfe, 2011. Evolutionary plant breeding in cereals-into a new era. Sustainability, 3: 1944-1971.
CrossRefDirect Link - Dudley, J.W. and R.H. Moll, 1969. Interpretation and use of estimates of heritability and genetic variances in plant breeding. Crop Sci., 9: 257-262.
CrossRefDirect Link - Hildermann, I., A. Thommen, D. Dubois, T. Boller, A. Wiemken and P. Mader, 2009. Yield and baking quality of winter wheat cultivars in different farming systems of the DOK long‐term trial. J. Sci. Food Agric., 89: 2477-2491.
CrossRefDirect Link - Khodadadi, M., M.H. Fotokian and M. Miransari, 2011. Genetic diversity of wheat (Triticum aestivum L.) genotypes based on cluster and principal component analyses for breeding strategies. Aust. J. Crop Sci., 5: 17-24.
Direct Link - Van Bueren, E.T.L., S.S. Jones, L. Tamm, K.M. Murphy, J.R. Myers, C. Leifert and M.M. Messmer, 2011. The need to breed crop varieties suitable for organic farming, using wheat, tomato and broccoli as examples: A review. NJAS-Wageningen J. Life Sci., 58: 193-205.
CrossRefDirect Link - Van Bueren, E.T.L., P.C. Struik and E. Jacobsen, 2002. Ecological concepts in organic farming and their consequences for an organic crop ideotype. NJAS-Wageningen J. Life Sci., 50: 1-26.
CrossRefDirect Link - Loschenberger, F., A. Fleck, H. Grausgruber, H. Hetzendorfer and G. Hof et al., 2008. Breeding for organic agriculture: The example of winter wheat in Austria. Euphytica, 163: 469-480.
CrossRefDirect Link - Tester, M. and P. Langridge, 2010. Breeding technologies to increase crop production in a changing world. Science, 327: 818-822.
CrossRef - Zhang, D.Y., G.J. Sun and X.H. Jiang, 1999. Donald's ideotype and growth redundancy: A game theoretical analysis. Field Crops Res., 61: 179-187.
CrossRefDirect Link - Murphy, K., D. Lammer, S. Lyon, B. Carter and S.S. Jones, 2005. Breeding for organic and low-input farming systems: An evolutionary-participatory breeding method for inbred cereal grains. Renewable Agric. Food Syst., 20: 48-55.
CrossRefDirect Link








