Research Article
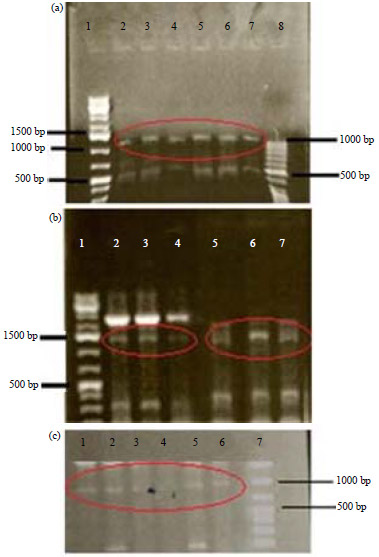
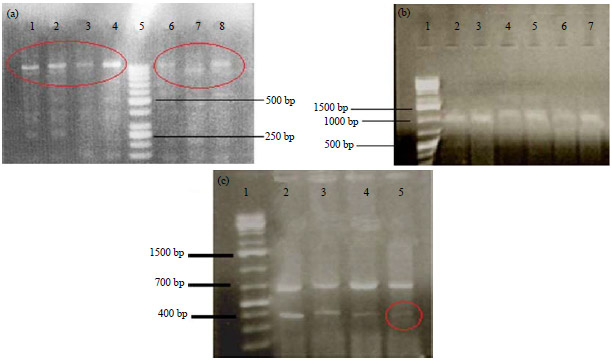
Genotypic Characterization of Phytophthora infestans from Mauritius using Random Amplified Polymorphic DNA (RAPD), Mitochondrial Haplotyping and Mating Type Analysis
Department of Biosciences, Faculty of Science, University of Mauritius, 80837 Reduit, Mauritius
Nawsheen Taleb-Hossenkhan
Department of Biosciences, Faculty of Science, University of Mauritius, 80837 Reduit, Mauritius
LiveDNA: 222.18548