Review Article
Clinical Applications of Epigenetic Markers in Diagnosis and Treatment of Cancer
Department of Biochemistry, Afzalipour School of Medicine, Kerman University of Medical Sciences, Kerman, Iran
Epigenetic changes, unlike genetic alterations defined as heritable changes in gene expression that are potentially reversible and have essential roles during embryonic development, however appear to contribute to the malignant transformation and progression of cancer (Wolffe and Matzke, 1999; Li, 2002). There are three main types of epigenetic information in the genome which are having important role in the regulation of gene expression as follow. Firstly, DNA methylation that is a covalent modification of DNA in which a methyl group is transferred from S-adenosyl-methionine to the C-5 position of cytosine by a family of cytosine methyl-transferases (DNAMTs). DNA methylation occurs almost exclusively at CpG nucleotides. Secondly, Histone modifications which are including acetylation, methylation and phosphorylation which are stably maintained during cell division. Finally, Genomic imprinting which is parent-of-origin-specific allele silencing or relative silencing of one parental allele compared with the other parental allele (Vaziri Gohar and Mohammadi, 2010).
The goal of this study is the background, promise, status of the applications of epigenetic alterations for the early detection, prevention, diagnostics, therapeutics and risk prediction of cancer.
DNA methylation: DNA methylation in mammalian genomes is a defense mechanism by which repetitive DNA which accounts for at least 50% of genome’s content is transcriptionally silenced to prevent it from propagating (Urnov, 2002).
Methyl-cytosine residues are often found in short stretches of CpG-rich regions (CpG islands) that are 0.5-2 kb long and found in the 5′-region of approximately 60% of genes (Gardiner-Garden and Frommer, 1987). When a CpG site is methylated, cytosines on both DNA strands are methylated. At DNA replication, the methylated status is transmitted to daughter DNAs by maintenance DNA methyl-transferase (DNMT), which is present at a replication fork and recognizes hemi-methylated CpG sites. Although a newly synthesized DNA strand does not have methyl groups, maintenance DNA methyl-transferase at the replication fork transfers a methyl group to the newly synthesized strand. Therefore, the methylated or unmethylated status of CpG sites is faithfully copied into daughter DNA in somatic cells (Jones and Baylin, 2002; Herman and Baylin, 2003).
Most CpG islands are unmethylated, with the exception of certain imprinted genes and genes on the inactive X chromosomes of females (Bird, 1986). DNA methylation aberrations can occur as either hypo- or hypermethylation. Both forms can lead to chromosomal instability and transcriptional gene silencing (Baylin et al., 1991).
DNMT1 is referred to as the maintenance methylase due to its preference for hemimethylated CpG sites in DNA (Pradhan et al., 1999). DNMT3a and DNMT3b are considered to be de novo methylases because they can methylate unmethylated DNA (Okano et al., 1999; Pradhan et al., 1999). However, all three DNMTs have been shown to act cooperatively and the functional differences between the methylases may to a large extent be due to the genomic regions that they act upon (Liang et al., 2002; El-Osta, 2003). Over-expression of the DNA methyltransferases 1 and 3A was found in the bone marrow of patients with myelodysplastic syndrome (MDS). Upregulation of DNMTs has also been shown in prostate cancer cell lines and tissues (Patra et al., 2002; Langer et al., 2005).
For detection of cancer cells in body fluids, a high-sensitivity method is necessary. One way is mutation detection in cells. Indeed, because the exact location of a mutation within a gene is usually unknown, many primer sets are necessary for complete analysis. In contrast, aberrant methylation of DNA molecule of cancer cells, even in very few in number, can be sensitively detected by using Methylation-Specific PCR method (MSP), only with one set of PCR primer can be performed on chemically stable DNA, not on RNA (Herman et al., 1996; Laird, 2003).
DNA hypermethylation: Methylation of unmethylated DNA, also known as hypermethylation, can repress the gene expression. The gene repression is caused by changes in chromatin structures due to binding of specific proteins to methylated DNA. This binding leads to decreased affinity for binding of some transcriptional factors to methylated CpG sites. Inappropriate silencing of genes can contribute to cancer initiation, progression, pathologic grade, invasion and metastasis (Ushijima et al., 2003; Laird et al., 2004). Nearly, all types of cancers have transcriptional inactivation of tumor suppressor genes due to DNA hypermethylation (Costello et al., 2000; Herman and Baylin, 2003; Ransohoff, 2003; Jones and Baylin, 2007).
DNA hypomethylation: Demethylation of normally methylated DNA, also known as hypomethylation, can disrupt such a defense mechanism, leading to structural and functional alterations of the genome. There are two types of hypomethylation: global and gene-specific hypomethylation, which refers to an overall decrease of 5-methyl-cytosine content in the genome or a gene, respectively (Dunn, 2003). Both global- and gene-specific hypomethylation have hazardous effects and may contribute in human cancer, like DNA hypermethylation. Global DNA hypomethylation has also been found in the premalignant or early stages of some neoplasms. Also, net decreases in the content of methyl-cytosines, often exceed the localized increases in DNA methylation. However, it is unclear whether this epigenetic alteration is a cause or consequence of tumorigenesis and also, whether hypomethylation induced by disrupting DNMT1 does inhibit or promote tumor growth (Feinberg et al., 1988; Cravo et al., 1996; Baylin et al., 2000; Robertson, 2001).
In a murine model of intestinal neoplasia, mice carrying a germ-line mutation in the APC gene (APCMin/_) crossed with mice heterozygous for the DNMT1 mutation had substantially fewer tumors than Min mice with wild-type DNMT1 (Laird et al., 1995; Cormier and Dove, 2000). In contrast, genomic hypomethylation has been associated with the induction of T-cell lymphomas in mice carrying a hypomorphic DNMT1 allele, which reduces DNMT1 expression to 10% of wild-type levels and results in substantial genome-wide hypomethylation in all tissues. So, hypomethylation-induced cancer might be related to differences in model systems or tissue specificity (Lengauer, 2003).
It is possible that demethylation could lead to the activation proto-oncogenes (Ehrlich, 2002; Fruhwald and Plass, 2002; Nishigaki et al., 2005). Also, genome-wide demethylation may promote genomic instability possibly by activating global demethylation of repetitive sequences such as satellite DNAs can lead to increased chromosomal rearrangements (Dunn, 2003).
Compared with adjacent normal tissues, different types of cancer tissues contain hypomethylated c-jun and c-MYC proto-oncogenes in liver cancer (Tsujiuchi et al., 1999; Tao et al., 2000), pS2 gene in breast cancer (Fruhwald and Plass, 2002) and PLAU gene in the prostate cancer (Van Veldhuizen et al., 1996).
Demethylating agents: Considering that some aberrant DNA methylation is present in early stages of carcinogenesis, there is a possibility that such demethylating agents may protect against some cancers (Laird et al., 1995). Demethylating agents are including DNMT1 inhibitors group (Azacitadine, Decitabine, Zebularine and MG98), procainamide, procaine and EGCG (epigallocatechin-3-gallate) (Fang et al., 2003; Villar-Garea et al., 2003). Inhibitors of DNMTs have been widely used in cell culture systems to reverse abnormal DNA hypermethylation and restore silenced gene expression. However, only limited success has been achieved in clinical trials with these drugs (Thibault et al., 1998; Goffin and Eisenhauer, 2002). Also, nucleosides analog inhibitors of DNMTs may promote genomic instability and increase the risk of cancer in other tissues, because have many potential side effects such as myelotoxicity, mutagenesis and tumorigenesis (Jones and Taylor, 1980; Jackson-Grusby et al., 1997; Gaudet et al., 2003). So, there is an attractive alternative for possible clinical use of non-nucleoside analog DNMT inhibitors.
The use of these drugs raises questions regarding their potential to affect non-cancerous cells epigenetically. However, normal cells divide at a slower rate than malignant cells and incorporate less of these drugs into their DNA resulting in less of an effect on DNA methylation. As shown in Table 1, methylation pattern of multiple genes have higher role in diagnostic and prognostic possibilities than that of a single gene. However, long-term negative effects of DNA methylation inhibitors in patients have not been found to date (Yang et al., 2003).
Azacitadine and decitabine are labil and have acute hematological toxicities. A next generation DNA methylation inhibitor, such as zebularine, might possibly overcome these problems (Marquez et al., 2005; Yoo et al., 2008). Also, the non-nucleoside analogue inhibitors are not as potent as the nucleoside analogues and therefore this issue needs for improvement (Chuang et al., 2005).
DNA methylation as a marker: It has been reported that the pattern of aberrant methylation of individual- or multiple genes can be associated with clinically useful information, such as cancer risk assessment, cancer prognoses, early detection and responses to therapeutics. Therefore, these features make DNA hypermethylation an excellent tumor biomarker candidate (Herman and Baylin, 2003).
In Table 2, aberrant DNA methylation can be applied to detect cancer cells or cancer-derived DNA cancer and diagnostics in several ways. Firstly, if aberrant methylation of some CGIs is specifically present in cancer cells, it can be used to detect cancer cells in biopsy samples or cancer-derived free DNA in plasma. On the other hand, incidences of aberrant methylation of specific CGIs are higher than those of mutations.
| Table 1: | DNA methylation-related drugs |
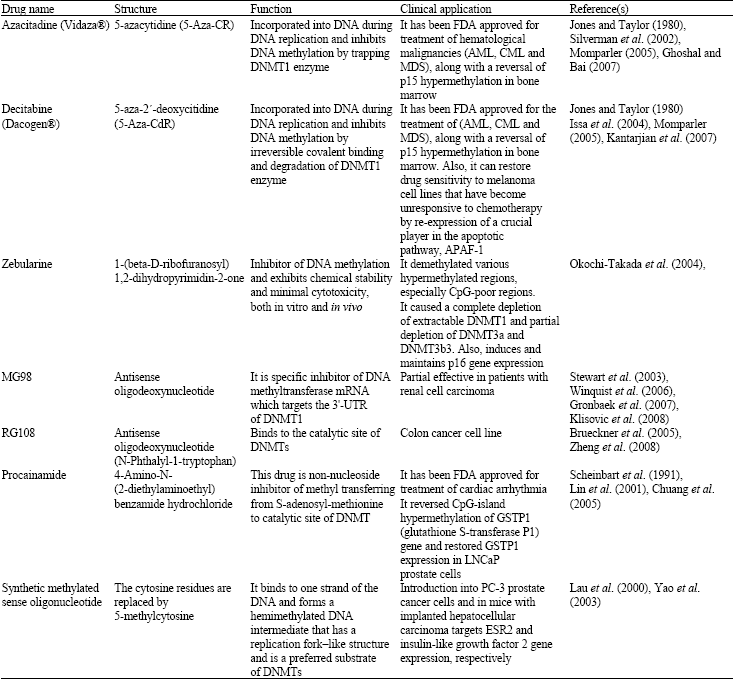 | |
| Table 2: | Clinical association of DNA methylation in diagnostic and prognostic of cancer |
 | |
Secondly, if aberrant methylation of some CGIs is associated with a disease phenotype, such as prognosis, responses to chemotherapies or occurrence of adverse effects, it can be used as a marker to predict the phenotype. Finally, if aberrant methylation of some CGIs in non-cancerous tissue is associated with a risk for cancer development, it can be used as a cancer risk marker (Kaneda et al., 2002; Miyamoto et al., 2003; Hagihara et al., 2004).
Histone modification: The N-terminal tails of histones, positioned peripheral to the nucleosome core, are subject to various covalent modifications, such as acetylation, methylation, phosphorylation and ubiquitination by specific chromatin modifying enzymes (Zhang and Reinberg, 2001).
The pattern of these modifications has been referred to as the histone code, and it acts as a second layer of epigenetic regulation of gene expression affecting chromatin structure and remodeling (Jenuwein and Allis, 2001). Histone modification is closely associated with DNA methylation status and is important for gene regulation (Jaenisch and Bird, 2003).
Histone methylation: Histone methylation of histone H3-k4 (Lys4) is associated with active gene transcription. But methylation of histone H3-k9 (Lys9) and H3K27 are normally present at transcriptionally inactive or heterochromatic regions is associated with gene repression (Cao et al., 2002; Nguyen et al., 2002). Lysine residues (lys) can accept up to three methyl groups, which are added by various histone methyltransferases (HMTs) (Fischle et al., 2003; Santos-Rosa and Caldas, 2005).
The methyltransferase MLL, which methylates H3K4, is involved in translocations that lead to the inappropriate expression of various homeotic (Hox) genes, which contributes to leukemic progression (Krivtsov and Armstrong, 2007).
The Polycomb group (PcG) complexes are chromatin modifiers that are crucial to development and have been implicated in the development of cancer (Tonini et al., 2008). These negative regulators of gene expression are very important in sustaining the repressive state of their target genes through the cell cycle (Kingston et al., 1996). Two of the PcG repressive complexes (PRC1 and PRC2) have both been shown to be involved in various cancers. Enhancer of zeste homologue 2 (EZH2), a component of PRC2 with H3K27 methyltransferase activity, is upregulated in mantle cell lymphoma, breast and prostate cancer (Visser et al., 2001; Kleer et al., 2003). RING1, a component of PRC1 that aids in the ubiquitylation of histone H2A lysine 119, is upregulated in prostate cancer (Van Leenders et al., 2007).
Histone acetylation: Histone acetylation is catalyzed by histone acetyltransferases (HATs) and associated with active gene transcription. The basic charges of the histone tails become neutralized upon acetylation. This causes increased accessibility for further modifications or access to the DNA for binding factors and transcriptional machinery (Hebbes et al., 1992; Turner, 1993; Kouzarides, 2007).
Histone deacetylation, mediated by three classes of HDACs which remove the acetyl group from lysine residues and associated with active gene transcription (Gray and Ekstrom, 2001; Yoshida et al., 2001; Marks et al., 2004). Inappropriate deacetylation can also contribute to cancer progression. HDACs are upregulated in various types of cancer, such as gastric, prostate, oral squamous cell and lung. Over-expression of HDACs can also lead to the transcriptional inactivation of tumor suppressors, such as p53 (Halkidou et al., 2004; Bartling et al., 2005; Song et al., 2005; Sakuma et al., 2006).
HDAC inhibitors are divided into 4 groups based on their structures including hydroxamic acids, cyclic peptides, short chain fatty acids and benzamides (Zheng et al., 2008). HDAC inhibitors have pleiotropic effects including inhibition of angiogenesis, induction of apoptosis and cell cycle arrest (Stearns et al., 2007). The hydroxamic acid group of HDAC inhibitors has been successful in treating both hematologic malignancies and solid tumors. The X-ray crystallography has shown that the catalytic site of HDACs contains a zinc atom. The hydroxamic acid can fit into the catalytic site of HDACs and bind to the zinc atom of this site thereby inhibiting the HDAC (Marks et al., 2000). The shortcoming of these HDAC inhibitors is that a high concentration of drug is required for efficacy resulting in limited use in the clinic (Johnstone, 2002). The list of the HDAC inhibitory agents was shown in Table 3.
| Table 3: | The anti-cancer drugs which exhibit HDAC inhibitory activity |
 | |
Neither of these drugs is as potent as the other classes of HDAC inhibitors and seems to have the greatest effect when used in a combinatorial treatment (Kouraklis and Theocharis, 2006). For example, sodium butyrate and TSA synergize with 1,25-(OH)2-vitamin D3 to inhibit the growth of LNCaP, PC-3 and DU-145 prostate cancer cells by inducing apoptosis (Fruhwald and Plass, 2002).
In prostate cancer, the expression of several genes may be potentially regulated by histone acetylation. Treatment of prostate cancer cells with HDAC inhibitors increased expression of specific genes such as insulin-like growth factor-binding protein 3 (Tsubaki et al., 2002) and carboxypeptidase A3 (Huang et al., 1999) and thus inferred a role for histone acetylation in gene regulation.
One such gene, coxsackie and adenovirus receptor (CAR), is the primary receptor for group C adenoviruses and is important for adenovirus attachment to the cell membrane. In urogenital cancer cells, including the prostate cancer cell line PC-3, activation of the CAR gene is modulated by histone acetylation and can be induced by depsipeptide, an HDAC inhibitor (Pong et al., 2003). Exposing cancer cells to low concentrations of depsipeptide has the functional consequence of preferentially increasing the efficiency of adenoviral transgene expression (Goldsmith et al., 2003).
Another gene regulated by histone modification is the vitamin D receptor. 1,25-(OH)2-vitamin D3 acts to antiproliferative effects in a variety of tumor cells, including those of the prostate (Moffatt et al., 1999; Zhao et al., 2000; Ikeda et al., 2003; Yang and Burnstein, 2003).
Prostate cancer cells that are insensitive to 1,25-(OH)2-vitamin D3 have increased levels of nuclear receptor corepressor SMRT (silencing mediator of retinoid and thyroid), which could result in increased deacetylase activity and decreased transcriptional activity of the vitamin D receptor. In addition, combined treatment of prostate cancer cell lines with the HDAC inhibitor trichostatin A (TSA) and 1,25-(OH)2-vitamin D3 synergistically inhibits cell proliferation. This finding may be useful in the clinical setting, in which use of 1,25-(OH)2-vitamin D3 and its analogs in combination with HDAC inhibitors could activate the vitamin D receptor while minimizing unwanted side effects associated with 1,25-(OH)2- vitamin D3, such as hypocalcemia (Banwell et al., 2003).
Loss of imprinting (LOI): Also, aberrant methylation of imprinted genes can disturb imprinting. LOI of insulin like growth factor 2 (IGF2) is causally involved in Wilms tumors and colorectal cancers through its overexpression (Feinberg and Tycko, 2004). Loss of IGF2 imprinting in colonic mucosae is associated with an elevated risk of colorectal cancers (Cui et al., 1998, 2003; Woodson et al., 2004) and LOI in peripheral lymphocyte was also associated with an increased risk (Cui et al., 2003). A study using a mouse model for loss of IGF2 imprinting showed that LOI caused less differentiation of normal intestinal epithelium (Sakatani et al., 2005).
The DNA methylation and histone modifications are as important interconnected epigenetic regulatory mechanisms that they can influence the gene regulation (Fuks et al., 2003). DNA methylation is involved in gene silencing through binding of methylated DNA binding proteins such as MeCP2 to gene promoter. This interaction then recruits HDAC to methylated promoters. Therefore, DNA methylation event happens first followed by histone deacetylation and then histone methylation (Antequera and Bird, 1999; Stirzaker et al., 2004).
Several studies have shown that the combination of HDAC and DNMT inhibitors can work synergistically to induce the re-expression of such genes like tumor suppressor genes and genes that involved in apoptosis, differentiation, cell growth arrest could enhance the antitumor effects, in vitro (Cameron et al., 1999; Weiser et al., 2001; Ghoshal et al., 2002). For example, phenylbutyrate and 5-Aza-CdR have synergistic effects on reducing lung tumor formation in mice more than 5-Aza-CdR alone (Belinsky et al., 2003). So, DNA methylation inhibitors and HDAC inhibitors are now used together in the clinic after garnering encouraging results in vitro and it is hypothesized that in a phase II trial, using a longer exposure could further increase the response rate (Gore et al., 2006). However, non-specific demethylation has the risk of inducing demethylation of normally methylated sequences, such as retrotransposons and thus retrotranspositions.
Pretreatment with an HDAC inhibitor can greatly increase cytotoxicity in various cell lines when followed by subsequent treatment of a chemotherapeutic drug (kim et al., 2003). Likewise, cisplatin resistant cells from head and neck cancer cell lines can be reprogrammed to become responsive after treatment with Phenyl butyrate (Burkitt and Ljungman, 2008). The increased sensitivity to other drugs after use of an epigenetic drug is encouraging since drug resistance does present a challenge in effective cancer treatment.
As the field of cancer epigenetic advances, a better understanding of the DNA methylation and post-translational histone modifications which play central roles in gene regulation is under developing. The future directions for the development of epigenetic drugs and epigenetic markers will depend on the elucidation of their mechanisms and the downstream effects of treatment.
The use of FDA approved epigenetic drugs has gained momentum and has proven useful in some tumors. The combinatorial use of DNA methylation inhibitors, HDAC inhibitors and non-epigenetic chemotherapeutic drugs in an effort increase response rates and maximize the efficacy of these drugs in the clinic and may have synergistic effects in re-establishing the expression of tumor suppressor genes. However, much work remains in designing drugs that will be more stable, less toxic and more specific in their enzyme inhibition. Future epigenetic drugs can also be designed to target histone methyltransferases, histone demethylases or other chromatin modifiers not yet discovered.
I am grateful to all the research scientists, especially to Professors. Andrew Feinberg and Stephen Baylin from School of Medicine at Johns Hopkins University for several years studies on cancer epigenetic.