ABSTRACT
The genetic heterogeneity of navel sweet orange (Citrus sinensis, Navel ) trees grown in Mazandaran province was evaluated with RAPD markers. Fifty two navel orange leaf samples from trees bearing each of the three morphologically distinct fruit types, soft rind, rough rind (old line), semi-rough rind (nucellar), were employed. Four of the 21 random primers used produced reproducible polymorphic bands. Of 864 fragments 150 to 2100 bp in size, amplified by the four primers, 70.13% were polymorphic. A similarity matrix using Nei coefficient, was generated and the genotypes were clustered with the UPGMA method by POPGEN32. The nucellar lines appeared to be closely similar to the standard Washington navel line. The highest levels of heterogeneity were observed within the soft rind and the rough rind lines. The observed heterogeneity may have resulted from the occasional budsports occurred and the selection methods variously employed by the nurserymen. The nucellar lines, which are gaining popularity only recently, have not been propagated extensively and thus remained comparatively homogeneous. It appears from the results of the present study, that the cultivar locally known as Thomson navel is misnamed and would probably be regarded as Washington navel.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajps.2007.1119.1124
URL: https://scialert.net/abstract/?doi=ajps.2007.1119.1124
INTRODUCTION
Citrus species are the most widely grown fruit crops in the world and have a large market worldwide. In Iran, Some of Citrus species have been cultivated traditionally since ancient times. In recent Decades their culture increased considerably and Iran is considered as the 8th largest Citrus producer in the world, with an annual production estimated in 3,037,000 tons, about 3% of total global production (FAO, 2006) and Mazandaran province produces more than 40% of total country production. navel orange cultivars, specially Thomson navel, are the most Important grown Citrus fruits in Mazandaran and because of ambiguous origin and uncertain date of parent rootstock introduction, their nomination is probably incorrect. The high variability observed in the Thomson navel orange orchards is not yet clear whether is genetically or influenced by environment conditions. Because of some difficulties that may occur in distinguishing real cultivar type and in order to certify the Citrus product for global markets, it was necessary to identify which cultivar navel sweet oranges grown in mazandaran province belong to.
The genus Citrus belongs to the subtribe Citrinae, tribe Citreae, subfamily Aurantioideae of the family Rutaceae. All Aurantioideae species are trees or shrubs with persistent leaves except for the three monotypic genera Poncirus, Aegle and Feronia, three species of Clausena and one of Murraya (Hodgson et al., 1967; Swingle, 1967; Tanaka, 1977). Citrus species have a high variation worldwide and hybridization, apomixes and many centuries of cultivation have complicated Citrus taxonomy with the result that very different numbers of species have been proposed. Although the genus Citrus has many species (Vardi and Spiegel-Roy, 1978; Herrero et al., 1996), the main cultivated species are sweet oranges (Citrus sinensis (L.) Osb), tangerines (C. clementina Hort. ex Tan. and C. unshiu (Mak.) Marc. mainly), grapefruits (C. paradisi Macf.) and lemons (C. limon L. Burm. f.).
The use of molecular markers has been a valuable and precise strategy to Identify Citrus species, cultivars and biotypes and to investigate the genetic diversity of Citrus species. Techniques like RFLP (Restriction Fragment Length Polymorphism), VNTR (Variable Number of Tandel Repeats), RAPD (Random Amplified Polymorphic DNA), Microsatellite or SSR (Simple Sequence Repeat), AFLP (Amplified Fragment Length Polymorphism), ISSR (Inter Simple Sequence Repeat) and other markers have been used in germplasm characterization, studies of genetic diversity, phylogenetic analysis and systematics (Coletta Filho et al., 1998; Roose and Fang, 1997; Fang et al., 1998; Federici et al., 1998; Nicolosi et al., 2000; Asins et al., 2001; Southwick et al., 2003; De Pasquale et al., 2006; Shahsavar et al., 2007). Among them, random amplified polymorphic DNA (RAPDs) has been employed most widely. RAPD have gained more attention due principally to the easiness of the procedure, being inexpensive and the very low amount of DNA required for analysis. In Citrus, RAPDs have been used for cultivar identification, genetic mapping, genetic diversity assessments and other breeding programs. Luro et al. (1992) described the use of RAPD marker for surveying the Citrus taxonomy and genetics. RAPDs have been used to generate linkage maps for Citrus (Cai et al., 1994). Deng et al. (1995), used RAPDs to Identify some in vivo and in vitro lemon mutants. Coletta Filho et al. (1998) implemented RAPD markers to study genetic diversity of some mandarins. Federici et al. (1998) used RAPDs to study the phylogenetic relationship among Citrus and related genera in combination with RFLP polymorphisms. Asins et al. (2001) reported the high efficiency of RAPDs for Clemantine mandarin genetic studies. Guo et al. (2002) used molecular markers to characterize some Citrus somatic hybrids.
The objective of this study was to assess genetic diversity of Citrus navel orange cultivars grown in Mazandaran Province through RAPD markers.
MATERIALS AND METHODS
Plant material: This research was conducted at Sari agriculture faculty, Mazandaran University, 2004-2005. Fifty-two genotypes belonging to Different navel Orange Cultivars (Thomson (Soft rind), Thomson (Rough rind), Thomson (Nucellar), Standard navel Cultivars (Navalate, Washington, Foyus) and some Unknown navel Genotypes) were sampled. They were collected from Ramsar Citrus Research center and different Orchards in Mazandaran Province (Table 1). Fresh leaves were collected and transferred to laboratory in an icebox and were stored at -20°C until use.
DNA extraction: Isolation of total genomic DNA for RAPD analysis was carried out through a new modified method based on Dellaporta et al. (1983). Leaf pieces were washed with tap water and rinsed twice in sterile distilled water and ground with liquid nitrogen in a sterile mortar and pestle. The powder obtained (about 200 mg) was transferred to a microcentrifuge 1.5 mL tube and resuspended in 600 mL extraction buffer (120 mM Tris (pH 8.2), 800 mM NaCl, 50 mM EDTA, 1% SDS, 0.2% β-Mercaptoethanol). Samples were incubated for 45 min at 65°C water bath and then 300 mL of 5 M KCl was added. The tubes were carefully inverted several times, left for 10 min at room temperature and centrifuged at 10000 g for 10 min. The supernatant was transferred to a new microcentrifuge tube, one volume of Chloroform/ isoamylalcohol (24:1) was added and after several inverting they were centrifuged at 12000 g for 10 min, supernatant was transferred to a new microcentrifuge tube and was precipitated with an equal volume of 2-isopropanol for 15 min at 4°C and centrifuged again (12000 g for 8 min). The crude DNA pellet was washed with ice-cold 80% ethanol, dried for 20 min at room temperature and resuspended in 70 μL of TE buffer. DNA was quantified by agarose electrophoresis gels after ethidium bromide staining by comparison with known quantities of lambda phage DNA used as a control. After dilution with sterile distilled water to a final concentration of 20 ng μL-1 the DNA solution was used directly for amplification.
| Table 1: | Fifty two Navel orange genotypes used in this study and their group |
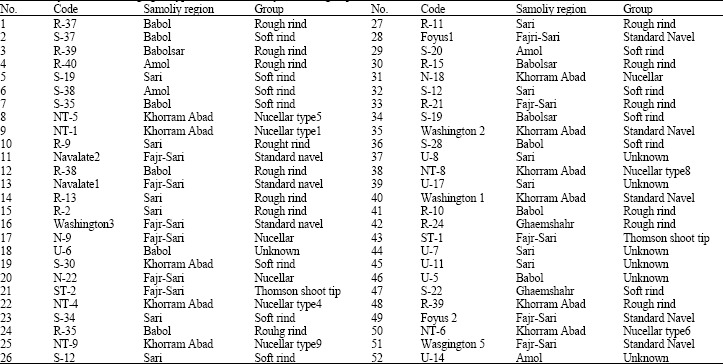 | |
Amplification protocol: A total of 21 decamer random primers previously selected as being suitable to generate complex and reproducible RAPD patterns in several Citrus biotypes, were used in this work. Amplifications with decamer primers were carried out in 25 μL final volume of a reaction mixture containing: 2.5 μL 10xTaq DNA polymerase buffer (100 mM Tris-HCl (pH 9.0); 500 mM KCl, 15 mM MgCl2) (Cinnagen Co., Iran), 200 mM each of dATP, dGTP, dCTP, Dttp (Fermentas Co. Lithuania); 0.5 μM of each of the primers (Operon Technologies Inc., USA), 20 ng genomic DNA and 1 Unit Taq DNA polymerase (Cinnagen Co., Iran). Amplification reactions were performed in a DNA thermal cycler (Eppendorf, mastercycler gradient) programmed for the following conditions: initial denaturation for 4 min at 94°C, 38 cycles of denaturation (60 s at 94°C), annealing (60 sec at 35°C) and extension (2 min at 72°C) followed by a final extension at 72°C for 10 min. Amplification products were analysed by submarine horizontal electrophoresis in 1.5% agarose gels, stained in ethidium bromide solution and visualised by transillumination with UV light after ethidium bromide staining. Gels were recorded using Vilber-Lourmat Gel Documentation unit (Vilber-Lourmat Co., France).
RESULTS AND DISCUSSION
In preliminary experiments the 21 selected primers generated reproducible, quite polymorphic bands, of which 4 primers were used for data analysis. These 4 primers could favourably differentiate between the 5 populations under study (Table 2).
A total of 29 distinct bands had been generated for 52 samples (Fig. 1-3) and a mean band number of 7.25 were scored for each polymorphic primer. All of these 29 bands were polymorphic in 5 populations. Size of the bands ranged from 150 to 2100 base pairs. Study of the banding pattern showed that primers OPA-06 and OPQ-11 generated the most polymorphic bands, while primer OPH-15 had the least polymorphic bands.
In general, a total of 864 RAPD bands were produced in 52 navel orange genotypes using the 4 primers.
Data processing: Each gel analysed by scoring the presence (1) or absence (0) of polymorphic bands in individual lanes. The POPGENE 32 statistical package (Yeh et al., 1997) was used to estimate genetic similarities through Nei`s similarity coefficient (Nei, 1978). The matrix of generated similarities was analysed by the Unweighted Pair Group Method with Arithmetic average (UPGMA) and a cluster was produced. For data analysis by POPGENE32 software, data of each of 5 populations (soft rind, rough rind, nucellars, standard navels and unknown group) were inserted separately, with the assumption that they are certain groups.
Gene frequency for each locus was also estimated. The highest and lowest gene frequency of 0.5953 and 0.2418 was observed in OPA-06-04 and OPQ-18-07 loci, respectively. The most polymorphic locus was in soft rind and rough rind groups (100% polymorph) while the lowest polymorphism was obtained for nucellar group with apolymorphism percentage of 48.28 (Table 3). So, it can be assumed the nucellar population had lower genetic variability than other populations and thus may be more homogen.
| Table 2: | The 4 primers that were used for analysis and level of polymorphism they revealed |
 | |
| Table 3: | Mean genetic variability and level of polymorphism in 52 navel orange genotypes grouped in 5 populations |
 | |
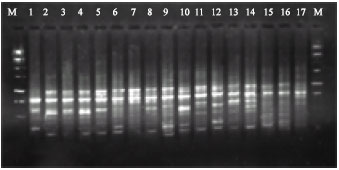 | |
| Fig. 1: | RAPD amplified with the arbitrary primer OPA-06 using DNAs of different Navel orange genotypes, M: weight marker |
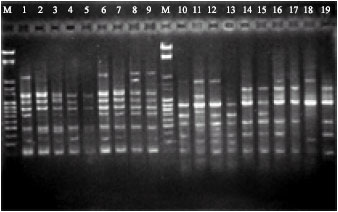 | |
| Fig. 2: | RAPD amplified with the arbitrary primer OPB-18 using DNAs of different Navel orange genotypes, M: weight marker |
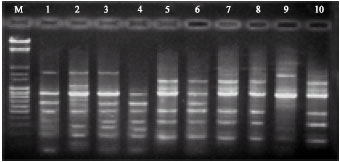 | |
| Fig. 3: | RAPD amplified with the arbitrary primer OPQ-11using DNAs of different Navel orange genotypes, M: weight marker |
Nucellar population had the lowest genetic variability with a mean of 0.2045 while that of rough rind group was 0.4044 showing the highest genetic variation indicating its wide genetic structure. According to these data we can assume nucellar population as the lowest homogenous population with a narrow genetic base and rough rind population as the most heterogenous one.
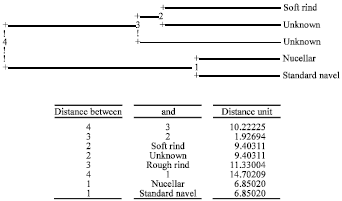 | |
| Fig. 4: | UPGMA dendrogram of 5 Navel orange populations (52 samples) based on Nei`s similarity coefficient |
| Table 4: | Genetic distance (genetic similarity) between 5 distinct populations of Navel oranges genotypes under study |
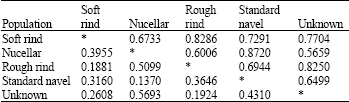 | |
With use of Nei`s similarity coefficient, nucellars and unknown populations had the highest genetic distance (0.5693) while nucellar and standard navel populations had the lowest (0.1370) (Table 4).
Cluster analysis of 5 populations (Fig. 4) grouped nucellars and standard navels in a separate cluster, while soft rind group and Unknown group constituted another cluster and the Rough rind group was located in the same cluster with a low distance.
As it can be observed in cluster, the groups nucellars and standard navels showed the highest genetic similarity (0.8720) indicating their close genetic relationship. In contrast, the highest genetic distance was obtained between nucellars and unknown group (0.5693) indicating their considerable genetic differences.
According to genetic analysis of 5 populations, nucellars and standard navel populations had a low genetic diversity and were more homogen that is in accordance with the fact that nucellars were propagated through parent type nucellar seeds and standard navels were propagated through grafting from mother rootstocks that inhibits heterogeneity. Rough rind, soft rind and unknown groups were located in a single cluster with a low distance from each other. The high variation observed between rough rind and soft rind individuals may be due to frequent occurrence of bud sport and subsequent selections leading to diversification of these populations. Motohashi et al. (1992) had previously reported a high mutation rate and varability among Citrus species. They reported that the role of hybridization in genetic diversification is low while the frequent mutations and subsequent selection are the major factors of diversification. This study showed that differences observed between populations of Thomson navel subgroups and other navel cultivars, have been also produced through mutations and subsequent propagation from mutant part of mother rootstock. This process will result in different progenies from the mother plant through time. Result of several studies performed by Motohashi et al. (1992), Sugawara et al. (1995) and Asins et al. (2001) supports this finding.
In general, RAPD markers classified into 5 populations, but they were not able to differentiate between individuals in each population. This is in accordance with Coletta Filho et al. (1998) that attributed the difficulties of classification to the low level of polymorphism. In this study, we estimated low levels of polymorphism for different navel orange cultivars. It may be due to some reasons discussed below. There is naturally very low variation among navel oranges. Roose et al. (1998) in a experiment studying several Citrus genotypes, reported a low efficiency for RFLP, RAPD and ISSR markers. ISSR markers could not differentiate several Washington navel genotypes that were originated from different regions (Roose and Fang, 1997). Southwick et al. (2003) after evaluating several microsatellite loci in Citrus species reported a single SSR primer that was able to produce distinct bands distinguishing navel groups from Valencia cultivars. According to Asins et al. (2001), genetic diversity of some Citrus populations like clementine and most of oranges is so low. These cultivars were not developed through Hybridization but through selection of favorable mutations, so they show a limited genetic variation. By use of a higher number of RAPD random primers it may be possible to distinguish between different genotypes.
In this study, the resultant dendrogram reveals the significant level of genetic similarity between standard navels and the genotypes presumed as Thomson navel. There was a high variation among the genotypes known as Thomson naevl and these genotypes experienced a progressive and steady diversification and were gradually transformed to different off type forms. The nucellars are more homogen with a low variation due to their few cycles of propagation, so they show a high similarity to parent Washington Navel rootstock they were derived of as shown in the cluster, they are classified in the same cluster. It appears that different genotypes belonging to cultivar known as Thomson navel are so close to parent Washington navel or may be the same as Washington navel. Morphological characteristics are also supporting RAPD results. Because of morphological relationships between cultivar known as Thomson navel and standard Washington navel, it can be assumed that with a high probability Nucellar lines and different Thomson navel genotypes are the same. In other words, the parent rootstock that Nucellar line was derived from belongs to Washington navel group.
REFERENCES
- Breto, M.P., C. Ruiz, J.A. Pina and M.J. Asins, 2001. The diversification of Citrus clemantina Hort. ex Tan, a vegetatively propagated crop species. Mol. Phylogenet. Evol., 21: 285-293.
CrossRef - Colettafilho, H.D., M.A. Machado, M.L.P.N. Targon, M.C.P.Q.D.G. Moreira and J. Pompeu, 1998. Analysis of the genetic diversity among mandarians (Citrus spp.) using RAPD markers. Euphytica, 102: 133-139.
CrossRef - Dellaporta, S.L., J. Wood and J.B. Hicks, 1983. A plant DNA minipreparation: Version II. Plant Mol. Biol. Rep., 1: 19-21.
CrossRefDirect Link - De Pasquale, F., M. Siragusa, L. Abbate, N. Tusa, C. De Pasquale and G. Alonzo, 2006. Characterization of five sour orange clones through molecularmarkers and leaf essential oils analysis. Scientia Hortic., 109: 54-59.
CrossRef - Federici, C.T., D.Q. Fang, R.W. Scora and M.L. Roose, 1998. Phylogenetic relationships within the genus Citrus (Rutaceae) and related genera as revealed by RFLP and RAPD analysis. Theor. Applied Genet., 94: 812-822.
CrossRef - Guo, W.W., Y.J. Cheng and X.X. Deng, 2002. Regeneration and molecular characterization of intergeneric somatic hybrids between Citrus reticulata and Poncirus trifoliata. Plant Cell Rep., 20: 829-834.
Direct Link - Herrero, R., M.J. Asins, E.A. Carbonell and L. Navarro, 1996. Genetic diversity in the orange subfamily Auarantioideae L. intraspecies and intragenous genetic variability. Theor. Applied Genet., 92: 599-609.
Direct Link - Motohashi, R., T. Matsuyama and T. Akihama, 1992. DNA fingerprinting in citrus cultivars. Proc. Int. Soc. Citriculture, 1: 221-224.
Direct Link - Nei, M., 1978. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: 583-590.
PubMedDirect Link - Nicolosi, E., Z.N. Deng, A., Gentile, S. La Malfa, G. Continella and E. Tribulato, 2000. Citrus phylogeny and genetic origin of important species as investigated by molecular markers. Theor. Applied Genet., 100: 1155-1166.
Direct Link - Roose, M.L. and D.Q. Fang, 1997. Identification of closely related citrus cultivars with inter-simple sequence repeat markers. Theor. Applied Genet., 95: 408-417.
Direct Link - Shahsavar, A.R., K. Izadpanah, E. Tafazoli and B.E. Sayed Tabatabaei, 2007. Characterization of citrus germplasm including unknown variants by Inter-Simple Sequence Repeat (ISSR) markers. Scientia Hortic., 112: 310-314.
CrossRef - Norheim, G., A. Aase, D.A. Caugant, E.A. Hoiby and E. Eritzsonn et al., 2005. Development and characterization of outer membrane vesicle vaccines against serogroup A Neisseria meningitids. Vaccine, 23: 3762-3774.
CrossRef - Welsh, J. and M. McClelland, 1990. Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res., 18: 7213-7218.
CrossRefPubMedDirect Link








