Research Article
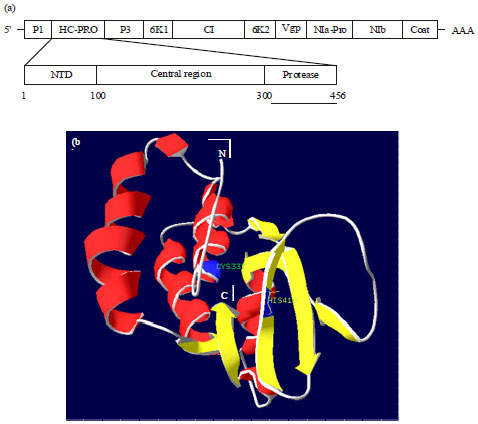
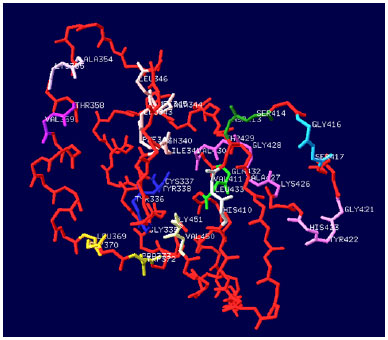
Sequence Motif Comparison and Homology Modeling of Helper Component Proteinase (HC-Pro) of Banana bract mosaic virus
ICAR-National Research Centre for Banana, Thogamalai Road Thayanur Post Tiruchirapalli, Tamil Nadu, India
Balasubramanian Velusamy
ICAR-National Research Centre for Banana, Thogamalai Road Thayanur Post Tiruchirapalli, Tamil Nadu, India
Selvarajan Ramasamy
ICAR-National Research Centre for Banana, Thogamalai Road Thayanur Post Tiruchirapalli, Tamil Nadu, India