Research Article
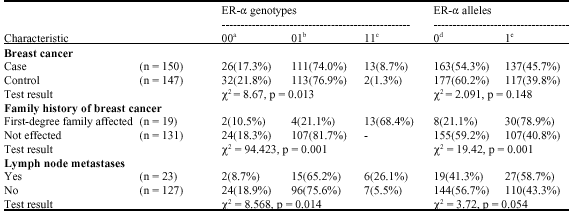
Estrogen Receptor-α Gene Codon 10 (T392C) Polymorphism in Iranian Women with Breast Cancer: A Case Study
Department of Biomedical Science, Faculty of Medicine and Health Sciences, Universiti Putra Malaysia, Malaysia
C. Azimi
Department of Genetics, Cancer Institute, School of Medicine, Imam Khomeini Hospital Complex, Tehran University of Medical Sciences, Tehran, Iran
F. Othman
Department of Biomedical Science, Faculty of Medicine and Health Sciences, Universiti Putra Malaysia, Malaysia
M.R. Noori Daloii
Department of Medical Genetics, School of Medicine, Medical Sciences/University of Tehran, Tehran, Iran
Z.O. Ashtiani
Department of Medical Genetics, School of Medicine, Medical Sciences/University of Tehran, Tehran, Iran
M. Mojarrad
Department of Medical Genetics, School of Medicine, Medical Sciences/University of Tehran, Tehran, Iran
S.A. Oskouei
Department of Medical Genetics, School of Medicine, Medical Sciences/University of Tehran, Tehran, Iran
F.M. Nejad
Department of Medical Genetics, School of Medicine, Medical Sciences/University of Tehran, Tehran, Iran
P. Ismail
Department of Biomedical Science, Faculty of Medicine and Health Sciences, Universiti Putra Malaysia, Malaysia










Mina Rasoli Reply
It was one best detaid article that I was looking for. I am interested in this subject in my reasearch work. It could be much help for me.
mohammad mehdi
iwill work on leptin GenePolymorphism in Iranian Women with Breast Cancer.please help.