Research Article
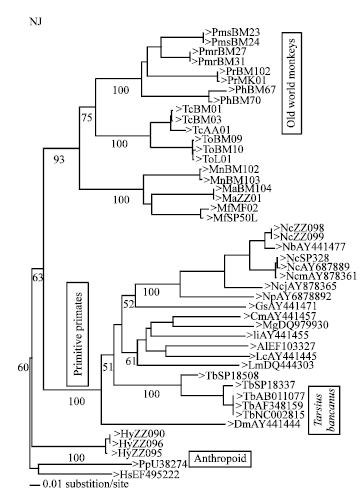
Phylogenetic Position of Tarsius bancanus Based on Partial Cytochrome b DNA Sequences
School of Environmental and Natural Resource Sciences, Faculty of Science and Technology, Universiti Kebangsaan Malaysia, 43600 Bangi, Selangor, Malaysia
S.J. Lee
School of Environmental and Natural Resource Sciences, Faculty of Science and Technology, Universiti Kebangsaan Malaysia, 43600 Bangi, Selangor, Malaysia
M. Lakim
The Board of Trustees of Sabah Parks, Kinabalu Park, Kota Kinabalu, Sabah, Malaysia
A. Ampeng
School of Environmental and Natural Resource Sciences, Faculty of Science and Technology, Universiti Kebangsaan Malaysia, 43600 Bangi, Selangor, Malaysia
M.C. Mahani
School of Environmental and Natural Resource Sciences, Faculty of Science and Technology, Universiti Kebangsaan Malaysia, 43600 Bangi, Selangor, Malaysia