Research Article
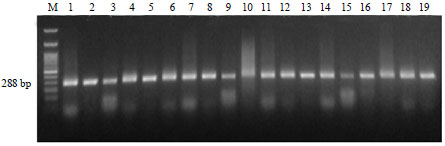
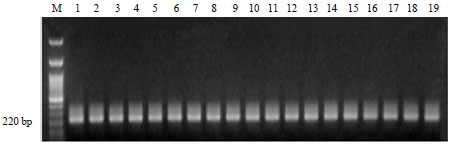
Detection and Molecular Characterization of some Egyptian Isolates of Ralstonia solanacearum by Nested-PCR and PCR-RFLP Analyses
Department of Plant Pathology, Faculty of Agriculture, Alexandria University, El-Shatby, 21545, Alexandria, Egypt