ABSTRACT
This study aimed to isolate and identify a new local bacterial strain, able to completely degrade keratin-rich wastes into soluble and useful materials which can be used in many proposes. Bacterial keratinases are of particular interest because of their action on insoluble keratin substrates and generally on a broad range of protein substrates. These enzymes have been studied for de-hairing processes in the leather industry and hydrolysis of feather and keratin. Samples from poultry industry wastes, soil, water, fodder and feather were collected from different places in Jeddah, Saudi Arabia. Each sample was plated on feather meal agar plates containing 5 g L-1 feather as the sole carbon and nitrogen source and the obtained colonies were selected, purified and their growth were detected on skimmed milk agar and feather meal broth media. The well grown isolates on feather meal agar which producing the largest clearing zone on skimmed milk plate were selected for keratinase assays. Out of 23 bacterial isolates, 7 isolates were selected. The best keratinase producing bacterium kera MS21 was selected and identified based on morphological, physiological and some biochemical characteristics. It was recorded as a species belonging to the genus Pseudomonas and identified as Pseudomonas sp. The results of identification were confirmed by 16S rDNA studies. Precipitation and purification of the keratinase enzyme in addition to factors affecting enzyme activity were studied. The enzyme molecular weight was determined to be of 30 KDa using sodium dodecyl sulfate polyacrylamide gel electrophoresis analysis. The optimum temperature and pH were determined to be 37°C and pH 8.0, respectively. The effect of some proteases inhibitors and activators were also studied.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajbkr.2010.1.13
URL: https://scialert.net/abstract/?doi=ajbkr.2010.1.13
INTRODUCTION
Keratins represent the major constituents of skin and its appendages such as nail, hair, feather and wool ( Lin et al., 1996). Keratins are grouped into hard keratins (feather, hair, hoof and nail) and soft keratins (skin and callus) according to the sulphur content. Hard keratins like chicken feather which are recognized as a solid wastes generated from poultry processing industry are insoluble and resistant to degradation by common proteolytic enzymes, such as trypsin, pepsin and papain because of their high degree of cross-linking by disulfide bonds, hydrogen bonding and hydrophobic interactions (Annapurna et al., 1996). Keratinous wastes are increasingly accumulating in the environment generated from various industries. To recycle of such wastes, biotechnological alternatives are developing to hydrolyze those materials to soluble form using specific enzyme of microorganisms named keratinase which is an efficient proteolytic enzyme, able to hydrolyze insoluble keratin. Keratinases have enormous potential applications in processing waste in the poultry and leather industries. They are used in detergent production, medicine, cosmetics, leather and feed industries and in newer fields like prion degradation for treatment of dreaded mad cow disease. Keratinase produced by insects and mostly by some microorganisms including bacteria, saprophytic and parasitic fungi and actinomycetes. Bacterial keratinases are of particular interest because of their action on the degradation of the insoluble keratin substrates and generally on a broad range of protein substrates ( Lin et al., 1996). Many studies monitored the production of keratinase by Bacillus licheniformis PWD-1, Fervidobacterium, Chryseobacterium sp., Streptomyces, (Sangali and Brandelli, 2000; Langeveld et al., 2003; Riffel et al., 2003). Chryseobacterium sp., strain kr6 was isolated from waste of a poultry industry and was capable to completely degrade chicken feathers. Its extracellular keratinase is a metalloprotease with great potential for biotechnological applications ( Riffel et al., 2003). Another work describes the effect of temperature, initial pH and substrates on keratinase production by Chryseobacterium sp., kr6 during growth on native feathers (Brandelli and Riffel, 2005). More recently, Kansoh et al. (2009) isolated two bacterial strains producing keratinase from Egypt soil.
The aims of this study were to isolate and identify a new efficient feather-degrading bacterium, which uses feather as carbon and nitrogen sources, from poultry decomposed wastes, also aimed to extraction, isolation, purification and characterization of keratinase enzyme from this new isolate.
MATERIALS AND METHODS
Sample Collection and Bacterial Isolation
Thirty samples were collected from different sources and places including natural composting wastes from chicken farms, soil, feather and chicken fodder. Samples (about 100 g) were collected from Jeddah, Saudi Arabia in sterile plastic bags and transferred directly to the laboratory and spreaded on paper sheet until air dried. Serial dilutions from each sample were prepared in sterile buffer composed of K2HPO4, 2 g L-1; KH2PO4, 1 g L-1; NaCl, 0.5 g L-1. About 0.1 mL of the suitable dilution was placed on minimal medium supplemented with feather as the sole carbon and nitrogen source (feather meal plate).
Water samples were collected from Red sea marine water, Jeddah, Saudi Arabia, serial dilutions were performed and 1 mL of the suitable dilution was used to inoculate feather meal plate. On some cases, the sea water was plated directly on the agar plates without dilutions.
Preparation of Native Chicken Feather
Native chicken feathers were cut with scissors to small pieces of 1-3 cm long, then, washed several times with tap water. Defatted of feather pieces was done by soaking them in a mixture of chloroform: methanol (1:1 v/v) for 2 days followed by chloroform: acetone: methanol (4:1:3 v/v/v) for 2 days. The solvent was replaced every day. Finally, the feathers were washed several times with tap water to eliminate the solvent residual, dried for 3 days at 50°C and grinded using electrical blinder (model Hg135XP). Grinded keratin which looked like cotton was maintained in a sterile bottle at 4°C until used.
Bacterial Isolation, Purification and Maintenance
The feather meal medium (basic medium) used for isolation and growth of the feather-degrading microorganisms contained the following constituents (g L-1): NaCl (0.5), KH2PO4 (0.7), K2HPO4 (1.4), MgSO4 (0.1) and ground keratin (10) as sole carbon and nitrogen source. To prepare the solid medium 15-20 g agar were used. The pH was adjusted to 7.2 ( Lin et al., 1992). Five plates of the basic medium were used for each sample. From the suitable solution, 1 mL was spread on the feather meal plates which were incubated at 37°C for 5-7 days. After incubation, colonies were picked up using sterile needle and streaked again on the same medium until complete purification of the colonies which maintained on nutrient agar medium at 4°C.
Measurement of Bacterial Growth
The abilities of bacterial isolates to produce keratinase were confirmed by cultivation on agar plates containing skimmed milk. Skimmed milk plates were composed of (g L-1): Tryptone (5.0), Yeast Extract (2.5), Dextrose (1.0), Skimmed Milk Powder (1.0). To observe the zone of hydrolysis, 10% (v/v) Trichloroacetic Acid (TCA) was flooded on the milk agar plate (Saran et al., 2007). The bacterial growth in feather meal broth medium was measured by counting the colony forming units (cfu mL-1) on nutrient agar.
Identification of Bacterial Isolate Kera MS21
Morphological characterization of the best keratinase-producing organism was studied. Different physiological testes and carbohydrates fermentation were carried out using API 20E. Oxidase and catalase production by the selected bacterial isolate were also performed. Sensitivity to different antibiotics was tested and presence of inhibition zones indicated the sensitivity to the antibiotic present.
Electron Microscopy
To prepare specimens for transmission electron microscopy, cells were harvested by centrifugation at 5,000x g for 10 min and then fixed overnight in a solution containing 2.5% (w/v) glutaraldehyde in 0.1 M Na2HPO4/KH2PO4 (pH 7.2) buffer at 4°C and post-fixed with 1% osmium tetroxide (OsO4). The cells were then dehydrated with ethanol and embedded in Spurr. The sections were stained with 1% (w/v) uranyl acetate and 1% (w/v) citrate and examined with a Philips model CM-10 electron microscope at an accelerating voltage of 80 KV. The examination was performed at National Research Center, Dokki, Egypt.
Phylogenetic Analysis of 16S rDNA Sequence
Genomic DNA from the selected isolate kera MS 21 was obtained by using QIAamp DNA Mini Kit (QIAGEN, USA). Three microliter of genomic DNA were amplified in a 100 μL reaction by using the hot star master mix from QIAGEN (2X). The primers were designed based on the highly conserved region of 16S rDNA from various bacteria (Weisberg et al., 1991). The negative control was done using the hot star master mix with the two primers only.
The 16S rDNA gene was amplified by PCR using forward primer 5'- AGTTTGATCATGGTCAG-3' and reverse primer 5'-GGTTACCTTGTTACGACT 3’. The PCR was performed by using DNA thermal cycler (Perkin Elmer, USA). Conditions consisted of one cycle with 5 min DNA denaturation at 95°C, 35 cycles of 95°C (2 min), 48°C (1 min) and 72°C (4 min), plus one additional cycle with a final 20 min chain elongation at 72°C. The PCR products were purified using QIAquick PCR purification kit (QIAGEN). The PCR product was analyzed in 1.5% agarose gel stained with ethidium bromide. Sequencing analysis step was performed at Gene Analysis unit, VACSERA. Cycle sequencing was carried out using a Bigdye terminator cycle sequencing kit (Applied Biosystems, Foster City). Sequencing products were resolved on an applied Biosystems Model 310 automated genetic analyzer (Perkin Elmer, USA). Data were submitted to GenBank database. The DNA sequence was compared to the GenBank database in the National Center for Biotechnology Information (NCBI) using the BLAST program. A blast search was used to find close relations through sequence similarity. The phylogenetic and molecular evolutionary analyses of the keratin-degrading bacteria based on 16S r-RNA genes were conducted using MEGA version (Tamura et al., 2007).
Keratinase Assay
Keratinase activity was determined by the modified method of Letourneau et al. (1982). An enzyme sample (0.2 mL) was incubated with 0.8 mL of 0.4% (w/v) of keratin azure (Sigma Aldrich, Germany) in 50 mM phosphate buffer at 60°C with a pH 8.5 and 150 rev min-1 for 1 h. One unit of keratinase activity was defined as the amount of enzyme that resulted in an increase in absorbance at 595 nm (A595) of 0.1 after reaction with the keratin azure for 1 h (Scott and Untereinen, 2004).
Keratinase Purification
Bacterial isolate kera MS 21 was cultivated in feather meal broth for 72 h at 37°C. Culture medium was prefilterated through glass wool to remove undegraded feathers. Bacterial cells were removed by using centrifuge under cooling (4°C) at 5000 rpm for 10 min. Protein was precipitated by adding solid ammonium sulphate (40%) at 4°C for 24 h. The precipitate was collected by centrifugation at 5000x g for 40 min and was dissolved in a minimal volume of 25 mM potassium phosphate buffer, pH 5.8, the enzyme solution was desalted by dialysis with a cellulose dialysis bag (3500 Da pore-size) in 25 mM potassium phosphate buffer at pH 5.8. After dialysis, the enzyme was dissolved in 2 mL dist. water and applied on carboxymethyl cellulose. The active fractions were collected, lyophilized and applied on sephadex G75. The active fractions were used to determine the molecular weight and enzyme assays. Protein content (mg mL-1) was determined by Bradford assay (Bradford, 1976).
Molecular Weight Determination
Molecular weight of keratinase was detected according to Laemmli (1970) on a 0.75 mm thick polyacrylamide slab gel (7-8 cm) using mini gel system Bio-Rad. Low molecular weight protein standard (Amersham Pharmacia) ranged from 14-97 KD was used to determine the molecular weight of the purified keriatinase. The protein markers were run along side of the sample (25 μg protein sample was used) and protein bands were visualized by Coomassie blue R350 staining method.
Effect of Different pH, Temperature, Some Enzyme Additives and Inhibitors on Enzyme Activity
The effect of pH on enzyme activity was studied by using 50 mM Tris/HCl (pH 4-8) and 50 mM potassium phosphate (pH 8-10). The effect of temperature was studied with a temperature range from 25 to 80°C. The experiments were carried out by incubating the enzyme solution in different pH and at different temperature for 60 min and the enzyme activity was measured. The effect of some additives (glycerol, mannitol and sorbitol), metal ions (including 10 mM of MnCl2.4H2O, ZnCl2, HgCl, BaCl.2H2O, CuSO4.5H2O, MgCl2 or CaCl2 and inhibitors (0.5% Triton X 100, 10 mM EDTA, 0.1, 0.5% DMSO, 0.5% isopropanol, or 0.5% mercaptoethanol) on enzyme activity were studied as described by El-Sabbagh et al. (2003) and Tapia and Simoes (2008).
Statistical Analysis
Each experiment had three replicates. Means of variable and standard deviation were recorded and t-student test was carried out to detect any significant differences between the results of control and the treated samples.
RESULTS
Screening for Feather-Degrading Bacteria
Twenty three keratinase producing bacteria were isolated from different sources on medium containing feather meal as sole carbon and nitrogen source. The source of isolation, Gram reaction and growth using different media were shown in Table 1. Eight isolates were from soil, 7 isolates were from marine water, 3 isolates were from feather, 3 isolates were from poultry wastes and one isolate from fodder and one isolate from air. Three isolates were belonging to Gram negative bacteria and 20 isolates were Gram positive bacteria. All isolates were screened for their proteolytic activities on skimmed milk agar plates. The isolates kera MS 3, kera MS 13, kera MS 15, kera MS 17, kera MS 19 and kera MS 21 showed the highest growth rate on keratin meal agar and skimmed milk agar. The largest clearing zone on skimmed milk was observed for isolate kera MS21, with a zone diameter equal 2.8 cm.
| Table 1: | The isolated keratinase producing bacterial strains on feather meal agar, the source of isolation, Gram reaction and the growth of bacterial isolate, on feather meal agar, number of cell forming units mL-1 and diameter of the clear zone on skimmed milk plates |
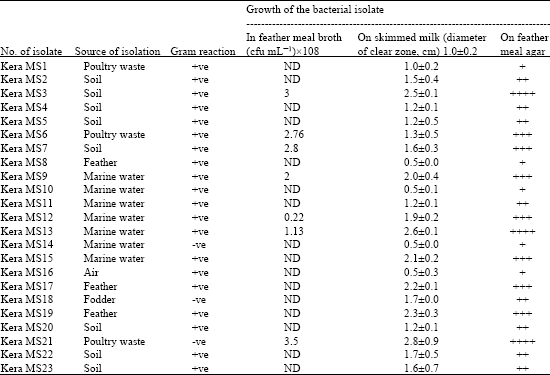 | |
| *The differences in the mean values of the growth on skimmed milk among the isolates is statistically highly significant difference (p<0.001) | |
| Table 2: | Keratinolytic activity of some bacterial isolates in keratin meal broth medium at different incubation period |
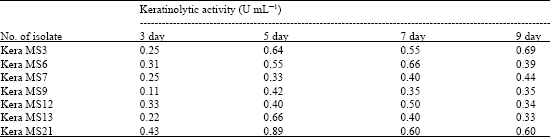 | |
 | |
| Fig. 1: | (a) Growth of kera MS 21 on skimmed milk agar plate at 37°C for 24 h and (b) ker MS 21 cultivated in feather meal broth at 37°C for one week |
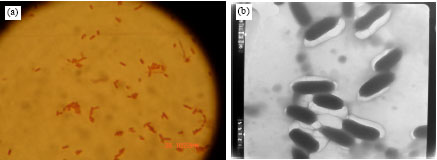 | |
| Fig. 2: | (a) Gram staining of kera MS 21 under light microscope and (b) transmission electron microscope X 60 000 of kera MS 21 |
The diameter of the obtained inhibition zones varied significantly with the tested organisms at p-values <0.05. The quantities of keratinase were determined for these isolates. The isolate kera MS21 was the most active isolate in keratinase production after 5 days of growth followed by kera MS 3, kera MS13 and kera MS6 (Table 2). Kera MS21 was selected for more studies.
Identification and Characterization of Feather Degrading Isolate (Kera MS21)
Microscopic observation of the isolate showed a straight rod shaped bacterium, non sporulated, non acid fast and Gram negative cells. It has milky like colonies with lobated margin on feather meal agar (Fig. 1a, b). The bacterium grew aerobically, strongly catalase, oxidase positive and was highly motile. The bacterial dimensions were ranged from 0.3-0.5 and 0.7-1.1 μm and a polar flagellum was observed by transmission electron microscopy as shown in Fig. 2a and b.
| Table 3: | Physiological characteristics of keratinase-producing bacterial (kera MS21) |
 | |
| Table 4: | Purification of keratinase enzyme from Pseudomonas sp., MS21 |
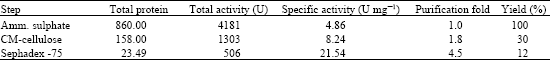 | |
Additional morphological, physiological and biochemical tests were conducted for ker MS21 and summarized in Table 3. The morphological examinations and physiological tests (API 20E: Analytical profile index) proposed that kera MS21 belongs to Pseudomonas group. It was identified as Pseudomonas sp., MS21. Characters of the selected Pseudomonas sp., MS21 were compared with data from Bergey’s Manual of Systematic Bacteriology (Bergey, 1984). These results were confirmed using 16S rDNA studies. The 16S rDNA sequence was compared to the GenBank database in the National Center for Biotechnology Information (NCBI) using the BLAST program. The 16S rDNA sequence of this isolate kera MS21 showed high levels of sequence similarity with members of the genus Pseudomonas, such as Pseudomonas sp., CL3.5 (95%), Pseudomonas sp., CL3.1 (94%), Pseudomonas putita strain FW C30 (94%) Pseudomonas sp. me-11 (91%) and Pseudomonas lini (90%) (Fig. 3).
Purification and Molecular Weight of Keratinase Enzyme
Purification of microbial enzyme was carried out using different column chromatography and the purified enzyme had more specific activity after the final step. Purification was carried out using two types of column chromatography. Total and specific activities were summarized in Table 4. Importantly, the second column sephadex G75 column chromatography used for enzyme purification provided the best yield with an increased purity. This step eliminated proteins that might inhibit keratinase activity. A single protein band was shown on SDS-PAGE gel electrophoresis at 30 KDa (Fig. 4).
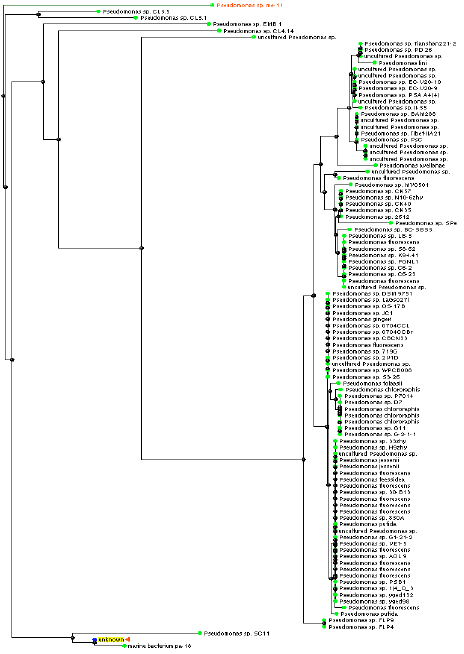 | |
| Fig. 3: | Phylogenetic tree based on 16S rDNA sequence comparisons of strain kera MS2 and selected bacteria. The branching pattern was generated by fast minimum evolution tree method, maximum sequence difference = 0.05 |
Characterization of the Purified Enzyme
The purified enzyme was suspended in buffers at different pH values; the highest activity was recorded at pH 8.0 (Fig. 5a). The effect of different temperature on the enzyme activity was estimated, the best activity was detected at 37°C (Fig. 5b). The effect of some additives was achieved in Table 5. The maximum activity, found using glycerol (0.5%) and sorbitol (0.5%), have significant values compared with the control at p<0.05. In contrast, mannitol and EDTA mainly at 10 mM concentration had an inhibitory effect.
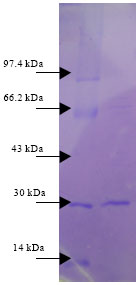 | |
| Fig. 4: | Molecular weight of keratinase isolated from Pseudomonas sp., by SDS-PAGE, first lane: molecular mass marker proteins. Rabbit phosphorylase B (97.4 KDa), bovine serum albumin (66.2 KDa), rabbit actin (43 KDa), bovine carbonic anhydrase (30 KDa) and hen egg white lysozyme (14 KDa). Second lane: Purified keratinase |
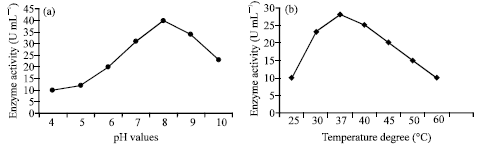 | |
| Fig. 5: | Effect of pH and temperature on keratinase activity using keratin azure as substrate |
| Table 5: | Effect of some chemicals or additive on keratinolytic activity |
 | |
| *Significant results using studied-t-test at p<0.05 compared with control without addition | |
No significant effect on keratinase activity was found by the addition of triton X100, EDTA mainly at 10 mM concentration, isopropannol or mercaptanol. The effects of some metal ions on enzyme activity are monitored on. The results indicated that the activity of the enzyme was slightly reduced by the following metal ions; CaCl2 and ZnCl2. No significant effect on keratinase activity was found by the addition of MnCl2, HgCl2, BaCl2, CuSO4 and MgCl2.
DISCUSSION
Keratinaceous materials such as feather, wool and hair are insoluble and resistant of degradation by common proteolytic enzymes. Feathers contain over 90% protein, the major component being a keratin, a fibrous and insoluble structural protein extensively cross-linked by disulfide, hydrogen and hydrophobic bonds. Owing to their insoluble nature, feathers are resistant to degradation by common microbial proteases. Thus, the several million tons of feathers generated annually by the poultry industry leads to troublesome environmental pollution and wastage of protein-rich reserve (Onifade et al., 1998; Gousterova et al., 2005; Grazziotin et al., 2006). Keratinase enzymes have been found to be an effective feather degrading enzyme useful for degradation of feather, hair and chicken wastes. It can act converting keratin (insoluble materials) into soluble amino acids which can be introduced into animal feeds. The Keratin-degrading microorganism thrives under different ecological and environmental conditions including soil, air, fodder, sand and different environmental wastes ( Kansoh et al. (2009)). In this study, we tried to isolate and identify a new local bacterial strain, able to completely degrade keratin rich wastes into soluble and useful materials and can be used in many proposes. Out of 24 bacterial isolates recovered from different habitat, the isolate kera MS21, which was isolated from poultry waste, has been found to be the best producer of keratinase. This may be due to its high metabolic rate and adaptation to use keratin as carbon and nitrogen source. Several bacterial isolates belongs to the genus Bacillus ( Lin et al., 1992) and Streptomyces sp. (Bockle et al., 1995; Bressollier et al., 1999; Kansoh et al. (2009)) were isolated from chicken waste feathers at 37°C and pH 7. It degraded feather keratin by using it as a primary source of energy, carbon, nitrogen and sulfur. It was clear that presence of feather in the growth medium act as an enzyme inducer. Riffel and Brandelli (2006) reported that keratinase producing bacteria had different characteristics, such as a broad temperature range of growth. The optimal proteolytic activities were detected between 30 and 37°C (Atalo and Gashe, 1993). An optimum keratin-degrading activity at mesophilic temperatures should be a desirable characteristic because these microorganisms may achieve hydrolysis with reduced energy input. The identification of this bacterial isolate (Kere MS21) was based on cell morphology, colony morphology, growth characteristics and several physiological and biochemical tests. Microscopic observation of the isolate showed a straight rod without endospores and it was Gram negative and non-acid fast. Preliminary identification tests, physiological tests and API 20E indicate that strain kera MS21 belongs to the proteobacteria. The isolated strain was oxidase and catalase positive; it belongs to Pseudomonas group (Brenner et al., 2005) and it was identified as Pseudomonas sp., MS21. Phylogenetic analysis using small subunit rDNA is considered the most reliable method for determination of the taxonomic relationships between microorganisms available at present (Iwamoto et al., 2001). The results of 16S rDNA sequence of this isolate showed high levels of sequence similarity with members of the genus Pseudomonas, such as Pseudomonas sp., CL3.5 (95%), Pseudomonas sp., CL3.1 (94%), Pseudomonas putita strain FW C30 (94%) Pseudomonas sp., me-11 (91%) and Pseudomonas lini (90%). Thus, Strain Kera MS21 was belonging to genus Pseudomonas and identified as Pseudomonas sp., MS21.
Similar results were found by Burrell and MacDiarmid (1984). They isolated Pseudomonas aeruginosa from fleece rot which was a producer of keratinase. Studies on Gram-negative bacteria are relatively scarce and keratinase activity has been described for Vibrio sp. (Sangali and Brandelli, 2000) Chryseobacterium sp. ( Riffel et al., 2003) Xanthomonas sp. (De-Toni et al., 2002) and Burkholderia, Chryseobacterium, Pseudomonas (Riffel and Brandell, 2006). For this reason, Gram negative isolate was selected for the further work.
The molecular weight Pseudomonas keratinase was detected to be 30 KDa and maximum enzyme activities were detected at pH 8 and 37°C. Similar results were obtained for Bacillus licheniformis PWD where its keratinase has 33 KDa ( Lin et al., 1992) and the maximum activity of the enzyme at pH 6.5 and 50°C. Cao et al. (2009) isolated keratinase with 32.3 KDa from Stenotrophomonas maltophilia grown on raw feathers at different temperatures and initial pH. They added that, maximum growth and feather-degrading activity of the bacterium were observed at 40°C and initial pH ranging from 7.5 to 8.0. Lin et al. (2009) isolated keratinase which was nonglycosylated protein with a molecular mass of 33 KDa and had an optimal pH and temperature at 8.0 and 60°C, respectively by expression of Keratinase-coding gene from Pseudomonas aeruginosa in Pichia pastoris.
Anionic detergent such as Triton X-100 and the organic solvent DMSO increased the keratinolytic activity. On contrast, β-mercaptanol and isopropanol had no clear effect but EDTA deceased significantly the keratinase activity. Present results are similar to that obtained by Korkmaz et al. (2003) using Streptomyces strain BA7. An increase in the enzyme activity was also noticed when DMSO was used (Bockle et al., 1995) and an antagonistic effect was found with EDTA. Keratinase activity was decreased significantly by some metal ions, especially CaCl2 and ZnCl2. On contrast, MgCl2, BaCl.2H2O, HgCl2 and MnCl2.4H2O did not show any relevant effect. Results of Lin et al. (2009) reported that Ba2+, Ca2+, Mg2+, Mn2+, Zn2+, dithiothreitol, glutathione and β-mercaptoethanol activated, while Cu2+, Fe2+, Hg2+, Fe3+, ethylene glycol tetraacetic acid, ethylene diamine tetraacetic acid and p-chloromercuribenzoate inhibited keatinase activity.
From the obtained results, we can conclude that the chicken wastes are rich sources of bacteria which hydrolyze keratin. Present isolated strain can utilized in large to overcome the problem of environmental contamination with poultry wastes.
ACKNOWLEDGMENT
We would like to express our gratitude to all those who gave us the possibility to complete this project. We want to thank the Deanship of Scientific Research and Research Center and Recommendations, King Abdul Aziz University, Ministry of Higher Education for the finical supports of this project (No. 428/523), great encouragement, keep interest and their efforts during writing and revising of the manuscript. I would like to give my special thanks to Mrs. Fizah Bachimail for her assistance in the practical work.
REFERENCES
- Atalo, K. and B.A. Gashe, 1993. Protease production by a thermophilic Bacillus species (P-001A) which degrades various kinds of fibrous proteins. Biotechnol. Lett., 15: 1151-1156.
CrossRefDirect Link - Bockle, B., B. Galunsky and R. Muller, 1995. Characterization of a keratinolytic serine proteinase from Streptomyces pactum DSM 40530. Applied Environ. Microbiol., 61: 3705-3710.
PubMedDirect Link - Brandelli, A. and A. Riffel, 2005. Production of an extracellular keratinase from Chryseobacterium sp. growing on raw feathers. Electron. J. Biotechnol., 8: 35-42.
Direct Link - Bressollier, P., F. Letourneau, M. Urdaci and B. Verneuil, 1999. Purification and characterization of a keratinolytic serine proteinase from Streptomyces albidoflavus. Applied Environ. Microbiol., 65: 2570-2576.
Direct Link - Burrell, D.H. and M.A., MacDiarmid, 1984. Characterization of isolates of Pseudomonas aeruginosa from sheep. Aust. Vet. J., 61: 277-279.
PubMed - Cao, Z., Q. Zhang, D. Wei, D. Chen, J. Wang, X. Zhang and M. Zhou, 2009. Characterization of a novel Stenotrophomonas isolate with high keratinase activity and purification of the enzyme. J. Indian Microbiol. Biotechnol., 36: 181-188.
PubMed - De Toni, C.H., M.F. Richter, J.R. Chagas, J.A.P. Henriques and C. Termignoni, 2002. Purification and characterization of an alkaline serine endopeptidase from a feather-degrading Xanthomonas maltophilia strain. Can. J. Microbiol., 48: 342-348.
Direct Link - Gousterova, A., D. Braikova, I. Goshev, P. Christov and K. Tishinov et al., 2005. Degradation of keratin and collagen containing wastes by newly isolated thermoactinomycetes or by alkaline hydrolysis. Lett. Applied Microbiol., 40: 335-340.
CrossRefDirect Link - Grazziotin, A., F.A. Pimentel, E.V. de Jong and A. Brandelli, 2006. Nutritional improvement of feather protein by treatment with microbial keratinase. Anim. Feed Sci. Technol., 126: 135-144.
CrossRefDirect Link - Iwamoto, K., T. Yoshikawa, T. Sakata and H. Maeda, 2001. Phylogenetic analysis of marine algicidal filamentous bacteria inferred from ssu rdna and intergenic spacer regions. Microbes Environ., 16: 213-221.
CrossRef - Korkmaz, H., M.N. Unaldi, B. Aslan, G. Coral, B. Arikan, S. Dincer and O. Colak, 2003. Keratinolytic activity of Streptomyces satrin BA7 a new isolate from Turkey. Ann. Microbiol., 53: 85-93.
Direct Link - Laemmli, U.K., 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227: 680-685.
CrossRefDirect Link - Langeveld, J.P.M., J.J. Wang, D.F.M. van de Wiel, G.C. Shih, J. Garssen, A. Bossers and J.C.H. Shih, 2003. Enzymatic degradation of prion protein in brain stem from infected cattle and sheep. J. Infect. Dis., 188: 1782-1789.
Direct Link - Lin, H.H., L.J. Yin and S.T. Jiang, 2009. Functional expression and characterization of keratinase from Pseudomonas aeruginosa in Pichia pastoris. J. Agric. Food Chem., 57: 5321-5325.
CrossRefDirect Link - Lin, X., C.G. Lee, E.S. Casale and J.C.H. Shih, 1992. Purification and characterization of a keratinase from a feather-degrading Bacillus licheniformis strain. Applied Environ. Microbiol., 58: 3271-3275.
Direct Link - Lin, X., J. Shih and H.E. Swaisgood, 1996. Hydrolysis of feather keratin by immobilized keratinase. Applied Environ. Microbiol., 62: 4273-4275.
PubMedDirect Link - Onifade, A.A., N.A. Al-Sane, A.A. Al-Musallam and S. Al-Zarban, 1998. A review: Potentials for biotechnological applications of keratin-degrading microorganisms and their enzymes for nutritional improvement of feathers and other keratins as livestock feed resources. Bioresour. Technol., 66: 1-11.
CrossRefDirect Link - Riffel, A. and A. Brandelli, 2006. Keratinolytic bacteria isolated from feather waste. Braz. J. Microbiol., 37: 395-399.
CrossRefDirect Link - Riffel, A., F. Lucas, P. Heeb and A. Brandelli, 2003. Characterization of a new keratinolytic bacterium that completely degrades native feather keratin. Arch. Microbiol., 179: 258-265.
CrossRefDirect Link - Sangali, S. and A. Brandelli, 2000. Feather keratin hydrolysis by a Vibrio sp. strain kr2. J. Applied Microbiol., 89: 735-743.
CrossRef - Scott, J.A. and W.A. Untereinen, 2004. Determination of keratin degradation by fungi using keratin azure. Med. Mycol., 42: 239-246.
CrossRef - Tamura, K., J. Dudley, M. Nei and S. Kumar, 2007. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol., 24: 1596-1599.
CrossRefPubMedDirect Link - Tapia, D.M.T. and M.L.G. Simoes, 2008. Production and partial characterization of keratinase produced by a microorganism isolated from poultry processing plant wastewater. Afr. J. Biotechnol., 7: 296-300.
Direct Link - Weisberg, W.G., S.M. Barns, D.A. Pelletier and D.J. Lane, 1991. 16S ribosomal DNA amplfication for phylogenetic study. J. Bacteriol., 173: 697-703.
PubMed - Bradford, M.M., 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem., 72: 248-254.
CrossRefPubMedDirect Link








