Research Article
A Phenylalanine Ammonia-lyase Gene from Asparagus: cDNA Cloning, Sequence and Expression in Response to Wounding
Not Available
Toshiyuki Matsui
Not Available
Haruo Suzuki
Not Available
Yusuke Kosugi
Not Available
The toughening of asparagus is due to lignification of fibrovascular tissues and occurs within hours after harvest in spears stored at ambient conditions (Isherwood, 1963). Lignin, the substance that lends fibers their toughness, is polymerized from cinnamyl alcohols derived from shikimic acid pathway (Gross, 1981). PAL catalyzing the conversion of L-phenylalanine to trans-cinnamic acid in this pathway has been considered to play an important role as a key enzyme. Fluctuation in PAL activity has been shown to be a key element controlling the synthesis of lignin and in many cases, an increase in the amount of PAL mRNA has been shown to underlie the increase of PAL activity (Edwards et al., 1985; Fritzemeier et al., 1987; Lawton and Lamb, 1987; Orr et al., 1993). In elicited cultured pine cells for instance, the induction of PAL activity occurs concurrently with increases in the activities of lignin-specific enzymes and is followed by the deposition of apparently genuine gymnosperm lignin in the cell walls (Campbell and Ellis, 1992a, 1992b). PAL transcripts increased in response to fruit ripening and wounding in the mesocarp tissue of Cucurbita maxima (Kato et al., 2000). In freshly harvested asparagus spears PAL activity increased upon excision and incubation (Goldstein et al., 1972). Increased PAL activity was also observed in intact spears given an extended period of cold treatment. However, PAL activity in asparagus spears have not yet been studied in relation to fiber development and no PAL gene expression is detected during storage of asparagus spears.
This study reports the cloning and characterization of a partial PAL cDNA from harvested asparagus spears. By RNA blot analysis and in situ RNA hybridization the pattern of expression of PAL gene has also been reported in relation to fiber development and PAL enzyme activity during storage.
Materials and Methods
Plant materials: Green asparagus spears (Asparagus officinalis L. cv. Welcome) harvested from a commercial crop in Kagawa, Japan in the year 2002 were obtained directly from farmer’s field. Spears were hand harvested and trimmed to approximately 25 cm length. The spears, which were of good quality, straight with closed bracts, were put in polyethylene bags and held at 200C for up to 48 hours. Fiber content in both top and bottom portions of the spears was measured at harvest (0 h) and after 8, 16, 24, 36 and 48 h. For PAL enzyme analysis and RNA extraction the spears were frozen at -800C.
Texture measurement: Texture was measured rheologically based on the measurement of resistant to pressure or shearing. Breaking force to indicate the fiber content in spears was determined with a creep meter YAMADEN RHEONER RE-3305 equipped with software Ver. 2.0 for automatic analysis. The thickness of the blade was 0.04 mm and it sheared at the rate of 1 mm per second with a pressure of 20 kgf. Spears were cut into two equal pieces and breaking force readings were made separately in the mid point of top and bottom portions of the spears.
Extraction and Assay of PAL activity: Two g of spear tissues were homogenized at 20C with a mortar and pestle in 10 ml of 0.1 M borate buffer, pH 8.8 and 1.0 g PVPP. A further 10 ml of the buffer was added to the homogenate which was then centrifuged at 14, 000 rpm for 20 min and the supernatant was used for enzymatic assays. Protein concentration was measured by Lowry (Lowry et al., 1951) method. PAL activity was determined spectrophotometrically by measuring the increase in A290 due to the formation of trans-cinnamic acid. The reaction mixture consisted of 50 mM borate buffer, pH 8.8, 20 mM L-phenylalanine and the enzyme preparation in a total volume of 3 ml. A sample without L-phenylalanine was used as a blank. The activity was expressed as nmol trans-cinnamic acid formed per h per mg of protein.
RNA extraction and amplification of poly (A)+ RNA by RT-PCR: Total RNA was extracted according to the Hot Borate method of Wan and Wilkins (1994). The first strand cDNA was synthesized from 2 g of the total RNA by reverse transcriptase with Oligo-(dT) primer according to the instruction of SUPER SCRIPTTM Preamplification System for First Strand cDNA Synthesis (GIBCOBRL, Tokyo, Japan). PCR was performed in a total volume of 25μl containing the first strand cDNA reaction products, 10xPCR Buffer, MgCl2, dNTP, First Start Taq DNA Polymerase (Roche) and primers. The primers (5’-ATYGAGGCTGCTGCYATTATG-3’ as the upstream primer and 5’-ACATCTTGGTTGTGYTGCTC-3’ as the downstream primer) were designed and synthesized on the basis of amino acid domains (IEAAAIM and AEQHNQD respectively) conserved in various PAL genes. The Sal 1 and Not1 restriction site sequences were also included at 5’-end of the sense and antisense primer, to facilitate cloning of PCR product. The PCR procedure started with 10 min at 950C and was carried out 35 cycles of 30 s at 950C, 30 s at 500C and 30 s at 720C, and 10 min at 720C with ASTEC Program Temperature Control System PC-700. The PCR products were confirmed by agarose gel electrophoresis.
Cloning and sequencing of cDNA: The amplified cDNA was ligated to the plasmid pSPORT1 and cloned into Escherichia coli (DH-5α ) Not 1- Sal 1 – cut (BRL, Tokyo, Japan). Sequencing was performed by the cycle sequencing method using GATCR-Bio Cycle sequencing Kit and a DNA sequencer GATC 1500 Long-Run system (GATC Gmbh, Konstanz, Germany).
Sequence data analysis: Sequence analysis was performed using computer software GENETYX-MAC Ver.7. Homology searches with the Genbank and the EMBL databases were performed using the homology program in the software. The phylogenetic tree was also constructed with the UPGMA method in the software.
Preparation of the digoxigenin (DIG)-UTP-labeled RNA probe: The cloned RT-PCR product including the encoded region of PAL gene was cleaved by Not 1 and Sal 1 from the pSPORT1 vector that had been amplified in Escherichia coli (DH-5α) and it was purified and recovered by gel electrophoresis. Antisense DIG-labeled RNA probe was prepared according to the instructions of DIG RNA Labeling Kit (Boehringer Mannheim) using SP6 RNA polymerase.
Northern blot analysis: Ten μg of total RNA was subjected to electrophoresis on a 1.0 % agarose (Type II) gel containing 20 x MOPS and 37 % formaldehyde. After electrophoresis for 30 min, RNA was visualized with ethidium bromide under UV light to confirm equal loading of RNA in each lane. RNA was transferred to a positively charged nylon membrane HybondTM- N+ (Amersham Pharmacia Biotech) by capillary action with 20 x SSC and then after drying the membrane RNA was fixed under UV light. The membrane was prehybridized at 500C with 5 x SSPE, 5 x Denhart’s solution, formamide and 10 % SDS for 3 h. Hybridization was performed at 500C using the gene specific antisense DIG-labeled RNA probe for 24 hours using the same prehybridization buffer. After hybridyzation, the membrane was washed twice with 2 x SSPE containing 0.1 % SDS for 10 min at room temperature, once with 1 x SSPE containing 0.05 % SDS for 15 min at 650C and once with 0.2 x SSPE for 10 min containing 0.05 % SDS at 650C. The membrane was further washed with Buffer A containing maleic acid and Tween 20 at room temperature and was blocked with 2 % blocking reagent in maleic acid buffer for 30 minutes. Subsequently, the membrane was incubated with Anti-Digoxygenin –AP, Fab fragments (Boehringer Mannheim) in the blocking buffer for for 30 min. Signals were detected by color reaction using 5-bromo-4-chloro-3-indolyl- phosphate and 4-nitro blue tetrazolium chloride as the substrate.
Results
Isolation and identification of cDNA clone: The cDNA pAS-PAL is a partial clone encoding a harvest-induced transcript from asparagus spears. The encoded mRNA is 527 bp long (Fig. 1) and is highly homologus to the gene of other plant (Fig. 2). The pAS-PAL sequence is 75.3 % identical to PAL from rice (X87946), 75.3 % identical to PAL from maize (L77912), and 77.0 % identical to the barley PAL (Z49145) all of which are wound-induced gene (Table 1).
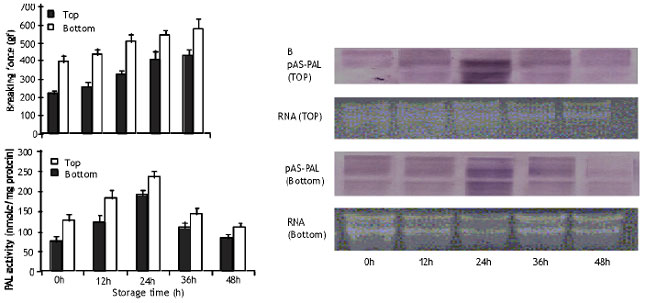 | |
| Fig. 1: | A changes in fiber content and PAL activity in the top and bottom portions asparagus Each point represent the mean of three replications. Verticle bars indicate SEs. B, Northere blot analysis of total RNA 10 (μg/lane) showing the expression of pAS-PAl. Equal loading of RNA was confirmed by staining a gel with ethidium bromide. |
 | |
| Fig. 2: | Necleotide sequence and deduced amino acid sequence of the cDNA clone corresponding ot pSA-PAL. The predicted amino acid sequence is given in single-letter code for each amino acid. The arrows indicate the positions of degenerated primers ( sense, antisense ) used for RT-PCR. Numbering reffers to total nucleotide residues on each line. |
 | |
| Fig. 3: | Comparison of the deduced amino acid sequences from asparagus (AB102677), barley (Z49145), maize (L77912), pea (10001), rice (X87946), tobacco (M84466) and tomato (M90692) by multi alignment. The amino acid residues are numbered at the beginning and end of the sequences on each line. Shaded letters denote the amino acid residues those are identical. Dashes in the amino acid sequences represent gaps introduced to maximize alignment of the polypeptides |
| Table 1: | Percentage of nucleotide and deduced amino acid homology between PAL from asparagus spear and other plants in the databases |
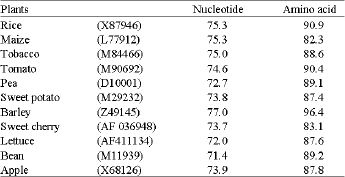 | |
| Asparagus PAL (AB102677) is calculated as 100 % | |
Allowing for conservative amino acid substitutions, the similarities are 90.9, 82.3 and 96.4 % for the rice, maize and barley sequences, respectively.
Induction of PAL activity: Fig. 3 shows the development of PAL activity in both top and bottom portions of asparagus spears. The increase in activity continued until 24 hours resulting in 30 and 55% increase over the initial level in top and bottom portions, respectively. Thereafter, the level of activity started to decline.
Fiber development: A general increase in fiber content in both the top and bottom portions of the spears was observed throughout the experimental period. Although the fiber development in both top and bottom portions followed almost the same pattern, the breaking force indicating the toughness of the spears was greater in the bottom than in the top portion.
Expression of PAL gene: The expression of pAS-PAL was induced by wounding due to harvest. Maximum pAS-PAL expression appeared after 24 hours of storage and coincided with the peak of enzyme activity (Fig. 3) A phylogenetic tree (Fig. 4) was generated from the alignment of the deduced amino acid sequences of pAS-PAL and other PAL gene in the database. The pAS-PAL (accession No. AB102677), PALfrom barley (Z49145) and maize (L77912) strongly clustered together in a subgroup belonged to the monocotyledon, having closest relationship with rice PAL gene.
The development of PAL activity in asparagus spears was similar in a number of respects to other wound induced tissue systems (Rubery and Fosket, 1969; Kato et al., 2000). There was an increase in PAL activity in both top and bottom portions of the spears which continued for about 24 hours. After that PAL activity started to decline. Although PAL activity in both top and bottom portions followed almost the same pattern the activity was higher in bottom than top portions.
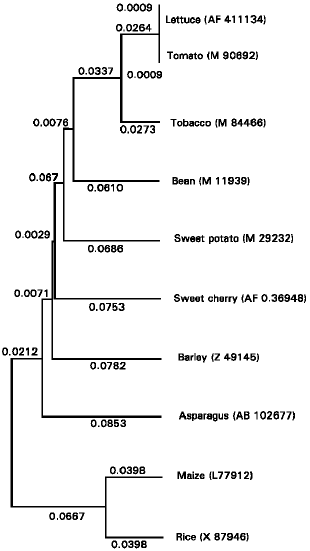 | |
| Fig. 4: | Phylogenetic tree of the alignment of pAS-PAL deduced amino acid sequence with other PAL gene in the database. Protein sequences were aligned using UPGMA, and a phylogenetic tree was constructed using GENETYX-MAC software. The GenBank accession numbers are shown in the parentheses |
High PAL activity associated with bottom portion, or tissue near to cut surfaces, such as we observed, has been noted previously (Goldstein et al., 1972; Engelsma, 1968). The basal tissue of the asparagus is highly lignified and PAL activity is known to be correlated with the degree of lignification in many tissues (Higuchi, 1966; Yoshida and Shimokoriyama, 1965). Alternatively, ethylene produced by the wounded tissue may induce PAL activity as observed in some other plant tissues (Imaseki et al., 1968; Kato et al., 2000).
To understand the molecular basis of induction of PAL activity in relation to fiber development of asparagus spears we have isolated a cDNA for PAL from harvested asparagus and found that it is wound induced. The clone, pAS-PAL had high sequence similarity to rice, maize and barley cDNA that were also associated with wounding. Phylogenetic analysis of PAL sequences has revealed the existence of at least two major branches that contain characteristic conserved amino acid sequences, monocotyledon and dicotyledon. The pAS-PAL belonged to the subgroup wound-induced monocotyledon as it was highly homologous to rice PAL and was closely related to PAL from Maize. Our results suggest that induction of PAL activity in harvested asparagus is regulated by transcription of pAS-PAL in response to the wounding associated with harvest.
Northern blot analysis revealed that the expression of pAS-PAL increased in harvested asparagus until 24 hours of storage period at 200C and afterwards it started to decline although the toughness determined as the value of breaking force was still increasing. The increase in fiber content after 24 hours might be due to the higher activity of other lignin-specific enzymes like peroxidase and cinnamyl alcohol dehydrogenase. Further research is necessary to investigate the role of other lignin-specific enzymes controlling the toughness of asparagus.
The financial support of the Ministry of Education, Science, Sports and Culture of Japan under scholarship program for foreign students is gratefully acknowledged.